Abstract
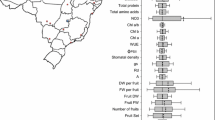
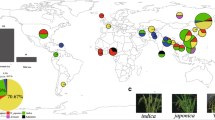
The aim of the study was to investigate the genetic distances and their relationships among pepper species using photosynthetic features under different stresses and genetic variability. The photosynthetic features under drought, waterlogging and low-temperature stresses, rDNA internal transcribed spacer (ITS) sequences of nuclear genome and trnH-psbA sequence of chloroplast genome of 25 varieties from 5 pepper species Capsicum annuum L. (CA), Capsicum baccatum L. (CB), Capsicum chinense Jacquin. (CC), Capsicum frutescens L. (CF) and Capsicum pubescens Ruiz & Pavon (CP) were analyzed and used to construct the dendrograms. The results showed the photosynthetic rate of different pepper species could be greatly but differentially decreased by stresses. For example, CB and CF had the smallest and the highest decrease to drought, CC had the highest decrease to waterlogging, and CP had the smallest decrease to low temperature. The ITS sequences of 25 pepper varieties are 591–619 bp in length and have GC% between 51.1% and 64.5%. Their trnH-psbA sequences are 537–558 bp in length and have GC% between 27.2% and 28.5%. The cluster analysis of the five pepper species based on the changes in P N under stresses is similar to that based on genetic variability, that is, CP clusters with CB, and CC clusters with CA after first clusters with CF. In addition, the clustering methods based on the photosynthetic stress responses and genetic variability are unable to completely distinguish pepper varieties within the same species. The results indicate that similarly to genetic variability, changes in P N under stresses (specifically the stress corresponding to the climate of plant’s original habitat) could be used to identify genetic distance of pepper species.
Similar content being viewed by others
Abbreviations
- te]CA:
-
Capsicum annuum L.
- CB:
-
Capsicum baccatum L.
- CC:
-
Capsicum chinense Jacquin.
- CF:
-
Capsicum frutescens L.
- CP:
-
Capsicum pubescens Ruiz &Pavon
- E :
-
transpiration rate
- g s :
-
stomatal conductance
- ITS:
-
rDNA internal transcribed spacer
- P N :
-
net photosynthetic rate
- WUE:
-
water-use efficiency
References
Bahadur, B.J.: Studies on interspecific and intraspecific genetic relationships in Capsicum using randomly amplified polymorphic DNA markers and chloroplast DNA sequences. — Ph.D. Thesis, New Mexico State Univ., New Mexico 2003.
Christopher, H. J., Kenneth, J. S., Harvey, E. B. J.: Evolutionary relationships, interisland biogeography, and molecular evolution in the Hawaiian violets (Viola: Violaceae). — Am. J. Bot. 96: 2087–2099, 2009.
Egawa, Y.: [Cytogenetical relationships between wild and cultivated Capsicum peppers and their phylogenetic differentiation.] — Ph. D. Thesis, Dep. Genet. Resources, Nat. Inst. Agrobiol. Resources, Tsukuba Sci. City, Tsukuba 1985. [In Jap.]
Egawa, Y., Tanaka, M.: Cytogenetical study of the interspecific hybrid between Capsicum annuum and Capsicum baccatum. — Jap. J. Breed. 36: 16–21, 1986.
Farquhar, G.D., Buckley, T.N., Miller, J.M.: Optimal stomatal control in relation to leaf area and nitrogen content. — Silva Fenn. 36: 625–637, 2002.
Fischer, R.A., Turner, N.C.: Plant productivity in the arid and semiarid zones. — Ann. Rev. Plant Physiol. 29: 227–317. 1978.
Font, M., Garcia-Jacas, N., Vilatersana, R., Roquet, C., Susanna, A.: Evolution and biogeography of Centaurea section Acrocentron inferred from nuclear and plastid DNA sequence analyses. — Ann. Bot. 103: 985–997, 2009.
Frankel, N., Hasson, E., Iusem, N.D., Rossi, M.S.: Adaptive evolution of the water stress-induced gene Asr2 in Lycopersicon species dwelling in arid habitats. — Mol. Biol. Evol. 20: 1955–1962, 2003.
Fu, Q.S., Li, H.L., Cui, J., Zhao, B., Guo, Y.D.: [Effects of water stress on photosynthesis and associated physiological characters of Capsicum annuum L.] — Sci. Agric. Sin. 42: 1859–1866, 2009. [In Chin.]
Huang, G.P., Gong, S., Xu, W.L., Li, P., Zhang, D.J., Qin, L. X., Li, W., Li, X.B. GhHyPRP4, a cotton gene encoding putative hybrid proline-rich protein, is preferentially expressed in leaves and involved in plant response to cold stress. — Acta Biochim. Biophys. Sin. 43: 519–527, 2011.
Jarret, R.L.: DNA barcoding in a crop genebank: resolving the Capsicum annuum species complex. — Open Biol. J. 1: 35–42, 2008.
John, K.W., Ericksona, D.L., Andrew, F.J., Nathan, G.S., Perezb, R., Sanjurb, O., Bermingham, E.: Plant DNA barcodes and a community phylogeny of a tropical forest dynamics plot in Panama. — PNAS 106: 18621–18626, 2009.
Lefebvre, V., Palloix, A., Max, R.: Nuclear RFLP between pepper species (Capsicum annuum L). — Euphytica 71: 189–199, 1993.
Liu, X.Y., Mao, X.F.: [Study on the effect of adversity on plant photosynthetic physiology.] — J. Hebei. Agric. Sci. 12: 12–14, 2008. [In Chin.]
McLeod, M.J., Guttman, S.I., Eshbaugh, W.H.: Early evolution of chili peppers (Capsicum). — Econ. Bot. 36: 361–8, 1982.
Ou, L.J., Dai, X.Z., Zhang, Z.Q., Zou, X.X.: Responses of pepper to waterlogging stress. — Photosynthetica 49: 339–345, 2011.
Peterson, A., John, H., Koch, E., Peterson, J.: A molecular phylogeny of the genus Gagea (Liliaceae) in Germany inferred from non-coding chloroplast and nuclear DNA sequences. — Plant Syst. Evol. 245: 145–162, 2004.
Pickersgill, B.: Relationships between weedy and cultivated forms in some species of chili peppers (genus Capsicum). — Evolution 25: 683–91, 1971.
Pickersgill, B.: Genetic resources and breeding of Capsicum spp. — Euphytica 96: 129–133, 1997.
Prince, J.P., Lackney, V.K., Angeles, C., Blauth, J.R., Kyle, M.M.: A survey of DNA polymorphism within the genus Capsicum and the fingerprinting of pepper species. — Genome 38: 224–231, 1995.
Rogers, O.S., Bendich, A.J.: Extraction of DNA plant tissue, plant molecular. — In: Gelvin, S.B., Schilpe, R.A., Verna, D.S. (ed.): Plant Molecular Biology Manual. Pp. A6: 1–10. Kluwer Academic Publishers, Dordecht 1998.
Sun, X.C., Hu, C.X., Tan, Q.L., Liu, J.S., Liu, H.G.: Effects of molybdenum on expression of cold-responsive genes in abscisic acid (ABA)-dependent and ABA-independent pathways in winter wheat under low-temperature stress. — Ann. Bot. 104: 345–356, 2009.
Vijaykumar, A., Saini, A., Jawali, N. Phylogenetic analysis of Subgenus vigna species using nuclear Ribosomal RNA ITS: Evidence of hybridization among vigna unguiculata subspecies. — J. Hered. 101: 177–188, 2010.
Weih, M., Bonosi, L., Ghelardini, L., Roönberg-Wästljung, a. c.: Optimizing nitrogen economy under drought: increased leaf nitrogen is an acclimation to water stress in willow (Salix spp.). — Ann. Bot. 108: 1347–1353, 2011.
White, T.J., Bruns, T., Lee, S., Taylor, J.W.: Amplification and direct sequencing of fungal ribosomal RNA genes for phylogenetics. — In: Innis, M.A., Gelfand, D.H., Sninsky, J.J., White, T.J. (ed.): PCR Protocols: A Guide to Methods and Applications. Pp. 315–322. Academic Press, New York 1990.
Yan, J., Deng, J., Zhou, C.J., Zhong, B. Y., Hao, F.: Phenotypic and molecular characterization of Madurella pseudomycetomatis sp. nov., a novel opportunistic fungus possibly causing black-grain mycetoma. — J. Clin. Microbiol. 48: 251–257, 2010.
Author information
Authors and Affiliations
Corresponding author
Additional information
Acknowledgements: This work was supported by China Agriculture Research System (CARS-25-A-8).
Rights and permissions
About this article
Cite this article
Ou, L.J., Zou, X.X. The photosynthetic stress responses of five pepper species are consistent with their genetic variability. Photosynthetica 50, 49–55 (2012). https://doi.org/10.1007/s11099-012-0008-8
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s11099-012-0008-8