Abstract
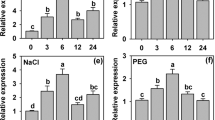
A novel CBL-interacting protein kinase (CIPK) gene, ZmCIPK16, was isolated from maize (Zea mays), which has been certified to have two copies in the genome. The ZmCIPK16 is strongly induced in maize seedlings by PEG, NaCl, ABA, dehydration, heat and drought, but not by cold. A yeast two-hybrid assay demonstrated that ZmCIPK16 interacted with ZmCBL3, ZmCBL4, ZmCBL5, and ZmCBL8. Bimolecular fluorescence complementation (BiFC) assays prove that ZmCIPK16 can interact with ZmCBL3, ZmCBL4, ZmCBL5, and ZmCBL8 in vivo. Subcellular localization showed that ZmCIPK16 is distributed in the nucleus, plasma membrane and cytoplasm; this is different from the specific localization of ZmCBL3, ZmCBL4, and ZmCBL5, which are found in the plasma membrane. The results also showed that overexpression of ZmCIPK16 in the Arabidopsis sos2 mutant induced the expression of the SOS1 gene and enhanced salt tolerance. These findings indicate that ZmCIPK16 may be involved in the CBL-CIPK signaling network in maize responses to salt stress.









Similar content being viewed by others
Abbreviations
- ABA:
-
Abscisic acid
- BiFC:
-
Bimolecular fluorescence complementation
- CBL:
-
Calcineurin B-like protein
- CIPK:
-
CBL-interacting protein kinase
- ORF:
-
Open reading frame
- PEG:
-
Polyethylene glycol
- RACE:
-
Rapid amplification of cDNA ends
- SOS:
-
Salt overly sensitive
References
Ahn S, Tanksley SD (1993) Comparative linkage maps of the rice and maize genomes. Proc Natl Acad Sci USA 90:7980–7984. doi:10.1073/pnas.90.17.7980
Albrecht V, Ritz O, Linder S, Harter K, Kudla J (2001) The NAF domain defines a novel protein–protein interaction module conserved in Ca2+-regulated kinase. EMBO J 20:1051–1063. doi:10.1093/emboj/20.5.1051
Albrecht V, Weinl S, Blazevic D, Angelo DC, Batistic O, Kolukisaoglu U, Bock R, Schulz B, Harter K, Kudla J (2003) The calcium sensor CBL1 integrates plant responses to abiotic stresses. Plant J 36:457–470. doi:10.1046/j.1365-313X.2003.01892.x
Appel RD, Bairoch A, Hochstrasser DF (1994) A new generation of information retrieval tools for biologists: the example of the ExPASy www Server. Trends Biochem Sci 19:258–260. doi:10.1016/0968-0004(94)90153-8
Batistic O, Sorek N, SchuSltke S, Yalovsky S, Kudla J (2008) Dual fatty acyl modification determines the localization and plasma membrane targeting of CBL/CIPK Ca2+ signaling complexes in Arabidopsis. Plant Cell 20(5):1346–1362. doi:10.1105/tpc.108.058123
Cheong YH, Pandey GK, Grant JJ, Batistic O, Li L, Kim BG, Lee SC, Kudla J, Luan S (2007) Two calcineurin B-like calcium sensors, interacting with protein kinase CIPK23, regulate leaf transpiration and root potassium uptake in Arabidopsis. Plant J 52:223–239. doi:10.1111/j.1365-313X.2007.03236.x
Clough SJ, Bent AF (1998) Floral dip: A simplified method for Agrobacterium-mediated transformation of Arabidopsis thaliana. Plant J 16:735–743. doi:10.1046/j.1365-313x.1998.00343.x
D’Angelo C, Weinl S, Batistic O, Pandey GK, Cheong YH, Schultke S, Albrecht V, Ehlert B, Schulz B, Harter K, Luan S, Bock R, Kudla J (2006) Alternative complex formation of the Ca2+-regulated protein kinase CIPK1 controls abscisic acid dependent and independent stress responses in Arabidopsis. Plant J 48:857–872. doi:10.1111/j.1365-313X.2006.02921.x
Flowers TJ, Troke PF, Yeo AR (1977) The mechanism of salt tolerance in halopytes. Annu Rev Plant Physiol 28:89–121. doi:10.1146/annurev.pp.28.060177.000513
Fu RZ, Sun YR, Jia SR (1994) Plant genetic transformation. China Science and Technology Press, Beijing
Gao P, Zhao PM, Wang J, Wang HY, Du XM, Wang GL, Xia GX (2008) Co-expression and preferential interaction between two calcineurin B-like proteins and a CBL-interacting protein kinase from cotton. Plant Physiol Biochem 46(10):935–940. doi:10.1016/j.plaphy.2008.05.001
Gilroy S, Trewavas A (2001) Signal processing and transduction in plant cells: the end of the beginning. Nat Rev Mol Cell Biol 2:307–314. doi:10.1038/35067109
Gong D, Guo Y, JK Z, Schumaker KS (2004) The SOS3 family of calcium sensors and SOS2 family of protein kinases in Arabidopsis. Plant Physiol 134:919–926. doi:10.1104/pp.103.037440
Halford NG, Hardie DG (1998) SNF1-related protein kinase: global regulators of carbon metabolism in plants. Plant Mol Biol 37:735–748. doi:10.1023/A:1006024231305
Halfter U, Ishitan M, Zhu JK (2000) The Arabidopsis SOS2 protein kinase physically interacts with and is activated by the calcium-binding protein SOS3. Proc Natl Acad Sci USA 97:3735–3740. doi:10.1073/pnas.040577697
Hardie DG (1999) Plant protein serine/threonine kinases: classification and functions. Annu Rev Plant Physiol Plant Mol Biol 50:97–131. doi:10.1146/annurev.arplant.50.1.97
Harper JF (2001) Dissecting calcium oscillators in plant cells. Trends Plant Sci 6:395–397. doi:10.1016/S1360-1385(01)02023-4
Hwang YS, Bethke PC, Cheong YH, Chang HS, Zhu T, Jones RL (2005) A gibberellin-regulated calcineurin B in rice localizes to the tonoplast and is implicated in vacuole function. Plant Physiol 138:1347–1358. doi:10.1104/pp.105.062703
Jin YS, Ma DY DJL, Jin JC, Li DF, Deng CW, Wang T (2007) HC-Pro protein of Potato Virus Y can interact with three Arabidopsis 20S proteasome subunits in planta. J Virol 81(23):12881–12888. doi:10.1128/JVI.00913-07
Kay R, Chan A, Daly M, Mcpherson J (1987) Duplication of CaMV 35S promoter sequences creates a strong enhancer for plant genes. Science 236:1299–1302. doi:10.1126/science.236.4806.1299
Kim KN, Cheong YH, Gupta R, Luan S (2000) Interaction specificity of Arabidopsis calcineurin B-like calcium sensors and their target kinases. Plant Physiol 124:1844–1853. doi:10.1104/pp.124.4.1844
Kim KN, Cheong YH, Grant JJ, Pandey GK, Luan S (2003a) CIPK3, a calcium sensor-associated protein kinase that regulates abscisic acid and cold signal transduction in Arabidopsis. Plant Cell 15:411–423. doi:10.1105/tpc.006858
Kim KN, Lee JS, Han H, Choi SA, Go SJ, Yoon IS (2003b) Isolation and characterization of a novel rice Ca2+-regulated protein kinase ene involved in responses to diverse signals including cold, light, cytokinins, sugars and salts. Plant Mol Biol 52:1191–1202. doi:10.1023/B:PLAN.0000004330.62660.a2
Knight H, Knight MR (2001) Abiotic stress signalling pathways: specificity and cross-talk. Trends Plant Sci 6:262–267. doi:10.1016/S1360-1385(01)01946-X
Kolukisaoglu U, Weinl S, Blazevic D, Batistic O, Kudla J (2004) Calcium sensors and their interacting protein kinases: genomics of the Arabidopsis and rice CBL-CIPK signaling networks. Plant Physiol 134:43–58. doi:10.1104/pp.103.033068
Kudla J, Batisti O (2004) Integration and channeling of calcium signaling through the CBL calcium sensor/CIPK protein kinase network. Planta 219:915–924. doi:10.1007/s00425-004-1333-3
Kudla J, Xu Q, Harter K, Gruissem W, Luan S (1999) Genes for calcineurin B-like proteins in Arabidopsis are differentially regulated by stress signals. Proc Natl Acad Sci USA 96:4718–4723. doi:10.1073/pnas.96.8.4718
Liu J, Zhu JK (1998) A calcium sensor homolog required for plant salt tolerance. Science 280:1943–1945. doi:10.1126/science.280.5371.1943
Luan S, Kudla J, Rodriguez-Concepcion M, Yalovsky S, Gruissem W (2002) Calmodulins and calcineurin B-like proteins: calcium sensors for specific signal response coupling in plants. Plant Cell 14(suppl):S389–S400
Mahajan S, Sopory SK, Tuteja N (2006) Cloning and characterization of CBL-CIPK signaling components from a legume (Pisum sativum). FEBS J 273:907–925. doi:10.1111/j.1742-4658.2006.05111.x
Martínez-Atienza J, Jiang X, Garciadeblas B, Mendoza I, Zhu JK, Pardo JM, Quintero FJ (2007) Conservation of the salt overly sensitive pathway in rice. Plant Physiol 143:1001–1012. doi:10.1104/pp.106.092635
Murashige T, Skoog F (1962) A revised medium for rapid growth and bioassays with tobacco tissue culture. Physiol Plant 15:473–497. doi:10.1111/j.1399-3054.1962.tb08052.x
Ohba H, Steward N, Kawasaki S, Berberich T, Ikeda Y, Koizumi N, Kusano T, Sano H (2000) Diverse response of rice and maize genes encoding homologs of WPK4, an SNF1-related protein kinase from wheat, to light, nutrients, low temperature and cytokinins. Mol Gen Genet 263:359–366. doi:10.1007/s004380051179
Pandey GK, Cheong YH, Kim KN, Grant JJ, Li L, Hung W, D’Angelo C, Weinl S, Kudla J, Luan S (2004) The calcium sensor calcineurin B-Like 9 modulates abscisic acid sensitivity and biosynthesis in Arabidopsis. Plant Cell 16(7):1912–1924. doi:10.1105/tpc.021311
Qiu QS, Guo Y, Dietrich MA, Schumaker KS, Zhu JK (2002) Regulation of SOS1, a plasma membrane Na+/H+ exchanger in Arabidopsis thaliana, by SOS2 and SOS3. Proc Natl Acad Sci USA 99:8436–8441. doi:10.1073/pnas.122224699
Quan RD, Lin HX, Mendoza I, Zhang YG, Cao WH, Yang YQ, Shang M, Chen SY, Pardo JM, Guo Y (2007) SCABP8/CBL10, a putative calcium sensor, interacts with the protein kinase SOS2 to protect Arabidopsis shoots from salt stress. Plant Cell 19:1415–1431. doi:10.1105/tpc.106.042291
Quintero FJ, Ohta M, Shi H, Zhu JK, Pardo JM (2002) Reconstitution in yeast of the Arabidopsis SOS signaling pathway for Na+ homeostasis. Proc Natl Acad Sci USA 99:9061–9066. doi:10.1073/pnas.132092099
Sambrook J, Russell DW (2001) Molecular cloning: a laboratory manual, 3rd edn. Science Press, Beijing
Schinkmann K, Blenis J (1997) Cloning and characterization of a human STE20-like protein kinase with unusual cofactor requirements. J Biol Chem 272:28695–28703. doi:10.1074/jbc.272.45.28695
Shi J, Kim KN, Ritz O, Albrecht V, Gupta R, Harter K, Luan S, Kudla J (1999) Novel protein kinases associated with calcineurin B-like calcium sensors in Arabidopsis. Plant Cell 11:2393–2405
Shi H, Ishitani M, Kim C, Zhu JK (2000) The Arabidopsis thaliana salt tolerance gene SOS1 encodes a putative Na+/H+ antiporter. Proc Natl Acad Sci USA 97:6896–6901. doi:10.1073/pnas.120170197
Shi H, Quintero FJ, Pardo JM, Zhu JK (2002) The putative plasma membrane Na+/H+ antiporter SOS1 controls long-distance Na+ transport in plants. Plant Cell 14:465–477. doi:10.1105/tpc.010371
Shi H, Lee B, Wu SJ, Zhu JK (2003) Overexpression of a plasma membrane Na+/H+ antiporter improves salt tolerance in Arabidopsis. Nat Biotechnol 2:81–85. doi:10.1038/nbt766
Shinozaki K, Yamaguchi-Shinozaki K (1994) A novel cis-acting element in an Arabidopsis gene is involved in responsiveness to drought, low-temperature, or high-salt stress. Plant Cell 6:251–264
Waadt R and Kudla J (2008) In planta visualization of protein interactions using bimolecular fluorescence complementation (BiFC). CSH Protocols (online). doi:10.1101/pdb.prot4995
Wang MY, Gu D, Liu TS, Wang ZQ, Guo XY, Hou W, Bai YF, Chen XP, Wang GY (2007) Overexpression of a putative maize calcineurin B-like protein in Arabidopsis confers salt tolerance. Plant Mol Biol 65:733–746. doi:10.1007/s11103-007-9238-8
Walter M, Chaban C, Schutze K, Batistic O, Weckermann K, Nake C, Blazevic D, Grefen C, Schumacher K, Oecking C, Harter K, Kudla J (2004) Visualization of protein interactions in living plant cells using bimolecular fluorescence complementation. Plant J 40(3):428–438. doi:10.1111/j.1365-313X.2004.02219.x
Xu J, Li HD, Chen LQ, Wang Y, Liu LL, He L, Wu WH (2006) A protein kinase, interacting with two calcineurin B-like proteins, regulates K+ transporter AKT1 in Arabidopsis. Cell 125:1347–1360. doi:10.1016/j.cell.2006.06.011
Zheng J, Zhao JF, Tao YZ, Wang JH, Liu YJ, Fu JJ, Jin Y, Gao P, Zhang JP, Bai YF, Wang GY (2004) Isolation and analysis of water stress induced genes in maize seedlings by subtractive PCR and cDNA macroarray. Plant Mol Biol 55:807–823
Acknowledgments
The authors thank Dr. Zhizhong Chen (China Agricultural University) for providing Arabidopsis sos2 mutants, Dr. Klaus Harter (Botanisches Institut, Universitat zu Koln, Germany) for the vectors used in the BiFC assay, Liping Liang and Xiping Shi for help with the confocal microscope. This work was supported by the National High-tech Program of China (2006AA10Z103), the Natural Science Foundation of China (30500286) and the research grant from the Institute of Crop Sciences, CAAS (082060302–18).
Author information
Authors and Affiliations
Corresponding author
Additional information
Jinfeng Zhao and Zhenfei Sun are contributed equally to this work.
Electronic supplementary material
Below is the link to the electronic supplementary material.
11103_2008_9445_MOESM3_ESM.doc
Subcellular localization of GFP, ZmCIPK16-GFP, ZmCBL3-GFP,ZmCBL4-GFP and ZmCBL5-GFP in onion cells. Left, fluorescence images; right, Bright-field; they were taken from the same cell for each protein localization assay a/b, pBI221-GFP; c/d, ZmCIPK16-GFP; e/f, ZmCBL3-GFP; g/h, ZmCBL4-GFP; i/j, ZmCBL5-GFP. (DOC 27 kb)
11103_2008_9445_MOESM4_ESM.tif
Southern analysis of ZmCIPK16 in transgenic events of Arabidopsis T1 generation. Genomic DNA (10 μg) of different T1 transgenic line (9–2、17–3、19–1、20–4) was digested with EcoRI respectively, separated by electrophoresed on 0.8% (w/v) agarose gel and transferred onto Hybond-N+ nylon membrane. A 523 bp cDNA fragment located in the 5’ region of the ZmCIPK16 open reading frame was labeled with α-32p-dCTP as probe. (TIFF 11561 kb)
11103_2008_9445_MOESM5_ESM.tif
The expression of ZmCIPK16 in Arabidopsis wild type and transgenic plants. 10 μg total RNA from two-week-old seedlings were loading. A 523 bp cDNA fragment located in the 5’ region of the ZmCIPK16 open reading frame was labeled with α-32p-dCTP as probe. rRNA was visualized by staining with methylene blue as loading control. (TIFF 12788 kb)
Rights and permissions
About this article
Cite this article
Zhao, J., Sun, Z., Zheng, J. et al. Cloning and characterization of a novel CBL-interacting protein kinase from maize. Plant Mol Biol 69, 661–674 (2009). https://doi.org/10.1007/s11103-008-9445-y
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s11103-008-9445-y