Abstract
Background
Machine learning algorithms (MLAs) carry a huge potential in identifying predicting factors and are being explored for their utility in the field of personalized medicine.
Aim
We aimed to investigate MLAs for identifying predictors (clinical and genetic) of poor anticoagulation status (ACS) and stable weekly warfarin dose (SWWD).
Method
Clinical factors, in addition to the CYP2C9, VKORC1, and CYP4F2 genotypes, were obtained for patients receiving warfarin for at least the previous six months. The C5.0 decision tree classification algorithm was used to predict poor ACS while classification and regression tree analysis (CART), in addition to the Chi-square automatic interaction detector (CHAID), was used to predict SWWD. The percentage of patients within 20% of the actual dose, root mean squared error (RMSE), and area under the receiver-operating characteristics curve (AUROC) were identified as performance indicators of the models.
Results
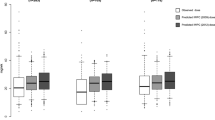
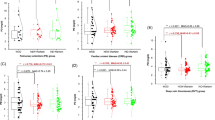
In the C5.0 classification decision tree, the CYP4F2 genotype was the strongest predictor of ACS (AUROC = 0.53). In the CART analysis of SWWD, VKORC1 polymorphisms were the most significant predictor, followed by the CYP2C9 genotype (percentage of patients within 20% of the actual dose = 38.2%, RMSE = 13.6). For the CHAID algorithm, the percentage of patients within 20% of the actual dose was 49%, while the RMSE was found to be 13.4.
Conclusion
Genetic and non-genetic predictive factors were identified by the MLAs for ACS and SWWD. Further, the need to externally validate the MLAs in a prospective study was highlighted.


Similar content being viewed by others
References
Lee MT, Klein TE. Pharmacogenetics of warfarin: challenges and opportunities. J Hum Genet. 2013;58(6):334–8.
Loebstein R, Yonath H, Peleg D, et al. Interindividual variability in sensitivity to warfarin-Nature or nurture? Clin Pharmacol Ther. 2001;70(2):159–64.
Yang W, Ma J, Hu W, et al. Associated factors and safety of the rapidly achieving first therapeutic target of warfarin in hospitalized patients: a retrospective cohort study. Int J Clin Pharm. 2022. https://doi.org/10.1007/s11096-022-01404-9.
Sridharan K, Al Banny R, Qader AM, et al. Health-related quality of life in patients receiving oral anti-coagulants: a cross-sectional study. Expert Rev Cardiovasc Ther. 2020;18(5):309–14.
Bader LA, Elewa H. The impact of genetic and non-genetic factors on warfarin dose prediction in MENA region: a systematic review. PLoS ONE. 2016;11(12):e0168732.
Bazan NS, Sabry NA, Rizk A, et al. Validation of pharmacogenetic algorithms and warfarin dosing table in Egyptian patients. Int J Clin Pharm. 2012;34(6):837–44.
Sridharan K, Al Banna R, Malalla Z, et al. Influence of CYP2C9, VKORC1, and CYP4F2 polymorphisms on the pharmacodynamic parameters of warfarin: a cross-sectional study. Pharmacol Rep. 2021;73(5):1405–17.
Asiimwe IG, Zhang EJ, Osanlou R, et al. Warfarin dosing algorithms: a systematic review. Br J Clin Pharmacol. 2021;87(4):1717–29.
Fahmi AM, Elewa H, El Jilany I. Warfarin dosing strategies evolution and its progress in the era of precision medicine, a narrative review. Int J Clin Pharm. 2022. https://doi.org/10.1007/s11096-022-01386-8.
Jiang T, Gradus JL, Rosellini AJ. Supervised machine learning: a brief primer. Behav Ther. 2020;51(5):675–87.
Knights J, Chanda P, Sato Y, et al. Vertical integration of pharmacogenetics in population PK/PD modeling: a novel information theoretic method. CPT Pharmacomet Syst Pharmacol. 2013;2(2):e25.
Lin E, Lin CH, Lane HY. Machine learning and deep learning for the pharmacogenomics of antidepressant treatments. Clin Psychopharmacol Neurosci. 2021;19(4):577–88.
Kalafi EY, Nor NAM, Taib NA, et al. Machine learning and deep learning approaches in breast cancer survival prediction using clinical data. Folia Biol (Praha). 2019;65(5–6):212–20.
Le Corre PA. Prescriptome analytics: an opportunity for clinical pharmacy. Int J Clin Pharm. 2019;41(6):1394–7.
Liu R, Li X, Zhang W, et al. Comparison of nine statistical model based warfarin pharmacogenetic dosing algorithms using the racially diverse international warfarin pharmacogenetic consortium cohort database. PLoS ONE. 2015;10(8):e0135784.
Ma Z, Wang P, Gao Z, et al. Ensemble of machine learning algorithms using the stacked generalization approach to estimate the warfarin dose. PLoS ONE. 2018;13(10):e0205872.
Gordon J, Norman M, Hurst M, et al. Using machine learning to predict anticoagulation control in atrial fibrillation: a UK clinical practice research datalink study. Inform Med Unlocked. 2021;25:100688.
Alsagheer RHA, Alharan AFH, Al-Haboobi ASA. Popular decision tree algorithms of data mining techniques: a review. Int J Comp Sci Mob Comp. 2017;6:133–42.
International Warfarin Pharmacogenetics Consortium, Klein TE, Altman RB, Eriksson N, et al. Estimation of the warfarin dose with clinical and pharmacogenetic data. N Engl J Med. 2009;360(8): 753–64.
Collins GS, Reitsma JB, Altman DG, et al. Transparent reporting of a multivariable prediction model for Individual prognosis or diagnosis (TRIPOD): the TRIPOD statement. J Clin Epidemiol. 2015;68(2):134–43.
Sridharan K, Al Banna R, Qader AM, et al. Evaluation of inter-patient variability in the pharmacodynamic indices of warfarin. Expert Rev Cardiovasc Ther. 2020;18(11):835–40.
International Warfarin Pharmacogenetics Consortium (IWPC). PharmGKB. Available at: https://www.pharmgkb.org/downloads. Accessed 14 Jun 2022.
Virdee MS, Stewart D. Optimizing the use of oral anticoagulant therapy for atrial fibrilation in primary care: a pharmacist-led intervention. Int J Clin Pharm. 2017;39(1):173–80.
Siddiqui S, DeRemer CE, Waller JL, et al. Variability in the calculation of time in therapeutic range for the quality control measurement of warfarin. J Innov Card Rhythm Manag. 2018;9(12):3428–34.
Dallalzadeh LO, Go AS, Chang Y, et al. Stability of high-quality warfarin anticoagulation in a community-based atrial fibrillation cohort: The anticoagulation and risk factors in atrial fibrillation (ATRIA) study. J Am Heart Assoc. 2016;5(7):e003482.
Sridharan K, Banny RA, Husain A. Evaluation of stable doses of warfarin in a patient cohort. Drug Res (Stuttg). 2020;70(12):570–5.
Sharabiani A, Darabi H, Bress A, et al. Machine learning based prediction of warfarin optimal dosing for African American patients. In: 2013 IEEE international conference on automation science and engineering (CASE). 2013. p. 623–8. https://doi.org/10.1109/CoASE.2013.6653999
Asiimwe IG, Blockman M, Cohen K, et al. Stable warfarin dose prediction in sub-Saharan African patients: A machine-learning approach and external validation of a clinical dose-initiation algorithm. CPT Pharmacometrics Syst Pharmacol. 2022;11(1):20–9.
Nguyen VL, Nguyen HD, Cho YS, et al. Comparison of multivariate linear regression and a machine learning algorithm developed for prediction of precision warfarin dosing in a Korean population. J ThrombHaemost. 2021;19(7):1676–86.
Li JX, Kim MH, Song K, et al. The influence of CYP4F2 polymorphisms on warfarin doses in Korean patients with a variety of diseases. Clin Exp ThrombHemost. 2018;4:7–10.
Caldwell MD, Awad T, Johnson JA, et al. CYP4F2 genetic variant alters required warfarin dose. Blood. 2008;111:4106–12.
Wei M, Ye F, Xie D, et al. A new algorithm to predict warfarin dose from polymorphisms of CYP4F2, CYP2C9 and VKORC1 and clinical variables: derivation in Han Chinese patients with non valvular atrial fibrillation. ThrombHaemost. 2012;108:1083–91.
Cha PC, Mushiroda T, Takahashi A, et al. Genome-wide association study identifies genetic determinants of warfarin responsiveness for Japanese. Hum Mol Genet. 2010;19:4735–44.
Sun X, Yu WY, Ma WL, et al. Impact of the CYP4F2 gene polymorphisms on the warfarin maintenance dose: A systematic review and meta-analysis. Biomed Rep. 2016;4(4):498–506.
Zhang JE, Klein K, Jorgensen AL, et al. Effect of Genetic Variability in the CYP4F2, CYP4F11, and CYP4F12 Genes on Liver mRNA Levels and Warfarin Response. Front Pharmacol. 2017;8:323.
Shendre A, Brown TM, Liu N, et al. Race-specific influence of CYP4F2 on dose and risk of hemorrhage among warfarin users. Pharmacotherapy. 2016;36(3):263–72.
Liew C-L, Yen J-H, Liu A-B, et al. Sex differences in the effective warfarin dosage in Han and aboriginal Taiwanese patients with the VKORC1-1639AA genotype. Tzu Chi Med J. 2013;25:213–7.
Absher RK, Moore ME, Parker MH. Patient-specific factors predictive of warfarin dosage requirements. Ann Pharmacother. 2002;36:1512–7.
Choi JR, Kim JO, Kang DR, et al. Proposal of pharmacogenetics-based warfarin dosing algorithm in Korean patients. J Hum Genet. 2011;56:290–5.
Acknowledgements
We thank the staff nurses in the anticoagulation clinic for their immense co-operation during the data collection.
Funding
The study was carried out as a part of warfarin pharmacogenetic study, with the Funding Number: AGU RCSI-2019-02.
Author information
Authors and Affiliations
Corresponding author
Ethics declarations
Conflicts of interest
All authors declare that there is no relevant financial or non-financial interest to disclose.
Additional information
Publisher's Note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary Information
Below is the link to the electronic supplementary material.
Rights and permissions
Springer Nature or its licensor holds exclusive rights to this article under a publishing agreement with the author(s) or other rightsholder(s); author self-archiving of the accepted manuscript version of this article is solely governed by the terms of such publishing agreement and applicable law.
About this article
Cite this article
Sridharan, K., Ramanathan, M. & Al Banna, R. Evaluation of supervised machine learning algorithms in predicting the poor anticoagulation control and stable weekly doses of warfarin. Int J Clin Pharm 45, 79–87 (2023). https://doi.org/10.1007/s11096-022-01471-y
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s11096-022-01471-y