Abstract
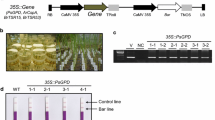
We present a simple and rapid method for screening second-generation transgenic rice plants (T1) to identify homozygous plants. The plasmid (pfd11) used for rice transformation contains a partially deleted cytochrome c gene (cyc) for comparing with the endogenous cyc for copy number. After polymerase chain reaction (PCR) amplification of a segment of the cyc in transgenic rice DNA followed by agarose gel electrophoresis, two specific bands are obtained. The upper band represents the endogenous cyc, and the lower band represents the partially deleted cyc in the transgene. The first-generation plants (T0) that harbor a single copy of the transgene are selected based on the fact that the density of the lower band is half as dense as the upper band. Next, only plants harboring a single copy of the transgene are advanced to the second generation (T1). The same PCR procedure is used again, and homozygous T1 plants are easily identified from samples in which the intensity of the two bands is the same.



Similar content being viewed by others
References
Bubner B, Gase K, Baldwin IT. Two-fold differences are the detection limit for determining transgene copy numbers in plants by real-time PCR. BMC Biotechnol 2004;13:4–14.
Chen X, Shang J, Chen D, Lei C, Zou Y, Zhai W, Liu G, Xu J, Ling Z, Cao G, Ma B, Wang Y, Zhao X, Li S, Zhu L. A B-lectin receptor kinase gene conferring rice blast resistance. Plant J 2006;46(5):794–804.
Choi HW, Lemaux PG, Cho MJ. Use of fluorescence in situ hybridization for gross mapping of transgenes and screening for homozygous plants in transgenic barley (Hordeum vulgare L.). Theor Appl Genet 2002;106(1):92–100.
Delmer DP. Agriculture in the developing world: connecting innovations in plant research to downstream applications. Proc Natl Acad Sci U S A 2005;102(44):15739–46.
Garg AK, Kim JK, Owens TG, Ranwala AP, Choi YD, Kochian LV, Wu JR. Trehalose accumulation in rice plants confers high tolerance levels to different abiotic stresses. Proc Natl Acad Sci USA 2002;99(25):15898–903.
Hu H, Da M, Yao J, Xiao B, Li X, Zhang Q, Xiong L. Overexpressing a NAM, ATAF, and CUC (NAC) transcription factor enhances drought resistance and salt tolerance in rice. Proc Natl Acad Sci USA 2006;103(35):12987–92.
Kathuria H, Giri J, Tyagi H, Tyagi AK. Advances in transgenic rice biotechnology. Crit Rev Plant Sci 2007;26:65–103.
Lee SC, Kim SH, An SH, Yi SY, Hwang BK. Identification and functional expression of the pepper pathogen-induced gene, CAPIP2, involved in disease resistance and drought and salt stress tolerance. Plant Mol Biol 2006;62:151–64.
Li F, Dey M, He CK, Sangwan V, Wu X, Wu R. Rapid PCR-based determination of transgene copy number in rice. Plant Mol Biol Report 2003;21:73–80.
Murashige T, Skoog F. A revised medium for rapid growth and bioassays with tobacco tissue cultures. Plant Physiol 1962;15:473–97.
Qin F, Sakuma Y, Li J, Liu Q, Li YQ, Shinozaki K, Yamaguchi-Shinozaki K. Cloning and functional analysis of a novel DREB1/CBF transcription factor involved in cold-responsive gene expression in Zea mays L. Plant Cell Physiol 2004;45(8):1042–52.
Roy M, Wu R. Arginine decarboxylase transgene expression and analysis of environmental stress tolerance in transgenic rice. Plant Sci 2001;160:869–75.
Umezawa T, Fujita M, Fujita Y, Yamaguchi-Shinozaki K, Shinozaki K. Engineering drought tolerance in plants: discovering and tailoring genes to unlock the future. Curr Opin Biotechnol 2006;17:113–22.
Acknowledgement
We thank Mu He, an undergraduate student at Cornell University, for the assistance in this study.
Author information
Authors and Affiliations
Corresponding author
Rights and permissions
About this article
Cite this article
Zhu, ZG., Wu, R. A Simple and Rapid PCR-based Procedure for Identifying Homozygous Transgenic Rice Plants. Plant Mol Biol Rep 26, 53–59 (2008). https://doi.org/10.1007/s11105-008-0022-y
Published:
Issue Date:
DOI: https://doi.org/10.1007/s11105-008-0022-y