Abstract
Objectives
Maximum a posteriori Bayesian estimation (MAP-BE) based on a limited sampling strategy and a population pharmacokinetic (POPPK) model is used to estimate individual pharmacokinetic parameters. Recently, we proposed a methodology that combined population pharmacokinetic and machine learning (ML) to decrease the bias and imprecision in individual iohexol clearance prediction. The aim of this study was to confirm the previous results by developing a hybrid algorithm combining POPPK, MAP-BE and ML that accurately predicts isavuconazole clearance.
Methods
A total of 1727 isavuconazole rich PK profiles were simulated using a POPPK model from the literature, and MAP-BE was used to estimate the clearance based on: (i) the full PK profiles (refCL); and (ii) C24h only (C24h-CL). Xgboost was trained to correct the error between refCL and C24h-CL in the training dataset (75%). C24h-CL as well as ML-corrected C24h-CL were evaluated in a testing dataset (25%) and then in a set of PK profiles simulated using another published POPPK model.
Results
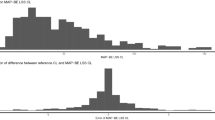
A strong decrease in mean predictive error (MPE%), imprecision (RMSE%) and the number of profiles outside ± 20% MPE% (n-out20%) was observed with the hybrid algorithm (decreased in MPE% by 95.8% and 85.6%; RMSE% by 69.5% and 69.0%; n-out20% by 97.4% and 100% in the training and testing sets, respectively. In the external validation set, the hybrid algorithm decreased MPE% by 96%, RMSE% by 68% and n-out20% by 100%.
Conclusion
The hybrid model proposed significantly improved isavuconazole AUC estimation over MAP-BE based on the sole C24h and may improve dose adjustment.
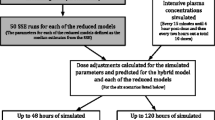
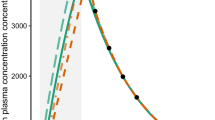
Graphical Abstract





Similar content being viewed by others
Data Availability
Data generated for this study are available upon request to the corresponding author.
References
Marquet P, Destère A, Monchaud C, Rérolle J-P, Buchler M, Mazouz H, et al. Clinical Pharmacokinetics and Bayesian Estimators for the Individual Dose Adjustment of a Generic Formulation of Tacrolimus in Adult Kidney Transplant Recipients. Clin Pharmacokinet. 2021;60:611–22.
Labriffe M, Woillard J, Debord J, Marquet P. Machine learning algorithms to estimate everolimus exposure trained on simulated and patient pharmacokinetic profiles. CPT Pharmacom & Syst Pharma. 2022;psp4.12810.
Woillard J-B, Saint-Marcoux F, Debord J, Åsberg A. Pharmacokinetic models to assist the prescriber in choosing the best tacrolimus dose. Pharmacol Res. 2018;130:316–21.
Destere A, Gandonnière CS, Åsberg A, Loustaud‐Ratti V, Carrier P, Ehrmann S, et al. A single Bayesian estimator for iohexol clearance estimation in ICU, liver failure and renal transplant patients. Brit J Clinical Pharma. 2021;bcp.15197.
Ponthier L, Ensuque P, Destere A, Marquet P, Labriffe M, Jacqz-Aigrain E, et al. Optimization of Vancomycin Initial Dose in Term and Preterm Neonates by Machine Learning. Pharm Res [Internet]. 2022 [cited 2022 Aug 11]; Available from: https://link.springer.com/10.1007/s11095-022-03351-6.
Franck B, Autmizguine J, Åsberg A, Théorêt Y, Marquet P, Ovetchkine P, et al. Thoroughly Validated Bayesian Estimator and Limited Sampling Strategy for Dose Individualization of Ganciclovir and Valganciclovir in Pediatric Transplant Recipients. Clin Pharmacokinet. 2021;60:1449–62.
Woillard J-B, Debord J, Benz-de-Bretagne I, Saint-Marcoux F, Turlure P, Girault S, et al. A Time-Dependent Model Describes Methotrexate Elimination and Supports Dynamic Modification of MRP2/ABCC2 Activity. Ther Drug Monit. 2017;39:12.
Benkali K, Rostaing L, Premaud A, Woillard J-B, Saint-Marcoux F, Urien S, et al. Population Pharmacokinetics and Bayesian Estimation of Tacrolimus Exposure in Renal Transplant Recipients on a New Once-Daily Formulation. Clin Pharmacokinet. 2010;49:683–92.
Goecks J, Jalili V, Heiser LM, Gray JW. How Machine Learning Will Transform Biomedicine. Cell. 2020;181:92–101.
Tang B-H, Guan Z, Allegaert K, Wu Y-E, Manolis E, Leroux S, et al. Drug Clearance in Neonates: A Combination of Population Pharmacokinetic Modelling and Machine Learning Approaches to Improve Individual Prediction. Clin Pharmacokinet. 2021;60:1435–48.
Koch G, Pfister M, Daunhawer I, Wilbaux M, Wellmann S, Vogt JE. Pharmacometrics and Machine Learning Partner to Advance Clinical Data Analysis. Clin Pharmacol Ther. 2020;107:926–33.
Hughes JH, Keizer RJ. A hybrid machine learning/pharmacokinetic approach outperforms maximum a posteriori Bayesian estimation by selectively flattening model priors. CPT Pharmacometrics Syst Pharmacol. 2021;10:1150–60.
Li Z, Li R, Niu W, Zheng X, Wang Z, Zhong M, et al. Population Pharmacokinetic Modelling Combined with Machine Learning Approach Improved Tacrolimus Trough Concentrations Prediction in Chinese Adult Liver Transplant Recipients. The Journal of Clinical Pharma. 2022;jcph.2156.
Destere A, Marquet P, Gandonnière CS, Åsberg A, Loustaud-Ratti V, Carrier P, et al. A Hybrid Model Associating Population Pharmacokinetics with Machine Learning: A Case Study with Iohexol Clearance Estimation. Clin Pharmacokinet [Internet]. 2022 [cited 2022 Jun 9]; Available from: https://link.springer.com/https://doi.org/10.1007/s40262-022-01138-x.
Rybak JM, Marx KR, Nishimoto AT, Rogers PD. Isavuconazole: Pharmacology, Pharmacodynamics, and Current Clinical Experience with a New Triazole Antifungal Agent. Pharmacotherapy. 2015;35:1037–51.
Townsend R, Dietz A, Hale C, Akhtar S, Kowalski D, Lademacher C, et al. Pharmacokinetic Evaluation of CYP3A4-Mediated Drug-Drug Interactions of Isavuconazole With Rifampin, Ketoconazole, Midazolam, and Ethinyl Estradiol/Norethindrone in Healthy Adults. Clin Pharmacol Drug Dev. 2017;6:44–53.
Sivasubramanian G, Chandrasekar PH. Efficacy and safety of Isavuconazole for the treatment of invasive Aspergillus infection - an update of the literature. Expert Opin Pharmacother. 2022;23:543–9.
Maertens JA, Raad II, Marr KA, Patterson TF, Kontoyiannis DP, Cornely OA, et al. Isavuconazole versus voriconazole for primary treatment of invasive mould disease caused by Aspergillus and other filamentous fungi (SECURE): a phase 3, randomised-controlled, non-inferiority trial. The Lancet. 2016;387:760–9.
Kaindl T, Andes D, Engelhardt M, Saulay M, Larger P, Groll AH. Variability and exposure–response relationships of isavuconazole plasma concentrations in the Phase 3 SECURE trial of patients with invasive mould diseases. J Antimicrob Chemother. 2019;74:761–7.
Furfaro E, Signori A, Di Grazia C, Dominietto A, Raiola AM, Aquino S, et al. Serial monitoring of isavuconazole blood levels during prolonged antifungal therapy. J Antimicrob Chemother. 2019;74:2341–6.
Kosmidis C, Otu A, Moore CB, Richardson MD, Rautemaa-Richardson R. Isavuconazole Therapeutic Drug Monitoring during Long-Term Treatment for Chronic Pulmonary Aspergillosis. Antimicrob Agents Chemother. 2020;65:e01511-e1520.
Wu X, Venkataramanan R, Rivosecchi RM, Tang C, Marini RV, Shields RK, et al. Population Pharmacokinetics of Intravenous Isavuconazole in Solid-Organ Transplant Recipients. Antimicrob Agents Chemother. 2020;64:e01728-e1819.
Elmokadem A, Riggs MM, Baron KT. Quantitative systems pharmacology and physiologically-based pharmacokinetic modeling with mrgsolve: a hands-on tutorial. CPT Pharmacometrics Syst Pharmacol. 2019;8:883–93.
Kuhn M, Wickham H. Tidymodels: a collection of packages for modeling and machine learning using tidyverse principles [Internet]. 2020. Available from: https://www.tidymodels.org. Accessed 17 Feb 2023.
Monolix version 2018R1. Antony. France: Lixoft SAS; 2018. http://lixoft.com/products/monolix/. Accessed 17 Feb 2023.
Desai A, Kovanda L, Kowalski D, Lu Q, Townsend R, Bonate PL. Population Pharmacokinetics of Isavuconazole from Phase 1 and Phase 3 (SECURE) Trials in Adults and Target Attainment in Patients with Invasive Infections Due to Aspergillus and Other Filamentous Fungi. Antimicrob Agents Chemother. 2016;60:5483–91.
Le Louedec F, Puisset F, Thomas F, Chatelut É, White-Koning M. Easy and reliable maximum a posteriori Bayesian estimation of pharmacokinetic parameters with the open-source R package mapbayr. CPT Pharmacometrics Syst Pharmacol. 2021;10:1208–20.
Warn PA, Sharp A, Parmar A, Majithiya J, Denning DW, Hope WW. Pharmacokinetics and Pharmacodynamics of a Novel Triazole, Isavuconazole: Mathematical Modeling, Importance of Tissue Concentrations, and Impact of Immune Status on Antifungal Effect. Antimicrob Agents Chemother. 2009;53:3453–61.
Seyedmousavi S, Brüggemann RJM, Meis JF, Melchers WJG, Verweij PE, Mouton JW. Pharmacodynamics of Isavuconazole in an Aspergillus fumigatus Mouse Infection Model. Antimicrob Agents Chemother. 2015;59:2855–66.
Lepak AJ, Marchillo K, VanHecker J, Diekema D, Andes DR. Isavuconazole Pharmacodynamic Target Determination for Candida Species in an In Vivo Murine Disseminated Candidiasis Model. Antimicrob Agents Chemother. 2013;57:5642–8.
Arendrup MC, Meletiadis J, Mouton JW, Guinea J, Cuenca-Estrella M, Lagrou K, et al. EUCAST technical note on isavuconazole breakpoints for Aspergillus, itraconazole breakpoints for Candida and updates for the antifungal susceptibility testing method documents. Clin Microbiol Infect. 2016;22:571.e1-571.e4.
Bououda M, Uster DW, Sidorov E, Labriffe M, Marquet P, Wicha SG, et al. A Machine Learning Approach to Predict Interdose Vancomycin Exposure. Pharm Res. 2022;39:721–31.
Chen T, Guestrin C. XGBoost: A Scalable Tree Boosting System. Proceedings of the 22nd ACM SIGKDD International Conference on Knowledge Discovery and Data Mining [Internet]. San Francisco California USA: ACM; 2016 [cited 2021 Dec 24]. p. 785–94. Available from: https://dl.acm.org/doi/https://doi.org/10.1145/2939672.2939785.
Uster DW, Wicha SG. Optimized sampling to estimate vancomycin drug exposure: Comparison of pharmacometric and equation-based approaches in a simulation-estimation study. CPT Pharmacom & Syst Pharma. 2022;11:711–20.
Acknowledgments
We gratefully thank Miss Karen Poole for manuscript editing.
Author information
Authors and Affiliations
Contributions
AD, JBW contributed to the conception and design of the study, analysis and interpretation of data, PM contributed to the conception and design of the study and interpretation of data, MLB and MD contributed to the interpretation of data. All the authors participated in drafting the article and approved the final version submitted.
Corresponding author
Ethics declarations
Consent for Publication
All the authors read and approved the final manuscript.
Conflicts of Interest
None of the author have any conflicts of interest to declare in relation to this work.
Additional information
Publisher's Note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Study Highlight
- What question did this study address?
This study evaluated the performances of a hybrid model combining population pharmacokinetics (POPPK) and machine learning to improve individual isavuconazole clearance estimation in comparison to POPPK alone.
- What does this study add to our knowledge?
A decreased by about 90% and 70% of the bias and imprecision was observed in comparison to the MAP-BE alone for prediction based on trough concentration.
- How might this change drug discovery, development, and/or therapeutics?
The hybrid model developed may spearhead a new generation of tools for MIPD in routine practice.
Supplementary Information
Below is the link to the electronic supplementary material.
Rights and permissions
Springer Nature or its licensor (e.g. a society or other partner) holds exclusive rights to this article under a publishing agreement with the author(s) or other rightsholder(s); author self-archiving of the accepted manuscript version of this article is solely governed by the terms of such publishing agreement and applicable law.
About this article
Cite this article
Destere, A., Marquet, P., Labriffe, M. et al. A Hybrid Algorithm Combining Population Pharmacokinetic and Machine Learning for Isavuconazole Exposure Prediction. Pharm Res 40, 951–959 (2023). https://doi.org/10.1007/s11095-023-03507-y
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s11095-023-03507-y