Abstract
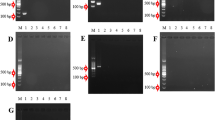
The wide development of single nucleotide polymorphism (SNP) markers also in non-model species increases the need for inexpensive methods that do not require sophisticated equipment and time for optimization. This work presents a new method for polymerase chain reaction (PCR) amplification of multiple specific alleles (PAMSA), which allows efficient discrimination of SNP polymorphisms in one reaction tube with standard PCR conditions. This improved PAMSA requires only three unlabeled primers: a common reverse primer and two allele-specific primers having a tail of different length to differentiate the two SNP alleles by the size of amplification products on agarose gel. A destabilizing mismatch within the five bases of the 3′ end is also added to improve the allele specificity. To validate the accuracy of this method, 94 full-sib individuals were genotyped with three SNPs and compared to the genotypes obtained by cleaved amplified polymorphic sequence (CAPS) or derived CAPS. This method is flexible, inexpensive, and well suited for high throughput and automated genotyping.


Similar content being viewed by others
Abbreviations
- ARMS:
-
amplification refractory mutation system
- AS-PCR:
-
allele-specific polymerase chain reaction
- Bi-PASA:
-
bidirectional-PASA
- CAPS:
-
cleaved amplification polymorphic site
- dCAPS:
-
derived cleaved amplification polymorphic site
- PAMSA:
-
PCR amplification of multiple specific alleles
- PASA:
-
PCR allele-specific amplification
- SNP:
-
single nucleotide polymorphisms
References
Ahmadian A, Gharizadeh B, O’Meara D, Odeberg J, Lundeberg J. Genotyping by apyrase-mediated allele-specific extension. Nucl Acids Res 2001;29:e121.
Bundock PC, Cross MJ, Shapter FM, Henry RJ. Robust allele-specific polymerase chain reaction markers developed for single nucleotide polymorphisms in expressed barley sequences. Theor Appl Genet 2005;112:358–65.
Day JP, Bergstrom D, Hammer RP, Barany F. Nucleotide analogs facilitate base conversion with 3′ mismatch primers. Nucl Acids Res 1999;27:1810–18.
Dutton C, Sommer SS. Simultaneous detection of multiple single-base alleles at a polymorphic site. BioTechniques 1991;11:700–7002.
Gaudet M, Jorge V, Paolucci I, Beritognolo I, Scarascia Mugnozza G, Sabatti M. Genetic linkage maps of Populus nigra L. including AFLPs, SSRs, SNPs, and sex trait. Tree Genet Genome 2007. DOI 10.1007/s11295-007-0085-1.
Germer S, Higuchi R. Single-tube genotyping without oligonucleotide probes. Genome Res 1999;9:72–8.
Gupta PK, Roy JK, Prasad M. Single nucleotide polymorphisms: a new paradigm for molecular marker technology and DNA polymorphism detection with emphasis on their use in plants. Curr Sci India 2001;80:524–35.
Gut IG. Automation in genotyping of single nucleotide polymorphisms. Hum Mutat 2001;17:475–92.
Hansson B, Kawabe A. A simple method to score single nucleotide polymorphisms based on allele-specific PCR and primer-induced fragment-length variation. Mol Ecol Notes 2005;5:692–6.
Hinten GN, Hale MC, Gratten J, Mossman JA, Lowder BV, Mann MK, Slate J. SNP-SCALE: SNP scoring by colour and length exclusion. Mol Ecol Notes 2007;7:377–88.
Imyanitov EN, Buslov KG, Suspitsin EN, Kuligina ESh, Belogubova EV, Grigoriev MY et al. Improved reliability of allele-specific PCR. BioTechniques 2002;33:484–90.
Ishiguro A, Kubota T, Soya Y, Sasaki H, Yagyu O, Takarada Y et al. High-throughput detection of multiple genetic polymorphisms influencing drug metabolism with mismatch primers in allele-specific polymerase chain reaction. Anal Biochem 2005;337:256–61.
Iwaki K, Nishida J, Yanagisawa T, Yoshida H, Kato K. Genetic analysis of Vrn-B1 for vernalization requirement by using linked dCAPS markers in bread wheat (Triticum aestivum L.). Theor Appl Genet 2002;104:571–6.
Kim MY, Van K, Lestari P, Moon JK, Lee SH. SNP identification and SNAP marker development for a GmNARK gene controlling supernodulation in soybean. Theor Appl Genet 2005;110:1003–10.
Kwok S, Kellogg DE, McKinney N, Spasic D, Goda L, Levenson C, Sninsky JJ. Effects of primer-template mismatches on the polymerase chain reaction: human immunodeficiency virus type 1 model studies. Nucl Acids Res 1990;18:999–1005.
Liu Q, Thorland EC, Heit JA, Sommer SS. Overlapping PCR for bidirectional PCR amplification of specific alleles: a rapid one-tube method for simultaneously differentiating homozygotes and heterozygotes. Genome Res 1997;7:389–98.
Michaels SD, Amisino RM. A robust method for the detecting single-nucleotide changes as polymorphic markers by PCR. Plant J 1998;14:381–5.
Neff MM, Neff JD, Chory J, Pepper AE. dCAPS, a simple technique for the genetic analysis of single nucleotide polymorphisms: experimental applications in Arabidopsis thaliana genetics. Plant J 1998;14:387–92.
Neff MM, Turk E, Kalishman M. Web-based primer design for single nucleotide polymorphism analysis. Trends Genet 2002;18:613–5.
Okimoto R, Dodgson JB. Improved PCR amplification of multiple specific alleles (PAMSA) using internally mismatched primers. BioTechniques 1996;21:20–6.
Sachidanandam R, Weissman D, Schmidt SC, Kakol JM, Stein LD, Marth G et al. A map of human genome sequence variation containing 1.42 million single nucleotide polymorphisms. Nature 2001;409:928–33.
Sasvari-Szekely M, Gerstner A, Ronai Z, Staub M, Guttman A. Rapid genotyping of factor V Leiden mutation using single-tube bidirectional allele-specific amplification and automated ultrathin-layer agarose gel electrophoresis. Electrophoresis 2000;21:816–21.
Soleimani VD, Baum BR, Johnson DA. Efficient validation of single nucleotide polymorphisms in plants by allele-specific PCR, with an example from barley. Plant Mol Biol Rep 2003;21:281–8.
Waterfall CM, Cobb BD. Single tube genotyping of sickle cell anaemia using PCR-based SNP analysis. Nucl Acids Res 2001;29:e119.
Waterfall CM, Cobb BD. SNP genotyping using single-tube fluorescent bidirectional PCR. BioTechniques 2002;33:80–90.
Wu WM, Tsai HJ, Pang JHS, Wang HS, Hong HS, Lee YS. Touchdown thermocycling program enables a robust single nucleotide polymorphism typing method based on allele-specific real-time polymerase chain reaction. Anal Biochem 2005;339:290–6.
Yamanaka S, Nakamura I, Watanabe KN, Sato YI. Identification of SNPs in the waxy gene among glutinous rice cultivars and their evolutionary significance during the domestication process of rice. Theor Appl Genet 2004;108:1200–4.
Yanagisawa T, Kiribuchi-Otobe C, Hirano H, Suzuki Y, Fujita M. Detection of single nucleotide polymorphism (SNP) controlling the waxy character in wheat by using a derived cleaved amplified polymorphic sequence (dCAPS) marker. Theor Appl Genet 2003;107:84–8.
Acknowledgments
The authors thank Véronique Jorge and Claudia Mattioni for their critical reading of the manuscript, Dermot Browne for correction of English writing, and Isacco Beritognolo for his technical advice and assistance. This work was supported by grants from the European Project QLK5-CT-2002-00953 (POPYOMICS) and the Italian project MIUR PRIN 2005072892_001.
Author information
Authors and Affiliations
Corresponding author
Rights and permissions
About this article
Cite this article
Gaudet, M., Fara, AG., Sabatti, M. et al. Single-reaction for SNP Genotyping on Agarose Gel by Allele-specific PCR in Black Poplar (Populus nigra L.). Plant Mol Biol Rep 25, 1–9 (2007). https://doi.org/10.1007/s11105-007-0003-6
Published:
Issue Date:
DOI: https://doi.org/10.1007/s11105-007-0003-6