Key message
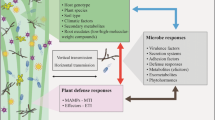
Rice- Phomopsis liquidambaris mutualistic interaction, under low nitrogen conditions, leads to increased plant growth and nitrogen uptake, as a consequence of host transcriptional reprogramming.
Abstract
Fungal endophytes establish symbiotic relationships with host plants, which results in a mutual growth benefit. However, little is known about the plant genetic response underpinning endophyte colonization. Phomopsis liquidambaris usually lives as an endophyte in a wide range of asymptomatic hosts and promotes biotic and abiotic stress resistance. In this study, we show that under low nitrogen conditions P. liquidambaris promotes rice growth in a hydroponic system, which is free of other microorganisms. In order to gain insights into the mechanisms of plant colonization by P. liquidambaris under low nitrogen conditions, we compared root and shoot transcriptome profiles of root-inoculated rice at different colonization stages. We determined that genes related to plant growth promotion, such as gibberellin and auxin related genes, were up-regulated at all developmental stages both locally and systemically. The largest group of up-regulated genes (in both roots and shoots) were related to flavonoid biosynthesis, which is involved in plant growth as well as antimicrobial compounds. Furthermore, genes encoding plant defense-related endopeptidase inhibitors were strongly up-regulated at the early stage of colonization. Together, these results provide new insights into the molecular mechanisms of plant-microbe mutualism and the promotion of plant growth by a fungal endophyte under nitrogen-deficient conditions.






Similar content being viewed by others
Availability of data and material
The RNA sequencing data generated in this work have been deposited to the NCBI’s Sequence Read Archive under BioProject PRJNA737734 (http://www.ncbi.nlm.nih.gov/bioproject/737734).
Code Availability
Not applicable.
References
Banhara A, Ding Y, Kuhner R, Zuccaro A, Parniske M (2015) Colonization of root cells and plant growth promotion by Piriformospora indica occurs independently of plant common symbiosis genes. Front Plant Sci 6:667. doi: https://doi.org/10.3389/fpls.2015.00667
Busby PE, Ridout M, Newcombe G (2016) Fungal endophytes: modifiers of plant disease. Plant Mol Biol 90:645–655. doi: https://doi.org/10.1007/s11103-015-0412-0
Canfield DE, Glazer AN, Falkowski PG (2010) The evolution and future of Earth’s nitrogen cycle. Science 330:192–196. doi: https://doi.org/10.1126/science.1186120
Chen Y, Ren CG, Yang B, Peng Y, Dai CC (2013) Priming effects of the endophytic fungus Phomopsis liquidambari on soil mineral N transformations. Microb Ecol 65:161–170. doi: https://doi.org/10.1007/s00248-012-0093-z
Christian N, Herre EA, Clay K (2019) Foliar endophytic fungi alter patterns of nitrogen uptake and distribution in Theobroma cacao. New Phytol 222:1573–1583. doi: https://doi.org/10.1111/nph.15693
de Hoon MJ, Imoto S, Nolan J, Miyano S (2004) Open source clustering software. Bioinformatics 20:1453–1454. doi: https://doi.org/10.1093/bioinformatics/bth078
De Vleesschauwer D, Xu J, Hofte M (2014) Making sense of hormone-mediated defense networking: from rice to Arabidopsis. Front Plant Sci 5:611. doi: https://doi.org/10.3389/fpls.2014.00611
Del Valle I, Webster TM, Cheng HY, Thies JE, Kessler A, Miller MK, Ball ZT, MacKenzie KR, Masiello CA, Silberg JJ, Lehmann J (2020) Soil organic matter attenuates the efficacy of flavonoid-based plant-microbe communication. Sci Adv 6:eaax8254. doi: https://doi.org/10.1126/sciadv.aax8254
Dinkins RD, Nagabhyru P, Graham MA, Boykin D, Schardl CL (2017) Transcriptome response of Lolium arundinaceum to its fungal endophyte Epichloe coenophiala. New Phytol 213:324–337. doi: https://doi.org/10.1111/nph.14103
Dodd IC, Ruiz-Lozano JM (2012) Microbial enhancement of crop resource use efficiency. Curr Opin Biotechnol 23:236–242. doi: https://doi.org/10.1016/j.copbio.2011.09.005
Dupont PY, Eaton CJ, Wargent JJ, Fechtner S, Solomon P, Schmid J, Day RC, Scott B, Cox MP (2015) Fungal endophyte infection of ryegrass reprograms host metabolism and alters development. New Phytol 208:1227–1240. doi: https://doi.org/10.1111/nph.13614
Eisen MB, Spellman PT, Brown PO, Botstein D (1998) Cluster analysis and display of genome-wide expression patterns. Proc Natl Acad Sci U S A 95:14863–14868
Fesel PH, Zuccaro A (2016) Dissecting endophytic lifestyle along the parasitism/mutualism continuum in Arabidopsis. Curr Opin Microbiol 32:103–112. doi: https://doi.org/10.1016/j.mib.2016.05.008
Foo E, Ross JJ, Jones WT, Reid JB (2013) Plant hormones in arbuscular mycorrhizal symbioses: an emerging role for gibberellins. Ann Bot 111:769–779. doi: https://doi.org/10.1093/aob/mct041
Gao Y, Wu M, Zhang M, Jiang W, Ren X, Liang E, Zhang D, Zhang C, Xiao N, Li Y, Dai Y, Chen J (2018) A maize phytochrome-interacting factors protein ZmPIF1 enhances drought tolerance by inducing stomatal closure and improves grain yield in Oryza sativa. Plant Biotechnol J 16:1375–1387. doi: https://doi.org/10.1111/pbi.12878
Gaudinier A, Rodriguez-Medina J, Zhang L, Olson A, Liseron-Monfils C, Bagman AM, Foret J, Abbitt S, Tang M, Li B, Runcie DE, Kliebenstein DJ, Shen B, Frank MJ, Ware D, Brady SM (2018) Transcriptional regulation of nitrogen-associated metabolism and growth. Nature 563:259–264. doi: https://doi.org/10.1038/s41586-018-0656-3
Hiruma K, Gerlach N, Sacristan S, Nakano RT, Hacquard S, Kracher B, Neumann U, Ramirez D, Bucher M, O’Connell RJ, Schulze-Lefert P (2016) Root Endophyte Colletotrichum tofieldiae Confers Plant Fitness Benefits that Are Phosphate Status Dependent. Cell 165:464–474. doi: https://doi.org/10.1016/j.cell.2016.02.028
Kanehisa M, Araki M, Goto S, Hattori M, Hirakawa M, Itoh M, Katayama T, Kawashima S, Okuda S, Tokimatsu T, Yamanishi Y (2008) KEGG for linking genomes to life and the environment. Nucleic Acids Res 36:D480–484. doi: https://doi.org/10.1093/nar/gkm882
Kawahara Y, de la Bastide M, Hamilton JP, Kanamori H, McCombie WR, Ouyang S, Schwartz DC, Tanaka T, Wu J, Zhou S, Childs KL, Davidson RM, Lin H, Quesada-Ocampo L, Vaillancourt B, Sakai H, Lee SS, Kim J, Numa H, Itoh T, Buell CR, Matsumoto T (2013) Improvement of the Oryza sativa Nipponbare reference genome using next generation sequence and optical map data. Rice 6:4. doi: https://doi.org/10.1186/1939-8433-6-4
Kiba T, Feria-Bourrellier AB, Lafouge F, Lezhneva L, Boutet-Mercey S, Orsel M, Brehaut V, Miller A, Daniel-Vedele F, Sakakibara H, Krapp A (2012) The Arabidopsis nitrate transporter NRT2.4 plays a double role in roots and shoots of nitrogen-starved plants. Plant Cell 24:245–258. doi: https://doi.org/10.1105/tpc.111.092221
Kiba T, Inaba J, Kudo T, Ueda N, Konishi M, Mitsuda N, Takiguchi Y, Kondou Y, Yoshizumi T, Ohme-Takagi M, Matsui M, Yano K, Yanagisawa S, Sakakibara H (2018) Repression of Nitrogen Starvation Responses by Members of the Arabidopsis GARP-Type Transcription Factor NIGT1/HRS1 Subfamily. Plant Cell 30:925–945. doi: https://doi.org/10.1105/tpc.17.00810
Kim JY, Park SC, Hwang I, Cheong H, Nah JW, Hahm KS, Park Y (2009) Protease inhibitors from plants with antimicrobial activity. Int J Mol Sci 10:2860–2872. doi: https://doi.org/10.3390/ijms10062860
Kim D, Langmead B, Salzberg SL (2015) HISAT: a fast spliced aligner with low memory requirements. Nat Methods 12:357–360. doi: https://doi.org/10.1038/nmeth.3317
Krapp A, Berthome R, Orsel M, Mercey-Boutet S, Yu A, Castaings L, Elftieh S, Major H, Renou JP, Daniel-Vedele F (2011) Arabidopsis roots and shoots show distinct temporal adaptation patterns toward nitrogen starvation. Plant Physiol 157:1255–1282. doi: https://doi.org/10.1104/pp.111.179838
Laluk K, Mengiste T (2011) The Arabidopsis extracellular UNUSUAL SERINE PROTEASE INHIBITOR functions in resistance to necrotrophic fungi and insect herbivory. Plant J 68:480–494. doi: https://doi.org/10.1111/j.1365-313X.2011.04702.x
Langmead B, Trapnell C, Pop M, Salzberg SL (2009) Ultrafast and memory-efficient alignment of short DNA sequences to the human genome. Genome Biol 10:R25. doi: https://doi.org/10.1186/gb-2009-10-3-r25
Li B, Dewey CN (2011) RSEM: accurate transcript quantification from RNA-Seq data with or without a reference genome. BMC Bioinformatics 12:323. doi: https://doi.org/10.1186/1471-2105-12-323
Li X, Zhou J, Xu RS, Meng MY, Yu X, Dai CC (2018) Auxin, Cytokinin, and Ethylene Involved in Rice N Availability Improvement Caused by Endophyte Phomopsis liquidambari. J Plant Growth Regul 37:128–143. doi: https://doi.org/10.1007/s00344-017-9712-8
Livak KJ, Schmittgen TD (2001) Analysis of relative gene expression data using real-time quantitative PCR and the 2(-Delta Delta C(T)) Method. Methods 25:402–408. doi: https://doi.org/10.1006/meth.2001.1262
Lyons PC, Evans JJ, Bacon CW (1990) Effects of the Fungal Endophyte Acremonium coenophialum on Nitrogen Accumulation and Metabolism in Tall Fescue. Plant Physiol 92:726–732. doi: https://doi.org/10.1104/pp.92.3.726
Martinez M, Abraham Z, Gambardella M, Echaide M, Carbonero P, Diaz I (2005) The strawberry gene Cyf1 encodes a phytocystatin with antifungal properties. J Exp Bot 56:1821–1829. doi: https://doi.org/10.1093/jxb/eri172
Peng X, Hu Y, Tang X, Zhou P, Deng X, Wang H, Guo Z (2012) Constitutive expression of rice WRKY30 gene increases the endogenous jasmonic acid accumulation, PR gene expression and resistance to fungal pathogens in rice. Planta 236:1485–1498. doi: https://doi.org/10.1007/s00425-012-1698-7
Pernas M, Lopez-Solanilla E, Sanchez-Monge R, Salcedo G, Rodriguez-Palenzuela P (1999) Antifungal activity of a plant cystatin. Mol Plant-Microbe Interact 12:624–627. Doi https://doi.org/10.1094/Mpmi.1999.12.7.624
Pieterse CM, Zamioudis C, Berendsen RL, Weller DM, Van Wees SC, Bakker PA (2014) Induced systemic resistance by beneficial microbes. Annu rev phytopathol 52:347–375. doi: https://doi.org/10.1146/annurev-phyto-082712-102340
Porras-Alfaro A, Bayman P (2011) Hidden fungi, emergent properties: endophytes and microbiomes. Annu rev phytopathol 49:291–315. doi: https://doi.org/10.1146/annurev-phyto-080508-081831
Qu LJ, Chen J, Liu M, Pan N, Okamoto H, Lin Z, Li C, Li D, Wang J, Zhu G, Zhao X, Chen X, Gu H, Chen Z (2003) Molecular cloning and functional analysis of a novel type of Bowman-Birk inhibitor gene family in rice. Plant Physiol 133:560–570. doi: https://doi.org/10.1104/pp.103.024810
Rasmussen S, Parsons AJ, Bassett S, Christensen MJ, Hume DE, Johnson LJ, Johnson RD, Simpson WR, Stacke C, Voisey CR, Xue H, Newman JA (2007) High nitrogen supply and carbohydrate content reduce fungal endophyte and alkaloid concentration in Lolium perenne. New Phytol 173:787–797. doi: https://doi.org/10.1111/j.1469-8137.2006.01960.x
Rehman S, Aziz E, Akhtar W, Ilyas M, Mahmood T (2017) Structural and functional characteristics of plant proteinase inhibitor-II (PI-II) family. Biotechnol Lett 39:647–666. doi: https://doi.org/10.1007/s10529-017-2298-1
Saikkonen K, Faeth SH, Helander M, Sullivan TJ (1998) Fungal endophytes: A continuum of interactions with host plants. Annu Rev Ecol Syst 29:319–343. DOI https://doi.org/10.1146/annurev.ecolsys.29.1.319
Saldanha AJ (2004) Java Treeview–extensible visualization of microarray data. Bioinformatics 20:3246–3248. doi: https://doi.org/10.1093/bioinformatics/bth349
Siddikee MA, Zereen MI, Li CF, Dai CC (2016) Endophytic fungus Phomopsis liquidambari and different doses of N-fertilizer alter microbial community structure and function in rhizosphere of rice. Sci Rep 6:32270. doi: https://doi.org/10.1038/srep32270
Stassen MJJ, Hsu SH, Pieterse CMJ, Stringlis IA (2021) Coumarin Communication Along the Microbiome-Root-Shoot Axis. Trends Plant Sci 26:169–183. doi: https://doi.org/10.1016/j.tplants.2020.09.008
Sun K, Zhang F-M, Kang N, Gong J-H, Zhang W, Chen Y, Dai C-C (2019) Rice carbohydrate dynamics regulate endophytic colonization of Diaporthe liquidambaris in response to external nitrogen. Fungal Ecol 39:213–224. doi: https://doi.org/10.1016/j.funeco.2019.02.010
Sun K, Zhang W, Yuan J, Song SL, Wu H, Tang MJ, Xu FJ, Xie XG, Dai CC (2020) Nitrogen fertilizer-regulated plant-fungi interaction is related to root invertase-induced hexose generation. FEMS Microbiol Ecol 96. doi: https://doi.org/10.1093/femsec/fiaa139
Tarazona S, Garcia-Alcalde F, Dopazo J, Ferrer A, Conesa A (2011) Differential expression in RNA-seq: a matter of depth. Genome Res 21:2213–2223. doi: https://doi.org/10.1101/gr.124321.111
Thomas J, Kim HR, Rahmatallah Y, Wiggins G, Yang Q, Singh R, Glazko G, Mukherjee A (2019) RNA-seq reveals differentially expressed genes in rice (Oryza sativa) roots during interactions with plant-growth promoting bacteria, Azospirillum brasilense. PLoS ONE 14:e0217309. doi: https://doi.org/10.1371/journal.pone.0217309
Tian B, Pei Y, Huang W, Ding J, Siemann E (2021) Increasing flavonoid concentrations in root exudates enhance associations between arbuscular mycorrhizal fungi and an invasive plant. ISME J 15:1919–1930. doi: https://doi.org/10.1038/s41396-021-00894-1
Waller F, Achatz B, Baltruschat H, Fodor J, Becker K, Fischer M, Heier T, Huckelhoven R, Neumann C, von Wettstein D, Franken P, Kogel KH (2005) The endophytic fungus Piriformospora indica reprograms barley to salt-stress tolerance, disease resistance, and higher yield. Proc Natl Acad Sci U S A 102:13386–13391. doi: https://doi.org/10.1073/pnas.0504423102
Wang W, Mauleon R, Hu Z, Chebotarov D, Tai S, Wu Z, Li M, Zheng T, Fuentes RR, Zhang F, Mansueto L, Copetti D, Sanciangco M, Palis KC, Xu J, Sun C, Fu B, Zhang H, Gao Y, Zhao X, Shen F, Cui X, Yu H, Li Z, Chen M, Detras J, Zhou Y, Zhang X, Zhao Y, Kudrna D, Wang C, Li R, Jia B, Lu J, He X, Dong Z, Xu J, Li Y, Wang M, Shi J, Li J, Zhang D, Lee S, Hu W, Poliakov A, Dubchak I, Ulat VJ, Borja FN, Mendoza JR, Ali J, Li J, Gao Q, Niu Y, Yue Z, Naredo MEB, Talag J, Wang X, Li J, Fang X, Yin Y, Glaszmann JC, Zhang J, Li J, Hamilton RS, Wing RA, Ruan J, Zhang G, Wei C, Alexandrov N, McNally KL, Li Z, Leung H (2018) Genomic variation in 3,010 diverse accessions of Asian cultivated rice. Nature 557:43–49. doi: https://doi.org/10.1038/s41586-018-0063-9
Wang S, Chen A, Xie K, Yang X, Luo Z, Chen J, Zeng D, Ren Y, Yang C, Wang L, Feng H, Lopez-Arredondo DL, Herrera-Estrella LR, Xu G (2020) Functional analysis of the OsNPF4.5 nitrate transporter reveals a conserved mycorrhizal pathway of nitrogen acquisition in plants. Proc Natl Acad Sci U S A 117:16649–16659. doi: https://doi.org/10.1073/pnas.2000926117
Wilson D (1995) Endophyte - the Evolution of a Term, and Clarification of Its Use and Definition. Oikos 73:274–276. Doi https://doi.org/10.2307/3545919
Xie XG, Zhang FM, Yang T, Chen Y, Li XG, Dai CC (2019) Endophytic Fungus Drives Nodulation and N2 Fixation Attributable to Specific Root Exudates. MBio 10:e00728–e00719. doi: https://doi.org/10.1128/mBio.00728-19
Xu XH, Wang C, Li SX, Su ZZ, Zhou HN, Mao LJ, Feng XX, Liu PP, Chen X, Hugh Snyder J, Kubicek CP, Zhang CL, Lin FC (2015) Friend or foe: differential responses of rice to invasion by mutualistic or pathogenic fungi revealed by RNAseq and metabolite profiling. Sci Rep 5:13624. doi: https://doi.org/10.1038/srep13624
Yang B, Ma HY, Wang XM, Jia Y, Hu J, Li X, Dai CC (2014a) Improvement of nitrogen accumulation and metabolism in rice (Oryza sativa L.) by the endophyte Phomopsis liquidambari. Plant Physiol Biochem 82:172–182. doi: https://doi.org/10.1016/j.plaphy.2014.06.002
Yang B, Wang XM, Ma HY, Jia Y, Li X, Dai CC (2014b) Effects of the fungal endophyte Phomopsis liquidambari on nitrogen uptake and metabolism in rice. Plant Growth Regul 73:165–179. doi: https://doi.org/10.1007/s10725-013-9878-4
Yang B, Wang XM, Ma HY, Yang T, Jia Y, Zhou J, Dai CC (2015) Fungal endophyte Phomopsis liquidambari affects nitrogen transformation processes and related microorganisms in the rice rhizosphere. Front Microbiol 6:982. doi: https://doi.org/10.3389/fmicb.2015.00982
Ye J, Fang L, Zheng H, Zhang Y, Chen J, Zhang Z, Wang J, Li S, Li R, Bolund L, Wang J (2006) WEGO: a web tool for plotting GO annotations. Nucleic Acids Res 34:W293–297. doi: https://doi.org/10.1093/nar/gkl031
Yu P, He X, Baer M, Beirinckx S, Tian T, Moya YAT, Zhang X, Deichmann M, Frey FP, Bresgen V, Li C, Razavi BS, Schaaf G, von Wiren N, Su Z, Bucher M, Tsuda K, Goormachtig S, Chen X, Hochholdinger F (2021) Plant flavones enrich rhizosphere Oxalobacteraceae to improve maize performance under nitrogen deprivation. Nat Plants 7:481–499. doi: https://doi.org/10.1038/s41477-021-00897-y
Yuan ZL, Dai CC, Li X, Tian LS, Wang XX (2007) Extensive host range of an endophytic fungus affects the growth and physiological functions in rice (Oryza sativa L.). Symbiosis 43:21–28
Zhang W, Yuan J, Cheng T, Tang MJ, Sun K, Song SL, Xu FJ, Dai CC (2019) Flowering-mediated root-fungus symbiosis loss is related to jasmonate-dependent root soluble sugar deprivation. Plant Cell Environ 42:3208–3226. doi: https://doi.org/10.1111/pce.13636
Zhang W, Li XG, Sun K, Tang MJ, Xu FJ, Zhang M, Dai CC (2020) Mycelial network-mediated rhizobial dispersal enhances legume nodulation. ISME J 14:1015–1029. doi: https://doi.org/10.1038/s41396-020-0587-5
Zhou J, Kang L, Wang HW, Yang T, Dai CC (2014a) Liquid laccase production by Phomopsis liquidambari B3 accelerated phenolic acids degradation in long-term cropping soil of peanut. Acta Agr Scand B-S P 64:683–693. doi: https://doi.org/10.1080/09064710.2014.953987
Zhou J, Yang T, Mei YZ, Kang L, Dai CC (2014b) Laccase production by Phomopsis liquidambari B3 cultured with food waste and wheat straw as the main nitrogen and carbon sources. J Air Waste Manage 64:1154–1163. doi: https://doi.org/10.1080/10962247.2014.930077
Zhou J, Li X, Chen Y, Dai CC (2017) De novo Transcriptome Assembly of Phomopsis liquidambari Provides Insights into Genes Associated with Different Lifestyles in Rice (Oryza sativa L.). Front Plant Sci 8:121. doi: https://doi.org/10.3389/fpls.2017.00121
Zhou J, Li X, Huang PW, Dai CC (2018) Endophytism or saprophytism: Decoding the lifestyle transition of the generalist fungus Phomopsis liquidambari. Microbiol Res 206:99–112. doi: https://doi.org/10.1016/j.micres.2017.10.005
Zipfel C, Oldroyd GE (2017) Plant signalling in symbiosis and immunity. Nature 543:328–336. doi: https://doi.org/10.1038/nature22009
Acknowledgements
We thank Dr. Theresa Catania and Dr. Madeleine Berger (University of York, UK) for proof reading the manuscript.
Funding
This work was supported by the National Natural Science Foundation of China (NSFC 32071638); Program for Jiangsu Excellent Scientific and Technological Innovation Team (17CXTD00014); and a project funded by the Priority Academic Program Development (PAPD) of the Jiangsu Higher Education Institutions of China.
Author information
Authors and Affiliations
Contributions
JZ designed experiments. JZ, PWH and XL performed experiments. JZ analyzed data. JZ and FEV wrote the manuscript. CCD supervised the work.
Corresponding author
Ethics declarations
Conflict of interest
The authors have no conflicts of interest to declare.
Ethics approval
Not applicable.
Consent to participate
Not applicable.
Consent for publication
All authors agreed to the publication and approved the final version.
Additional information
Publisher’s note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Electronic supplementary material
Below is the link to the electronic supplementary material.
Rights and permissions
About this article
Cite this article
Zhou, J., Huang, PW., Li, X. et al. Generalist endophyte Phomopsis liquidambaris colonization of Oryza sativa L. promotes plant growth under nitrogen starvation. Plant Mol Biol 109, 703–715 (2022). https://doi.org/10.1007/s11103-022-01268-7
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s11103-022-01268-7