Abstract
Purpose
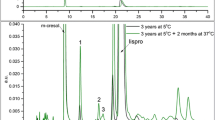
The main purposes of this manuscript are to report a surprising and interesting degradation reaction of glucagon from a specific vendor in which glucagon underwent cleavage among several peptide bonds quickly under near neutral to basic conditions, and to propose the root cause of mechanism for the degradation reaction.
Methods
The degradation reaction was monitored by HPLC and the fragment structures were confirmed by LC-MS. Possible impurities responsible for the degradation were either confirmed or excluded by a variety of techniques such as addition of chelator EDTA and transitional metal ions or separation by ultrafiltration.
Results
This type of degradation was rarely reported in literature, especially considering its extreme cleavage efficiency. Contamination by a thermostable high molecular impurity (such as a peptidase with molecular weight between 10 and 30 KDa) during the manufacturing process was the main reason for this interesting phenomenon.
Conclusions
The degradation phenomenon described here could be used as an excellent example showing that products ordered from vendors meeting the rudimentary quality standards might contain impurities which could cause significant degradation. We suggest that a simple solution, i.e. additional tests of stability under real or accelerated conditions by manufacturers and inclusion of the “accelerated stability criteria” in the Certificate of Analysis (CoAs), especially for sensitive biological reagents prone to faster degradation, would be very helpful for avoiding losses for both vendors and users.














Similar content being viewed by others
Abbreviations
- Amino acid:
-
Were abbreviated according to the rules of the IUPAC-IUB Joint Commission of Biochemical Nomenclature (Eur. J. Biochem. 1984, 138, 9–37)
- EDTA:
-
Ethylenediaminetetraacetic acid
- LC-MS:
-
Liquid chromatography-mass spectrometry
- MALDI-TOF MS:
-
Matrix-assisted laser desorption/ionization time-of-flight mass spectrometry
- RP-HPLC:
-
Reversed-phase high-performance liquid chromatography
- tR :
-
Retention time
References
Granner D, Pilkis S. The genes of hepatic glucose metabolism. J Biol Chem. 1990;265:10173–6.
Joshi AB, Rus E, Kirsch LE. The degradation pathways of glucagon in acidic solutions. Int J Pharm. 2000;203:115–25.
Joshi AB, Kirsch LE. The relative rates of glutamine and asparagine deamidation in glucagon fragment 22-29 under acidic conditions. J Pharm Sci. 2002;91:2331–45.
Joshi AB, Kirsch LE. The estimation of glutaminyl deamidation and aspartyl cleavage rates in glucagon. Int J Pharm. 2004;273:213–9.
Joshi AB, Sawai M, Kearney WR, Kirsch LE. Studies on the mechanism of aspartic acid cleavage and glutamine deamidation in the acidic degradation of glucagon. J Pharm Sci. 2005;94:1912–27.
Caputo N, Castle JR, Bergstrom CP, Carroll JM, Bakhtiani PA, Jackson MA, et al. Mechanisms of glucagon degradation at alkaline pH. Peptides. 2013;45:40–7.
Fang W-J, Qi W, Kinzell J, Prestrelski S, Carpenter JF. Effects of excipients on the chemical and physical stability of glucagon during freeze-drying and storage in dried formulations. Pharm Res. 2012;29:3278–91.
Ghodke S, Nielsen SB, Christiansen G, Hjuler HA, Flink J, Otzen D. Mapping out the multistage fibrillation of glucagon. FEBS J. 2012;279:752–65.
Moorthy BS, Ghomi HT, Lill MA, Topp EM. Structural transitions and interactions in the early stages of human glucagon amyloid fibrillation. Biophys J. 2015;108:937–48.
Pedersen JS, Dikov D, Otzen DE. N- and C-terminal hydrophobic patches are involved in fibrillation of glucagon. Biochemistry. 2006;45:14503–12.
Smith MA, Easton M, Everett P, Lewis G, Payne M, Riveros-Moreno V, et al. Specific cleavage of immunoglobulin G by copper ions. Int J Pept Protein Res. 1996;48:48–55.
Glover ZK, Basa L, Moore B, Laurence JS, Sreedhara A. Metal ion interactions with mAbs: part 1. mAbs. 2015;7:901–11.
Chin J. Developing artificial hydrolytic metalloenzymes by a unified mechanistic approach. Acc Chem Res. 1991;24:145–52.
Smith MA, Easton M, Everett P, Lewis G, Payne M. Specific cleavage of immunoglobulin G by copper ions. Int J Pept Protein Res. 1996;48:48–55.
Vlasak J, Ionescu R. Fragmentation of monoclonal antibodies. mAbs. 2011;3:253–63.
Gaza-Bulseco G, Liu H. Fragmentation of a recombinant monoclonal antibody at various pH. Pharm Res. 2008;25:1881–90.
Liu H, Gaza-Bulseco G, Sun J. Characterization of the stability of a fully human monoclonal IgG after prolonged incubation at elevated temperature. J Chromatogr B Analyt Technol Biomed Life Sci. 2006;837:35–43.
Kamerzell TJ, Li M, Arora S, Ji JA, Wang YJ. The relative rate of immunoglobulin gamma 1 fragmentation. J Pharm Sci. 2011;100:1341–9.
Ravuluri S, Bansal R, Chhabra N, Rathore AS. Kinetics and characterization of non-enzymatic fragmentation of monoclonal antibody therapeutics. Pharm Res. 2018;35:142.
Gao SX, Zhang Y, Stansberry-Perkins K, Buko A, Bai S, Nguyen V, et al. Fragmentation of a highly purified monoclonal antibody attributed to residual CHO cell protease activity. Biotechnol Bioeng. 2011;108:977–82.
Acknowledgments and Disclosures
We thank the Ministry of Science and Technology of China (Grant No. 2018ZX09J18107–002), National Natural Science Foundation of China (Grant No. 81741144), and the Fundamental Research Funds for the Central Universities (Grant No. 2019XZZX003–18) for their financial support. We would also like to extend our thanks to Zhiwei Ge and Youping Xu at the Analysis Center of Agrobiology and Environmental Sciences, Zhejiang University for their help on LC-MS and MALDI-TOF MS analysis, respectively. We have no personal financial or non-financial conflicts of interest.
Author information
Authors and Affiliations
Corresponding author
Additional information
Publisher’s Note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Rights and permissions
About this article
Cite this article
Zheng, HJ., Shen, BB., Wang, J. et al. Uncommon Peptide Bond Cleavage of Glucagon from a Specific Vendor under near Neutral to Basic Conditions. Pharm Res 36, 118 (2019). https://doi.org/10.1007/s11095-019-2647-y
Received:
Accepted:
Published:
DOI: https://doi.org/10.1007/s11095-019-2647-y