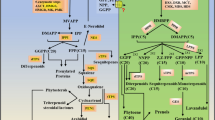
Key message
The core set of biosynthetic genes potentially involved in developmental lignification was identified in the model C4 grass Setaria viridis.
Abstract
Lignin has been recognized as a major recalcitrant factor negatively affecting the processing of plant biomass into bioproducts. However, the efficient manipulation of lignin deposition in order to generate optimized crops for the biorefinery requires a fundamental knowledge of several aspects of lignin metabolism, including regulation, biosynthesis and polymerization. The current availability of an annotated genome for the model grass Setaria viridis allows the genome-wide characterization of genes involved in the metabolic pathway leading to the production of monolignols, the main building blocks of lignin. Here we performed a comprehensive study of monolignol biosynthetic genes as an initial step into the characterization of lignin metabolism in S. viridis. A total of 56 genes encoding bona fide enzymes catalyzing the consecutive ten steps of the monolignol biosynthetic pathway were identified in the S. viridis genome. A combination of comparative phylogenetic studies, high-throughput expression analysis and quantitative RT-PCR analysis was further employed to identify the family members potentially involved in developmental lignification. Accordingly, 14 genes clustered with genes from closely related species with a known function in lignification and showed an expression pattern that correlates with lignin deposition. These genes were considered the “core lignin toolbox” responsible for the constitutive, developmental lignification in S. viridis. These results provide the basis for further understanding lignin deposition in C4 grasses and will ultimately allow the validation of biotechnological strategies to produce crops with enhanced processing properties.










Similar content being viewed by others
References
Abramson M, Shoseyov O, Hirsch S, Shani Z (2013) Genetic modifications of plant cell walls to increase biomass and bioethanol production. Advanced biofuels and bioproducts. Springer, New York, pp 315–338
Agarwal T, Grotewold E, Doseff AI, Gray J (2016) MYB31/MYB42 syntelogs exhibit divergent regulation of phenylpropanoid genes in maize, sorghum and rice. Sci Rep 6:28502
Anderson NA, Tobimatsu Y, Ciesielski PN, Ximenes E, Ralph J, Donohoe BS, Ladisch M, Chapple C (2015) Manipulation of guaiacyl and syringyl monomer biosynthesis in an Arabidopsis Cinnamyl Alcohol Dehydrogenase mutant results in atypical lignin biosynthesis and modified cell wall structure. Plant Cell 27:2195–2209. https://doi.org/10.1105/tpc.15.00373
Barakat A, Choi A, Yassin NBM, Park JS, Sun Z, Carlson JE (2011a) Comparative genomics and evolutionary analyses of the O-methyltransferase gene family in Populus. Gene 479:37–46. https://doi.org/10.1016/j.gene.2011.02.008
Barakat A, Yassin NBM, Park JS, Choi A, Herr J, Carlson JE (2011b) Comparative and phylogenomic analyses of cinnamoyl-CoA reductase and cinnamoyl-CoA-reductase-like gene family in land plants. Plant Sci 181:249–257. https://doi.org/10.1016/j.plantsci.2011.05.012
Barrière Y, Ralph J, Méchin V, Guillaumie S, Grabber JH, Argillier O, Chabbert B, Lapierre C (2004) Genetic and molecular basis of grass cell wall biosynthesis and degradability. II. Lessons from brown-midrib mutants. C R Biol 327:847–860. https://doi.org/10.1016/j.crvi.2004.05.010
Barros J, Serk H, Granlund I, Pesquet E (2015) The cell biology of lignification in higher plants. Ann Bot 115:1053–1074. https://doi.org/10.1093/aob/mcv046
Barros J, Serrani-Yarce JC, Chen F, Baxter D, Venables BJ, Dixon RA (2016) Role of bifunctional ammonia-lyase in grass cell wall biosynthesis. Nat Plants 2:16050. https://doi.org/10.1038/nplants.2016.50
Barros J, Escamilla-Trevino L, Song L, Rao X, Serrani-Yarce JC, Palacios MD, Engle N, Choudhury FK, Tschaplinski TJ, Venables BJ, Mittler R, Dixon RA (2019) 4-Coumarate 3-hydroxylase in the lignin biosynthesis pathway is a cytosolic ascorbate peroxidase. Nat Commun 10:1994. https://doi.org/10.1038/s41467-019-10082-7
Baucher M, Chabbert B, Pilate G, Doorsselaere JV, Tollier MT, Petit-Conil M, Cornu D, Monties B, Montagu MV, Inze D, Jouanin L, Boerjan W (1996) Red xylem and higher lignin extractability by down-regulating a Cinnamyl Alcohol Dehydrogenase in poplar. Plant Physiol 112:1479–1490. https://doi.org/10.1104/pp.112.4.1479
Bouvier d’Yvoire M, Bouchabke-Coussa O, Voorend W, Antelme S, Cézard L, Legée F, Lebris P, Legay S, Whitehead C, McQueen-Mason SJ, Gomez LD, Jouanin L, Lapierre C, Sibout R (2013) Disrupting the cinnamyl alcohol dehydrogenase 1 gene (BdCAD1) leads to altered lignification and improved saccharification in Brachypodium distachyon. Plant J 73:496–508. https://doi.org/10.1111/tpj.12053
Cannon SB, Mitra A, Baumgarten A, Young ND, May G (2004) The roles of segmental and tandem gene duplication in the evolution of large gene families in Arabidopsis thaliana. BMC Plant Biol 4:10. https://doi.org/10.1186/1471-2229-4-10
Carocha V, Soler M, Hefer C, Cassan-Wang H, Fevereiro P, Myburg AA, Paiva JAP, Grima-Pettenati J (2015) Genome-wide analysis of the lignin toolbox of Eucalyptus grandis. New Phytol 206:1297–1313. https://doi.org/10.1111/nph.13313
Cass CL, Peraldi A, Dowd PF, Mottiar Y, Santoro N, Karlen SD, Bukhman YV, Foster CE, Thrower N, Bruno LC, Moskvin OV, Johnson ET, Willhoit ME, Phutane M, Ralph J, Mansfield SD, Nicholson P, Sedbrook JC (2015) Effects of PHENYLALANINE AMMONIA LYASE (PAL) knockdown on cell wall composition, biomass digestibility, and biotic and abiotic stress responses in Brachypodium. J Exp Bot 66:4317–4335. https://doi.org/10.1093/jxb/erv269
Cesarino I (2019) Structural features and regulation of lignin deposited upon biotic and abiotic stresses. Curr Opin Biotechnol 56:209–214. https://doi.org/10.1016/j.copbio.2018.12.012
Cesarino I, Araujo P, Domingues Junior AP, Mazzafera P (2012) An overview of lignin metabolism and its effect on biomass recalcitrance. Braz J Bot 35:303–311
Cesarino I, Simões MS, dos Brito M, Fanelli A, da Franca Silva T, Romanel E (2016) Building the wall: recent advances in understanding lignin metabolism in grasses. Acta Physiol Plant 38:269. https://doi.org/10.1007/s11738-016-2293-5
de Oliveira DM, Finger-Teixeira A, Mota TR, Salvador VH, Moreira-Vilar FC, Molinari HBC, Mitchell RAC, Marchiosi R, Ferrarese-Filho O, dos Santos WD (2015) Ferulic acid: a key component in grass lignocellulose recalcitrance to hydrolysis. Plant Biotechnol J 13:1224–1232. https://doi.org/10.1111/pbi.12292
de Souza WR, Martins PK, Freeman J, Pellny TK, Michaelson LV, Sampaio BL, Vinecky F, Ribeiro AP, da Cunha BADB, Kobayashi AK, de Oliveira PA, Campanha RB, Pacheco TF, Martarello DCI, Marchiosi R, Ferrarese-Filho O, dos Santos WD, Tramontina R, Squina FM, Centeno DC, Gaspar M, Braga MR, Tiné MAS, Ralph J, Mitchell RAC, Molinari HBC (2018) Suppression of a single BAHD gene in Setaria viridis causes large, stable decreases in cell wall feruloylation and increases biomass digestibility. New Phytol 218:81–93. https://doi.org/10.1111/nph.14970
Del Río JC, Rencoret J, Prinsen P, Martinez AT, Ralph J, Gutierrez A (2012) Structural characterization of wheat straw lignin as revealed by analytical pyrolysis, 2D-NMR, and reductive cleavage methods. J Agric Food Chem 60:5922–5935. https://doi.org/10.1021/jf301002n
Do C-T, Pollet B, Thévenin J, Sibout R, Denoue D, Barrière Y, Lapierre C, Jouanin L (2007) Both caffeoyl Coenzyme A 3-O-methyltransferase 1 and caffeic acid O-methyltransferase 1 are involved in redundant functions for lignin, flavonoids and sinapoyl malate biosynthesis in Arabidopsis. Planta 226:1117–1129. https://doi.org/10.1007/s00425-007-0558-3
Ehlting J, Büttner D, Wang Q, Douglas CJ, Somssich IE, Kombrink E (1999) Three 4-coumarate:coenzyme A ligases in Arabidopsis thaliana represent two evolutionarily divergent classes in angiosperms. Plant J 19:9–20. https://doi.org/10.1046/j.1365-313X.1999.00491.x
Eloy N, Voorend W, Lan W, Saleme MD, Cesarino I, Vanholme R, Smith RA, Goeminne G, Pallidis A, Morreel K, Nicomedes J, Ralph J, Boerjan WA (2016) Silencing CHALCONE SYNTHASE impedes the incorporation of tricin in lignin and increases lignin content. Plant Physiol. https://doi.org/10.1104/pp.16.01108
Escamilla-Trevino LL, Shen H, Uppalapati SR, Ray T, Tang Y, Hernandez T, Yin Y, Xu Y, Dixon RA (2010) Switchgrass (Panicum virgatum) possesses a divergent family of cinnamoyl CoA reductases with distinct biochemical properties. New Phytol 185:143–155. https://doi.org/10.1111/j.1469-8137.2009.03018.x
Eudes A, Dutta T, Deng K, Jacquet N, Sinha A, Benites VT, Baidoo EEK, Richel A, Sattler SE, Northen TR, Singh S, Simmons BA, Loqué D (2017) SbCOMT (Bmr12) is involved in the biosynthesis of tricin-lignin in sorghum. PLoS ONE 12:e0178160. https://doi.org/10.1371/journal.pone.0178160
Ferreira SS, Hotta CT, de Poelking VG, Leite DCC, Buckeridge MS, Loureiro ME, Barbosa MHP, Carneiro MS, Souza GM (2016) Co-expression network analysis reveals transcription factors associated to cell wall biosynthesis in sugarcane. Plant Mol Biol 91(1–2):15–35. https://doi.org/10.1007/s11103-016-0434-2
Fornalé S, Capellades M, Encina A, Wang K, Irar S, Lapierre C, Ruel K, Joseleau J-P, Berenguer J, Puigdomènech P, Rigau J, Caparrós-Ruiz D (2012) Altered lignin biosynthesis improves cellulosic bioethanol production in transgenic maize plants down-regulated for Cinnamyl Alcohol Dehydrogenase. Mol Plant 5:817–830. https://doi.org/10.1093/mp/ssr097
Fornalé S, Rencoret J, Garcia-Calvo L, Capellades M, Encina A, Santiago R, Rigau J, Gutiérrez A, del Río J-C, Caparros-Ruiz D (2015) Cell wall modifications triggered by the down-regulation of Coumarate 3-hydroxylase-1 in maize. Plant Sci 236:272–282. https://doi.org/10.1016/j.plantsci.2015.04.007
Fornalé S, Rencoret J, García-Calvo L, Encina A, Rigau J, Gutiérrez A, del Río JC, Caparros-Ruiz D (2017) Changes in cell wall polymers and degradability in maize mutants lacking 3′- and 5′-O-methyltransferases involved in lignin biosynthesis. Plant Cell Physiol 58:240–255. https://doi.org/10.1093/pcp/pcw198
Gou M, Ran X, Martin DW, Liu C-J (2018) The scaffold proteins of lignin biosynthetic cytochrome P450 enzymes. Nat Plants 4:299–310. https://doi.org/10.1038/s41477-018-0142-9
Goujon T, Sibout R, Pollet B, Maba B, Nussaume L, Bechtold N, Lu F, Ralph J, Mila I, Barrière Y, Lapierre C, Jouanin L (2003) A new Arabidopsis thaliana mutant deficient in the expression of O-methyltransferase impacts lignins and sinapoyl esters. Plant Mol Biol 51:973–989. https://doi.org/10.1023/A:1023022825098
Grabber JH, Ralph J, Lapierre C, Barriere Y (2004) Genetic and molecular basis of grass cell-wall degradability. I. Lignin-cell wall matrix interactions. C R Biol 327:455–465. https://doi.org/10.1016/j.crvi.2004.02.009
Gui J, Shen J, Li L (2011) Functional characterization of evolutionarily divergent 4-Coumarate: Coenzyme A Ligases in rice. Plant Physiol 157:574–586. https://doi.org/10.1104/pp.111.178301
Guindon S, Dufayard J-F, Lefort V, Anisimova M, Hordijk W, Gascuel O (2010) New algorithms and methods to estimate maximum-likelihood phylogenies: assessing the performance of PhyML 3.0. Syst Biol 59:307–321. https://doi.org/10.1093/sysbio/syq010
Ha CM, Escamilla-Trevino L, Yarce JCS, Kim H, Ralph J, Chen F, Dixon RA (2016) An essential role of caffeoyl shikimate esterase in monolignol biosynthesis in Medicago truncatula. Plant J 86:363–375. https://doi.org/10.1111/tpj.13177
Handakumbura PP, Hazen SP (2012) Transcriptional regulation of grass secondary cell wall biosynthesis: playing catch-up with Arabidopsis thaliana. Plant Physiol 3:74. https://doi.org/10.3389/fpls.2012.00074
Hellemans J, Mortier G, De Paepe A, Speleman F, Vandesompele J (2007) qBase relative quantification framework and software for management and automated analysis of real-time quantitative PCR data. Genome Biol 8:R19. https://doi.org/10.1186/gb-2007-8-2-r19
Hirano K, Aya K, Kondo M, Okuno A, Morinaka Y, Matsuoka M (2012) OsCAD2 is the major CAD gene responsible for monolignol biosynthesis in rice culm. Plant Cell Rep 31:91–101. https://doi.org/10.1007/s00299-011-1142-7
Hoang DT, Chernomor O, von Haeseler A, Minh BQ, Vinh LS (2018) UFBoot2: improving the ultrafast bootstrap approximation. Mol Biol Evol 35:518–522. https://doi.org/10.1093/molbev/msx281
Hoffmann L, Besseau S, Geoffroy P, Ritzenthaler C, Meyer D, Lapierre C, Pollet B, Legrand M (2004) Silencing of Hydroxycinnamoyl-Coenzyme A Shikimate/Quinate Hydroxycinnamoyltransferase affects phenylpropanoid biosynthesis. Plant Cell Online 16:1446–1465. https://doi.org/10.1105/tpc.020297
Ho-Yue-Kuang S, Alvarado C, Antelme S, Bouchet B, Cézard L, Le Bris P, Legée F, Maia-Grondard A, Yoshinaga A, Saulnier L, Guillon F, Sibout R, Lapierre C, Chateigner-Boutin A-L (2016) Mutation in Brachypodium caffeic acid O-methyltransferase 6 alters stem and grain lignins and improves straw saccharification without deteriorating grain quality. J Exp Bot 67:227–237. https://doi.org/10.1093/jxb/erv446
Humphreys JM, Hemm MR, Chapple C (1999) New routes for lignin biosynthesis defined by biochemical characterization of recombinant ferulate 5-hydroxylase, a multifunctional cytochrome P450-dependent monooxygenase. Proc Natl Acad Sci USA 96:10045–10050. https://doi.org/10.1073/pnas.96.18.10045
Jun S-Y, Sattler SA, Cortez GS, Vermerris W, Sattler SE, Kang C (2018) Biochemical and structural analysis of substrate specificity of a Phenylalanine Ammonia-Lyase. Plant Physiol 176:1452–1468. https://doi.org/10.1104/pp.17.01608
Katoh K, Kuma KI, Toh H, Miyata K (2005) MAFFT version 5: improvement in accuracy of multiple sequence alignment. Nucleic Acids Res 33:511–518. https://doi.org/10.1093/nar/gki198
Kim RJ, Kim HJ, Shim D, Suh MC (2016) Molecular and biochemical characterizations of the monoacylglycerol lipase gene family of Arabidopsis thaliana. Plant J 85:758–771. https://doi.org/10.1111/tpj.13146
Koshiba T, Hirose N, Mukai M, Yamamura M, Hattori T, Suzuki S, Sakamoto M, Umezawa T (2013) Characterization of 5-hydroxyconiferaldehyde O-methyltransferase in Oryza sativa. Plant Biotechnol 30:157–167. https://doi.org/10.5511/plantbiotechnology.13.0219a
Lam PY, Zhu F-Y, Chan WL, Liu H, Lo C (2014) Cytochrome P450 93G1 is a flavone synthase II that channels flavanones to the biosynthesis of tricin O-linked conjugates in rice. Plant Physiol 165:1315–1327. https://doi.org/10.1104/pp.114.239723
Lam PY, Liu H, Lo C (2015) Completion of tricin biosynthesis pathway in rice: cytochrome P450 75B4 is a unique chrysoeriol 5′-hydroxylase. Plant Physiol 168:1527–1536. https://doi.org/10.1104/pp.15.00566
Lan W, Lu F, Regner M, Zhu Y, Rencoret J, Ralph SA, Zakai UI, Morreel K, Boerjan W, Ralph J (2015) Tricin, a flavonoid monomer in monocot lignification. Plant Physiol 167:1284–1295. https://doi.org/10.1104/pp.114.253757
Lan W, Morreel K, Lu F, Rencoret J, Del Río JC, Voorend W, Vermerris W, Boerjan WA, Ralph J (2016a) Maize tricin-oligolignol metabolites and their implications for monocot lignification. Plant Physiol. https://doi.org/10.1104/pp.16.02012
Lan W, Rencoret J, Lu F, Karlen SD, Smith BG, Harris PJ, del Río JC, Ralph J (2016b) Tricin-lignins: occurrence and quantitation of tricin in relation to phylogeny. Plant J 88:1046–1057. https://doi.org/10.1111/tpj.13315
Li P, Brutnell TP (2011) Setaria viridis and Setaria italica, model genetic systems for the Panicoid grasses. J Exp Bot 62:3031–3037. https://doi.org/10.1093/jxb/err096
Li L, Popko JL, Umezawa T, Chiang VL (2000) 5-hydroxyconiferyl aldehyde modulates enzymatic methylation for syringyl monolignol formation, a new view of monolignol biosynthesis in angiosperms. J Biol Chem 275:6537–6545
Li X, Yang Y, Yao J, Chen G, Li X, Zhang Q, Wu C (2009) FLEXIBLE CULM 1 encoding a cinnamyl-alcohol dehydrogenase controls culm mechanical strength in rice. Plant Mol Biol 69:685–697. https://doi.org/10.1007/s11103-008-9448-8
Li Y, Kim JI, Pysh L, Chapple C (2015) Four isoforms of Arabidopsis 4-coumarate: CoA ligase have overlapping yet distinct roles in phenylpropanoid metabolism. Plant Physiol 169:2409–2421. https://doi.org/10.1104/pp.15.00838
Liu C-J (2012) Deciphering the enigma of lignification: precursor transport, oxidation, and the topochemistry of lignin assembly. Mol Plant 5:304–317. https://doi.org/10.1093/mp/ssr121
Marita JM, Hatfield RD, Rancour DM, Frost KE (2014) Identification and suppression of the p-coumaroyl CoA:hydroxycinnamyl alcohol transferase in Zea mays L. Plant J 78:850–864. https://doi.org/10.1111/tpj.12510
Marriott PE, Gómez LD, McQueen-Mason SJ (2016) Unlocking the potential of lignocellulosic biomass through plant science. New Phytol 209:1366–1381. https://doi.org/10.1111/nph.13684
Martin AP, Palmer WM, Brown C, Abel C, Lunn JE, Furbank RT, Grof CPL (2016) A developing Setaria viridis internode: an experimental system for the study of biomass generation in a C4 model species. Biotechnol Biofuels 9:45. https://doi.org/10.1186/s13068-016-0457-6
Martins PK, Nakayama TJ, Ribeiro AP, Cunha BADB, Nepomuceno AL, Harmon FG, Kobayashi AK, Molinari HBC (2015a) Setaria viridis floral-dip: a simple and rapid Agrobacterium-mediated transformation method. Biotechnol Rep 6:61–63. https://doi.org/10.1016/j.btre.2015.02.006
Martins PK, Ribeiro AP, Cunha BADB, Kobayashi AK, Molinari HBC (2015b) A simple and highly efficient Agrobacterium-mediated transformation protocol for Setaria viridis. Biotechnol Rep 6:41–44. https://doi.org/10.1016/j.btre.2015.02.002
Martins PK, Mafra V, de Souza WR, Ribeiro AP, Vinecky F, Basso MF, da Cunha BADB, Kobayashi AK, Molinari HBC (2016) Selection of reliable reference genes for RT-qPCR analysis during developmental stages and abiotic stress in Setaria viridis. Sci Rep 6:28348. https://doi.org/10.1038/srep28348
Meyer K, Shirley AM, Cusumano JC, Bell-Lelong DA, Chapple C (1998) Lignin monomer composition is determined by the expression of a cytochrome P450-dependent monooxygenase in Arabidopsis. Proc Natl Acad Sci USA 95:6619–6623
Monte-Bello CC, Araujo EF, Martins MCM, Mafra V, da Silva VCH, Celente V, Caldana C (2018) A flexible low cost hydroponic system for assessing plant responses to small molecules in sterile conditions. JoVE J Vis Exp 138:e57800. https://doi.org/10.3791/57800
Mottiar Y, Vanholme R, Boerjan W, Ralph J, Mansfield SD (2016) Designer lignins: harnessing the plasticity of lignification. Curr Opin Biotechnol 37:190–200. https://doi.org/10.1016/j.copbio.2015.10.009
Muthamilarasan M, Khan Y, Jaishankar J, Shweta S, Lata C, Prasad M (2015) Integrative analysis and expression profiling of secondary cell wall genes in C4 biofuel model Setaria italica reveals targets for lignocellulose bioengineering. Front Plant Sci 6:965. https://doi.org/10.3389/fpls.2015.00965
Nelson RS, Stewart CN, Gou J, Holladay S, Gallego-Giraldo L, Flanagan A, Mann DGJ, Hisano H, Wuddineh WA, Poovaiah CR, Srivastava A, Biswal AK, Shen H, Escamilla-Treviño LL, Yang J, Hardin CF, Nandakumar R, Fu C, Zhang J, Xiao X, Percifield R, Chen F, Bennetzen JL, Udvardi M, Mazarei M, Dixon RA, Wang Z-Y, Tang Y, Mohnen D, Davison BH (2017) Development and use of a switchgrass (Panicum virgatum L.) transformation pipeline by the BioEnergy Science Center to evaluate plants for reduced cell wall recalcitrance. Biotechnol Biofuels 10:309. https://doi.org/10.1186/s13068-017-0991-x
Park HL, Bhoo SH, Kwon M, Lee S-W, Cho M-H (2017) Biochemical and expression analyses of the rice Cinnamoyl-CoA Reductase gene family. Front Plant Sci 8:2099. https://doi.org/10.3389/fpls.2017.02099
Parvathi K, Chen F, Guo D, Blount JW, Dixon RA (2001) Substrate preferences of O-methyltransferases in alfalfa suggest new pathways for 3-O-methylation of monolignols. Plant J 25:193–202. https://doi.org/10.1111/j.1365-313X.2001.00956.x
Petrik DL, Karlen SD, Cass CL, Padmakshan D, Lu F, Liu S, Le Bris P, Antelme S, Santoro N, Wilkerson CG, Sibout R, Lapierre C, Ralph J, Sedbrook JC (2014) p-Coumaroyl-CoA:monolignol transferase (PMT) acts specifically in the lignin biosynthetic pathway in Brachypodium distachyon. Plant J 77:713–726. https://doi.org/10.1111/tpj.12420
Piquemal J, Chamayou S, Nadaud I, Beckert M, Barrière Y, Mila I, Lapierre C, Rigau J, Puigdomenech P, Jauneau A, Digonnet C, Boudet A-M, Goffner D, Pichon M (2002) Down-regulation of Caffeic Acid O-Methyltransferase in maize revisited using a transgenic approach. Plant Physiol 130:1675–1685. https://doi.org/10.1104/pp.012237
Ponniah SK, Shang Z, Akbudak MA, Srivastava V, Manoharan M (2017) Down-regulation of hydroxycinnamoyl CoA: shikimate hydroxycinnamoyl transferase, cinnamoyl CoA reductase, and cinnamyl alcohol dehydrogenase leads to lignin reduction in rice (Oryza sativa L. ssp. japonica cv. Nipponbare). Plant Biotechnol Rep 11:17–27. https://doi.org/10.1007/s11816-017-0426-y
Raes J, Rohde A, Christensen JH, Van de Peer Y, Boerjan W (2003) Genome-wide characterization of the lignification toolbox in Arabidopsis. Plant Physiol 133:1051–1071. https://doi.org/10.1104/pp.103.026484
Rakoczy M, Femiak I, Alejska M, Figlerowicz M, Podkowinski J (2018) Sorghum CCoAOMT and CCoAOMT-like gene evolution, structure, expression and the role of conserved amino acids in protein activity. Mol Genet Genomics 293:1077–1089. https://doi.org/10.1007/s00438-018-1441-6
Ralph J, Lundquist K, Brunow G, Lu F, Kim H, Schatz PF, Marita JM, Hatfield RD, Ralph SA, Christensen JH, Boerjan W (2004) Lignins: natural polymers from oxidative coupling of 4-hydroxyphenyl-propanoids. Phytochem Rev 3:29–60. https://doi.org/10.1023/B:PHYT.0000047809.65444.a4
Rao X, Dixon RA (2018) Current models for transcriptional regulation of secondary cell wall biosynthesis in grasses. Front Plant Sci 9:399. https://doi.org/10.3389/fpls.2018.00399
Renault H, De Marothy M, Jonasson G, Lara P, Nelson DR, Nilsson I, André F, von Heijne G, Werck-Reichhart D (2017) Gene duplication leads to altered membrane topology of a cytochrome P450 enzyme in seed plants. Mol Biol Evol 34:2041–2056. https://doi.org/10.1093/molbev/msx160
Rinaldi R, Jastrzebski R, Clough MT, Ralph J, Kennema M, Bruijnincx PCA, Weckhuysen BM (2016) Paving the way for lignin valorisation: recent advances in bioengineering, biorefining and catalysis. Angew Chem Int Ed 55:8164–8215. https://doi.org/10.1002/anie.201510351
Rosler J, Krekel F, Amrhein N, Schmid J (1997) Maize phenylalanine ammonia-lyase has tyrosine ammonia-lyase activity. Plant Physiol 113:175–179. https://doi.org/10.1104/pp.113.1.175
Saballos A, Vermerris W, Rivera L, Ejeta G (2008) Allelic association, chemical characterization and saccharification properties of brown midrib mutants of sorghum (Sorghum bicolor (L.) Moench). BioEnergy Res 1:193–204. https://doi.org/10.1007/s12155-008-9025-7
Saballos A, Ejeta G, Sanchez E, Kang C, Vermerris W (2009) A genomewide analysis of the Cinnamyl Alcohol Dehydrogenase family in sorghum [Sorghum bicolor (L.) Moench] identifies SbCAD2 as the Brown midrib6 gene. Genetics 181:783–795. https://doi.org/10.1534/genetics.108.098996
Saballos A, Sattler SE, Sanchez E, Foster TP, Xin Z, Kang C, Pedersen JF, Vermerris W (2012) Brown midrib2 (Bmr2) encodes the major 4-coumarate:coenzyme A ligase involved in lignin biosynthesis in sorghum (Sorghum bicolor (L.) Moench). Plant J 70:818–830. https://doi.org/10.1111/j.1365-313X.2012.04933.x
Saha P, Blumwald E (2016) Spike-dip transformation of Setaria viridis. Plant J 86:89–101. https://doi.org/10.1111/tpj.13148
Saleme M, Cesarino I, Vargas L, Kim H, Vanholme R, Goeminne G, Acker RV, de Fonseca FC, Pallidis A, Voorend W, Junior JN, Padmakshan D, Doorsselaere JV, Ralph J, Boerjan W (2017) Silencing CAFFEOYL SHIKIMATE ESTERASE affects lignification and improves saccharification in poplar. Plant Physiol 175:1040–1057. https://doi.org/10.1104/pp.17.00920
Sattler SA, Walker AM, Vermerris W, Sattler SE, Kang C (2017) Structural and biochemical characterization of Cinnamoyl-CoA Reductases. Plant Physiol 173:1031–1044. https://doi.org/10.1104/pp.16.01671
Shih PM, Liang Y, Loqué D (2016) Biotechnology and synthetic biology approaches for metabolic engineering of bioenergy crops. Plant J 87:103–117. https://doi.org/10.1111/tpj.13176
Sibout R, Eudes A, Mouille G, Pollet B, Lapierre C, Jouanin L, Séguin A (2005) CINNAMYL ALCOHOL DEHYDROGENASE-C and -D are the primary genes involved in lignin biosynthesis in the floral stem of Arabidopsis. Plant Cell 17:2059–2076. https://doi.org/10.1105/tpc.105.030767
Sibout R, Proost S, Hansen BO, Vaid N, Giorgi FM, Ho-Yue-Kuang S, Legée F, Cézart L, Bouchabké-Coussa O, Soulhat C, Provart N, Pasha A, Bris PL, Roujol D, Hofte H, Jamet E, Lapierre C, Persson S, Mutwil M (2017) Expression atlas and comparative coexpression network analyses reveal important genes involved in the formation of lignified cell wall in Brachypodium distachyon. New Phytol 215:1009–1025. https://doi.org/10.1111/nph.14635
Smith RA, Cass CL, Mazaheri M, Sekhon RS, Heckwolf M, Kaeppler H, de Leon N, Mansfield SD, Kaeppler SM, Sedbrook JC, Karlen SD, Ralph J (2017) Suppression of CINNAMOYL-CoA REDUCTASE increases the level of monolignol ferulates incorporated into maize lignins. Biotechnol Biofuels 10:109. https://doi.org/10.1186/s13068-017-0793-1
Takeda Y, Koshiba T, Tobimatsu Y, Suzuki S, Murakami S, Yamamura M, Rahman MdM, Takano T, Hattori T, Sakamoto M, Umezawa T (2017) Regulation of CONIFERALDEHYDE 5-HYDROXYLASE expression to modulate cell wall lignin structure in rice. Planta 246:337–349. https://doi.org/10.1007/s00425-017-2692-x
Takeda Y, Suzuki S, Tobimatsu Y, Osakabe K, Osakabe Y, Ragamustari SK, Sakamoto M, Umezawa T (2019) Lignin characterization of rice CONIFERALDEHYDE 5-HYDROXYLASE loss-of-function mutants generated with the CRISPR/Cas9 system. Plant J 97:543–554. https://doi.org/10.1111/tpj.14141
Trabucco GM, Matos DA, Lee SJ, Saathoff AJ, Priest HD, Mockler TC, Sarath G, Hazen SP (2013) Functional characterization of cinnamyl alcohol dehydrogenase and caffeic acid O-methyltransferase in Brachypodium distachyon. BMC Biotechnol 13:61. https://doi.org/10.1186/1472-6750-13-61
Trifinopoulos J, Nguyen L-T, von Haeseler A, Minh BQ (2016) W-IQ-TREE: a fast online phylogenetic tool for maximum likelihood analysis. Nucleic Acids Res 44:W232–W235. https://doi.org/10.1093/nar/gkw256
Umezawa T (2018) Lignin modification in planta for valorization. Phytochem Rev 17:1305–1327. https://doi.org/10.1007/s11101-017-9545-x
van der Weijde T, Kamei CLA, Torres AF, Vermerris W, Dolstra O, Visser RGF, Trindade LM (2013) The potential of C4 grasses for cellulosic biofuel production. Front Plant Sci 4:107. https://doi.org/10.3389/fpls.2013.00107
Vandesompele J, De Preter K, Pattyn F, Poppe B, Van Roy N, De Paepe A, Speleman F (2002) Accurate normalization of real-time quantitative RT-PCR data by geometric averaging of multiple internal control genes. Genome Biol 3(7):research0034-1. https://doi.org/10.1186/gb-2002-3-7-research0034
Vanholme R, Ralph J, Akiyama T, Lu F, Pazo JR, Kim H, Christensen JH, Van Reusel B, Storme V, De Rycke R, Rohde A, Morreel K, Boerjan W (2010) Engineering traditional monolignols out of lignin by concomitant up-regulation of F5H1 and down-regulation of COMT in Arabidopsis. Plant J 64:885–897. https://doi.org/10.1111/j.1365-313X.2010.04353.x
Vanholme R, Morreel K, Darrah C, Oyarce P, Grabber JH, Ralph J, Boerjan W (2012) Metabolic engineering of novel lignin in biomass crops. New Phytol 196:978–1000. https://doi.org/10.1111/j.1469-8137.2012.04337.x
Vanholme B, Desmet T, Ronsse F, Rabaey K, Van Breusegem F, De Mey M, Soetaert W, Boerjan W (2013a) Towards a carbon-negative sustainable bio-based economy. Front Plant Sci 4:174. https://doi.org/10.3389/fpls.2013.00174
Vanholme R, Cesarino I, Rataj K, Xiao Y, Sundin L, Goeminne G, Kim H, Cross J, Morreel K, Araujo P, Welsh L, Haustraete J, McClellan C, Vanholme B, Ralph J, Simpson GG, Halpin C, Boerjan W (2013b) Caffeoyl Shikimate Esterase (CSE) is an enzyme in the lignin biosynthetic pathway in Arabidopsis. Science 341:1103–1106. https://doi.org/10.1126/science.1241602
Vargas L, Cesarino I, Vanholme R, Voorend W, Saleme MD, Morreel K, Boerjan W (2016) Improving total saccharification yield of Arabidopsis plants by vessel-specific complementation of caffeoyl shikimate esterase (cse) mutants. Biotechnol Biofuels 9:139. https://doi.org/10.1186/s13068-016-0551-9
Vogel J (2008) Unique aspects of the grass cell wall. Curr Opin Plant Biol 11:301–307. https://doi.org/10.1016/j.pbi.2008.03.002
Voxeur A, Wang Y, Sibout R (2015) Lignification: different mechanisms for a versatile polymer. Curr Opin Plant Biol 23:83–90. https://doi.org/10.1016/j.pbi.2014.11.006
Walker AM, Hayes RP, Youn B, Vermerris W, Sattler SE, Kang C (2013) Elucidation of the structure and reaction mechanism of sorghum hydroxycinnamoyltransferase and its structural relationship to other coenzyme A-dependent transferases and synthases. Plant Physiol 162:640–651. https://doi.org/10.1104/pp.113.217836
Walker AM, Sattler SA, Regner M, Jones JP, Ralph J, Vermerris W, Sattler SE, Kang C (2016) The structure and catalytic mechanism of Sorghum bicolor caffeoyl-CoA O-methyltransferase. Plant Physiol 172:78–92. https://doi.org/10.1104/pp.16.00845
Wang Y, Wang X, Paterson AH (2012) Genome and gene duplications and gene expression divergence: a view from plants. Ann N Y Acad Sci 1256:1–14. https://doi.org/10.1111/j.1749-6632.2011.06384.x
Watts KT, Mijts BN, Lee PC, Manning AJ, Schmidt-Dannert C (2006) Discovery of a substrate selectivity switch in tyrosine ammonia-lyase, a member of the aromatic amino acid lyase family. Chem Biol 13:1317–1326. https://doi.org/10.1016/j.chembiol.2006.10.008
Weng J-K, Li Y, Mo H, Chapple C (2012) Assembly of an evolutionarily new pathway for α-pyrone biosynthesis in Arabidopsis. Science 337:960–964. https://doi.org/10.1126/science.1221614
Wils CR, Brandt W, Manke K, Vogt T (2013) A single amino acid determines position specificity of an Arabidopsis thaliana CCoAOMT-like O-methyltransferase. FEBS Lett 587:683–689. https://doi.org/10.1016/j.febslet.2013.01.040
Wolfe D, Dudek S, Ritchie MD, Pendergrass SA (2013) Visualizing genomic information across chromosomes with PhenoGram. BioData Min 6:18. https://doi.org/10.1186/1756-0381-6-18
Xiong W, Wu Z, Liu Y, Li Y, Su K, Bai Z, Guo S, Hu Z, Zhang Z, Bao Y, Sun J, Yang G, Fu C (2019) Mutation of 4-coumarate: coenzyme A ligase 1 gene affects lignin biosynthesis and increases the cell wall digestibility in maize brown midrib5 mutants. Biotechnol Biofuels 12:82. https://doi.org/10.1186/s13068-019-1421-z
Xu B, Escamilla-Treviño LL, Sathitsuksanoh N, Shen Z, Shen H, Zhang Y-HP, Dixon RA, Zhao B (2011) Silencing of 4-coumarate:coenzyme A ligase in switchgrass leads to reduced lignin content and improved fermentable sugar yields for biofuel production. New Phytol 192:611–625. https://doi.org/10.1111/j.1469-8137.2011.03830.x
Yang Y, Tilman D, Furey G, Lehman C (2019) Soil carbon sequestration accelerated by restoration of grassland biodiversity. Nat Commun 10:718. https://doi.org/10.1038/s41467-019-08636-w
Zeng Y, Zhao S, Yang S, Ding S-Y (2014) Lignin plays a negative role in the biochemical process for producing lignocellulosic biofuels. Curr Opin Biotechnol 27:38–45. https://doi.org/10.1016/j.copbio.2013.09.008
Zhao Q (2016) Lignification: flexibility, biosynthesis and regulation. Trends Plant Sci 21:713–721. https://doi.org/10.1016/j.tplants.2016.04.006
Zhao X, Zhang L, Liu D (2012) Biomass recalcitrance. Part II: fundamentals of different pre-treatments to increase the enzymatic digestibility of lignocellulose. Biofuels Bioprod Biorefining-Biofpr 6:561–579. https://doi.org/10.1002/bbb.1350
Zhao Q, Tobimatsu Y, Zhou R, Pattathil S, Gallego-Giraldo L, Fu C, Jackson LA, Hahn MG, Kim H, Chen F, Ralph J, Dixon RA (2013) Loss of function of cinnamyl alcohol dehydrogenase 1 leads to unconventional lignin and a temperature-sensitive growth defect in Medicago truncatula. Proc Natl Acad Sci USA 110:13660–13665. https://doi.org/10.1073/pnas.1312234110
Zhong R, Ye Z-H (2012) MYB46 and MYB83 bind to the SMRE sites and directly activate a suite of transcription factors and secondary wall biosynthetic genes. Plant Cell Physiol 53:368–380. https://doi.org/10.1093/pcp/pcr185
Zhong R, Ye Z-H (2014) Complexity of the transcriptional network controlling secondary wall biosynthesis. Plant Sci 229:193–207. https://doi.org/10.1016/j.plantsci.2014.09.009
Zhong R, Lee C, Ye Z-H (2010) Global analysis of direct targets of secondary wall NAC master switches in Arabidopsis. Mol Plant 3:1087–1103. https://doi.org/10.1093/mp/ssq062
Acknowledgements
This work was supported by Fundação de Amparo à Pesquisa do Estado de São Paulo (FAPESP) via the BIOEN Young Investigators Awards research grant (Processo FAPESP no. 2015/02527-1). This study was financed in part by the Coordenação de Aperfeiçoamento de Pessoal de Nível Superior - Brasil (CAPES) - Finance Code 001. IC thanks FAPESP for the SPRINT Project grant number 2016/50189-0. IC is indebted to Conselho Nacional de Desenvolvimento Científico e Tecnológico (CNPq) for the research fellowship 302927/2018-2. SSF was funded by FAPESP for a postdoctoral fellowship (Processo FAPESP no. 2016/06917-1). MSS was funded by CAPES for a predoctoral fellowship. RMAS was funded by CNPq for a Master’s fellowship. GGC was funded by CNPq-PIBIC for a scientific initiation fellowship.
Author information
Authors and Affiliations
Contributions
SSF conducted most of the experiments, analyzed data and complemented the writing. MSS, GGC, LGAL and RMAS performed experiments and analyzed data. IC conceived and designed the experiments and wrote the article. All authors read and approved the final manuscript.
Corresponding author
Ethics declarations
Conflict of interest
The authors declare that they have no conflict of interest.
Additional information
Publisher's Note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Electronic supplementary material
Below is the link to the electronic supplementary material.
11103_2019_897_MOESM2_ESM.xlsx
Supplementary material 2 Table S1. Distribution of SNBEs, SMREs and AC elements in the promoter region of the 56 phenylpropanoid and monolignol biosynthetic genes in S. viridis. (XLSX 92 kb)
11103_2019_897_MOESM3_ESM.xlsx
Supplementary material 3 Table S2. List of bona fide genes involved in the phenylpropanoid and monolignol biosynthetic pathways in Arabidopsis and some grasses. (XLSX 63 kb)
11103_2019_897_MOESM4_ESM.xlsx
Supplementary material 4 Table S3. Protein sequence identity matrixes of all phenylpropanoid and monolignol gene families. (XLSX 60 kb)
11103_2019_897_MOESM5_ESM.xlsx
Supplementary material 5 Table S4. RNAseq expression data (FPKM) of all phenylpropanoid and monolignol biosynthetic genes in S. viridis retrieved from the work of Martin et al. (2016). (XLSX 61 kb)
Rights and permissions
About this article
Cite this article
Ferreira, S.S., Simões, M.S., Carvalho, G.G. et al. The lignin toolbox of the model grass Setaria viridis. Plant Mol Biol 101, 235–255 (2019). https://doi.org/10.1007/s11103-019-00897-9
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s11103-019-00897-9