Abstract
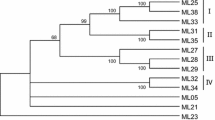
Based on conserved regions of a sequence of a previously isolated powdery mildew (Pm) resistance gene, Pm3b, from hexaploid wheat (Triticum aestivum L.), a pair of primers was designed, and 11 resistance gene analogs (RGAs) were obtained using a polymerase chain reaction-based cloning approach. Three RGAs were deemed as pseudogenes, while the remaining eight corresponded to protein-encoding genes. At the nucleotide level, all these RGAs shared a sequence identity of 99% and showed 86% sequence identity with Pm3b. Phylogenetic analysis revealed that the eight protein-encoding genes were paraphyletic with Pm3 alleles. Positively selected sites were identified using the Selecton 2.1 program, and these were located on secondary structures. Based on these findings, the following two inferences could be made on the mode of evolution of the nucleotide-binding site and leucine-rich repeat (LRR) class of resistance genes. First, the majority of evolution events must have occurred primarily in the LRR domain, and this might have contributed to an increase in the proportion of evolution events in other domains. Second, these evolutionary events in the LRR domain must have occurred initially in secondary structures and then in the β-sheet. The crystalline structure models of RGAs were constructed.




Similar content being viewed by others
Abbreviations
- BLAST:
-
Basic local alignment and search tool
- CC:
-
Coiled coil
- DNA:
-
Deoxyribonucleic acid
- InDel:
-
Insertion/deletion
- LRR:
-
Leucine-rich repeat
- NBS:
-
Nucleotide-binding site
- PCR:
-
Polymerase chain reaction
- Pm:
-
Powdery mildew
- RGAs:
-
Resistance gene analogs
- TIR:
-
Toll-interleukin receptor-like
References
Altschul SF, Madden TL, Schäffer AA et al (1997) Gapped BLAST and PSI-BLAST: a new generation of protein database search programs. Nucleic Acids Res 25:3389–3402, doi:10.1093/nar/25.17.3389
Bai J, Pennill LA, Ning J, Lee SW et al (2002) Diversity in nucleotide binding site-leucine-rich repeat genes in cereals. Genome Res 12:1871–1884, doi:10.1101/gr.454902
Choe J, Kelker MS, Wilson IA (2005) Crystal structure of human toll-like receptor 3 (TLR3) ectodomain. Science 309:581–585, doi:10.1126/science.1115253
Dangl JL, Jones JDG (2001) Plant pathogens and integrated defence responses to infection. Nature 411:826–833, doi:10.1038/35081161
Doron-Faigenboim A, Pupko T (2006) A combined empirical and mechanistic codon model. Mol Biol Evol 24:388–397, doi:10.1093/molbev/msl175
Doron-Faigenboim A, Stern A, Mayrose I et al (2005) Selecton: a server for detecting evolutionary forces at a single amino-acid site. Bioinformatics 21(9):2101–2103, doi:10.1093/bioinformatics/bti259
Ellis J, Dodds P, Pryor T (2000a) Structure, function and evolution of plant disease resistance genes. Curr Opin Plant Biol 3:278–284, doi:10.1016/S1369-5266(00)00080-7
Ellis J, Dodds P, Pryor T (2000b) The generation of plant disease resistance gene specificities. Trends Plant Sci 5:373–379, doi:10.1016/S1360-1385(00)01694-0
Endo T, Ikeo K, Gojobori T (1996) Large-scale search for genes on which positive selection may operate. Mol Biol Evol 13:685–690
Flavell RB, Bennett MD, Smith JB et al (1974) Genome size and proportion of repeated nucleotide sequence DNA in plants. Biochem Genet 12:257–269, doi:10.1007/BF00485947
Huang L, Brooks SA, Li W et al (2003) Map-based cloning of leaf rust resistance gene Lr21 from the large and polyploid genome of bread wheat. Genetics 164:655–664
Jia Y, McAdams SA, Bryan GT et al (2000) Direct interaction of resistance gene and avirulence gene products confers rice blast resistance. EMBO J 19:4004–4014, doi:10.1093/emboj/19.15.4004
Jones DA, Jones JDG (1997) The role of leucine-rich repeat proteins in plant defences. Adv Bot Res 24:90–167
Kobe B, Kajava AV (2001) The leucine-rich repeat as a protein recognition motif. Curr Opin Struct Biol 11:725–732, doi:10.1016/S0959-440X(01)00266-4
Kumar S, Tamura K, Jakobsen IB et al (2001) MEGA2: molecular evolutionary genetics analysis software. Arizona State University, Tempe, Arizona, USA
Lupas A, Van Dyke M, Stock J (1991) Predicting coiled coils from protein sequences. Science 252:1162–1164, doi:10.1126/science.252.5009.1162
Meyers BC, Shen KA, Rohani P et al (1998) Receptor-like genes in the major resistance locus of lettuce are subject to divergent selection. Plant Cell 11:1833–1846
Meyers BC, Dickerman AW, Michelmore RW et al (1999) Plant disease resistance genes encode members of an ancient and diverse protein family within the nucleotide-binding superfamily. Plant J 20:317–332, doi:10.1046/j.1365-313X.1999.t01-1-00606.x
Meyers BC, Kozik A, Griego A et al (2003) Genome-wide analysis of NBS–LRR-encoding genes in Arabidopsis. Plant Cell 15:809–834, doi:10.1105/tpc.009308
Mondragón-Palomino M, Meyers BC, Michelmore RW et al (2002) Pattern of positive selection in the complete NBS–LRR gene family of Arabidopsis thaliana. Genome Res 12:1305–1315, doi:10.1101/gr.159402
Murray MG, Thompson WF (1980) The isolation of high molecular weight plant DNA. Nucleic Acids Res 8:4321–4325, doi:10.1093/nar/8.19.4321
Sambrook J, Frisch EF, Maniatis T (1989) Molecular cloning: a laboratory manual, 2nd edn. Cold Spring Harbour Laboratory, New York, pp 19–22
Srichumpa P, Brunner S, Keller B et al (2005) Allelic series of four powdery mildew resistance genes at the Pm3 locus in hexaploid bread wheat. Plant Physiol 139:885–895, doi:10.1104/pp.105.062406
Swanson WJ, Nielsen R, Yang Q (2003) Pervasive adaptive evolution in mammalian fertilization proteins. Mol Biol Evol 20:18–20
Tian DC, Wang Q, Zhang PF et al (2008) Single-nucleotide mutation rate increases close to insertions/deletions in eukaryotes. Nature 455:105–108, doi:10.1038/nature07175
Wang GL, Ruan DL, Song WY, Sideris S et al (1998) Xa21D encodes a receptor-like molecule with a leucine-rich repeat domain that determines race-specific recognition and is subject to adaptive evolution. Plant Cell 10:765–779
Yahiaoui N, Srichumpa P, Dudler R et al (2004) Genome analysis at different ploidy levels allows cloning of the powdery mildew resistance gene Pm3b from hexaploid wheat. Plant J 37:528–538, doi:10.1046/j.1365-313X.2003.01977.x
Yahiaoui N, Brunner S, Keller B (2006) Rapid generation of new powdery mildew resistance genes after wheat domestication. Plant J 47:85–98, doi:10.1111/j.1365-313X.2006.02772.x
Yan N, Chai J, Lee ES et al (2005) Structure of the CED-4–CED-9 complex provides insights into programmed cell death in Caenorhabditis elegans. Nature 437:831–837, doi:10.1038/nature04002
Yang Z, Nielsen R, Goldman N, Pedersen AM (2000) Codon-substitution models for heterogeneous selection pressure at amino acid sites. Genetics 155:431–449
Acknowledgments
The authors thank Prof. Mark E. Sorrels (College of Agriculture and Life Sciences, Cornell University) for the critical review of the manuscript. We also wish to thank two anonymous reviewers for helpful comments that improved the manuscript. This project was supported by the Early Stage of China Key Development Project for Basic Research (973) (2007CB116208).
Author information
Authors and Affiliations
Corresponding author
Additional information
De-Shun Feng and Yan Li contributed equally to this paper.
Sequence data of 2Q2, 2Q3, 2Q4, 2Q7, 2Q9, 2Q11, 2Q12, 12Q11, 15Q1, and 15Q7 from this article have been deposited at GenBank under accession numbers EF157980, EF157981, EF157982, EF157983, EF157984, EF157985, EF157986, EF157987, EF157988, EF157989, and EF157990.
Rights and permissions
About this article
Cite this article
Feng, DS., Li, Y., Wang, HG. et al. Isolation and Evolution Mode Analysis of NBS–LRR Resistance Gene Analogs from Hexaploid Wheat. Plant Mol Biol Rep 27, 266–274 (2009). https://doi.org/10.1007/s11105-008-0080-1
Published:
Issue Date:
DOI: https://doi.org/10.1007/s11105-008-0080-1