Abstract
Purpose
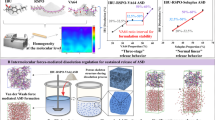
The pH-responsive copolymer micelles are widely used as carriers in drug delivery system, but there are few micro-level mechanistically explorations on the pH-triggered drug release. Here we elucidate the relationship between drug release behavior of four/six-arms star copolymer micelles and the copolymer structures.
Method
The net cumulative drug release percentage (En) was taken as the dependent variables, block unit autocorrelation descriptors as independent variables. The quantitative structure-property relationship models of drug release from block copolymers were developed at pH 7.4 and 5.0 of two periods (stage I: 0~12 h, stage II: 12~96 h).
Results
The models built are of good fitting ability, internal predictive ability, stability and statistically significance. Drug diffusion is mainly influenced by the intra-block force, and micellar erosion by inter-block force. At pH 5.0, lowest unoccupied molecular orbital energy of copolymer unit is the main factor influencing the En. Stage I of drug release is affected by hydrophobic property and stage II by regional polar of copolymer molecules.
Conclusion
The models present good performance, factors affecting drug release behavior at different pH conditions can offer guidance for the design of copolymer structures to control the drug release behavior of micelles in a targeted and quantitatively way.



Similar content being viewed by others
Abbreviations
- δ :
-
Standardized residual of the sample to be predicted
- σ :
-
Standard residual of the samples
- BUA:
-
Block unit autocorrelation
- E n :
-
Net cumulative drug release percentage during a stage
- E r :
-
Cumulative drug release percentage at time points
- F :
-
Value of F in F-test
- f k :
-
Polymeric molecular characteristic
- f ki :
-
Characteristic value of unit i
- h* :
-
Critical value of leverage h
- h i :
-
Leverage of the ith sample to be predicted
- l :
-
Block unit interval
- LUMO:
-
Lowest unoccupied molecular orbital
- m :
-
Number of samples in the training set
- n :
-
Number of query samples
- N :
-
Unit counts in the summation
- p :
-
Possibility
- q :
-
Total number of descriptors contained in the models
- q’ :
-
Number of descriptors plus one
- Q 2 LOO-CV :
-
Leave-one-out cross-validation correlation coefficient
- QSPR:
-
Quantitative structure-property relationship
- R 2 :
-
Multiple correlation coefficient
- R i 2 :
-
Multiple correlation coefficient in a regression analysis between independent variables xi and other independent variables
- R 2 Y-rand, Q 2 Y-rand :
-
Average values of R2 or Q2LOO-CV in Y-randomization test
- s :
-
Standardized error
- x i :
-
Characteristic matrix of the ith sample to be predicted
- X T :
-
Transpose of matrix X
References
Chen Z, Zhang P, Cheetham AG, Moon JH, Moxley JW, Lin Y, et al. Controlled release of free doxorubicin from peptide–drug conjugates by drug loading. J Control Release. 2014;191:123–30.
Nie SY, Lin WJ, Yao N, Guo XD, Zhang LJ. Drug release from pH-sensitive polymeric micelles with different drug distributions: insight from coarse-grained simulations. ACS Appl Mater Interfaces. 2014;6:17668–78.
Chen H, Ruckenstein E. Formation and degradation of multicomponent multicore micelles: insights from dissipative particle dynamics simulations. Langmuir. 2013;29:5428–34.
Rodríguez-Hidalgo M-R, Soto-Figueroa C, Vicente L. Mesoscopic simulation of the drug release mechanism on the polymeric vehicle P(ST-DVB) in an acid environment. Soft Matter. 2011;7:8224.
Wang Y, Huang J-J, Zhou N, Cao D-S, Dong J, Li H-X. Incorporating PLS model information into particle swarm optimization for descriptor selection in QSAR/QSPR. J Chemom. 2015;29:627–36.
Kale SP, Garg S. Prediction of the mutual diffusion coefficient for controlled drug delivery devices. Comput Chem Eng. 2012;39:186–98.
Gafourian T, Safari A, Adibkia K, Parviz F, Nokhodchi A. A drug release study from hydroxypropylmethylcellulose (HPMC) matrices using QSPR modeling. J Pharm Sci. 2007;96:3334–51.
Pajander J, Korhonen O, Laamanen M, Ryynänen E-L, Grimsey I, van Veen B, et al. Effect of formulation parameters and drug–polymer interactions on drug release from starch acetate matrix tablets. J Pharm Sci. 2009;98:3676–90.
Huang X, Xiao Y, Lang M. Self-assembly of pH-sensitive mixed micelles based on linear and star copolymers for drug delivery. J Colloid Interface Sci. 2011;364:92–9.
Cao W, Zhu L. Synthesis and unimolecular micelles of amphiphilic dendrimer-like star polymer with various functional surface groups. Macromolecules. 2011;44:1500–12.
Liu J, Huang W, Pang Y, Zhu X, Zhou Y, Yan D. Self-assembled micelles from an amphiphilic Hyperbranched copolymer with polyphosphate arms for drug delivery. Langmuir. 2010;26:10585–92.
Yang YQ, Zhao B, Li ZD, Lin WJ, Zhang CY, Guo XD, et al. pH-sensitive micelles self-assembled from multi-arm star triblock co-polymers poly(ε-caprolactone)-b-poly(2-(diethylamino)ethyl methacrylate)-b-poly(poly(ethylene glycol) methyl ether methacrylate) for controlled anticancer drug delivery. Acta Biomater. 2013;9:7679–90.
Lin WJ, Nie SY, Chen Q, Qian Y, Wen XF, Zhang LJ. Structure-property relationship of pH-sensitive (PCL) 2 (PDEA- b -PPEGMA) 2 micelles: experiment and DPD simulation. AIChE J. 2014;60:3634–46.
Lin W, Nie S, Zhong Q, Yang Y, Cai C, Wang J, et al. Amphiphilic miktoarm star copolymer (PCL)3-(PDEAEMA-b-PPEGMA)3 as pH-sensitive micelles in the delivery of anticancer drug. J Mater Chem B. 2014;2:4008.
Katritzky AR, Sild S, Karelson M. Correlation and prediction of the refractive indices of polymers by QSPR. J Chem Inf Comput Sci. 1998;38:1171–6.
García-Domenech R, de Julián-Ortiz JV. Prediction of indices of refraction and glass transition temperatures of linear polymers by using graph theoretical indices. J Phys Chem B. 2002;106:1501–7.
Wu W, Zhang R, Peng S, Li X, Zhang L. QSPR between molecular structures of polymers and micellar properties based on block unit autocorrelation (BUA) descriptors. Chemom Intell Lab Syst. 2016;157:7–15.
Karcher W, Devillers J, editors. Practical applications of quantitative structure-activity relationships (QSAR) in environmental chemistry and toxicology, vol. 1. New York: Springer Science & Business Media; 1990.
Yousefinejad S, Hemmateenejad B, Mehdipour AR. New autocorrelation QTMS-based descriptors for use in QSAM of peptides. J Iran Chem Soc. 2012;9:569–77.
Imran M, Baig AQ, Ali H. On molecular topological properties of hex-derived networks. J Chemom. 2016;30:121–9.
Todeschini R, Consonni V. Molecular Descriptors for Chemoinformatics: Volume I: Alphabetical Listing / Volume II: Appendices, References. New York: Wiley; 2009.
Karcher W, Devillers J. Practical applications of quantitative structure-activity relationships (QSAR) in environmental chemistry and toxicology. New York: Springer Science & Business Media; 1990.
Mo YZ, Xu JC. Studies on mechanical properties and optimization model of PI/SiO2 nanocomposite based on materials studio. Adv Mater Res. 2014;1049–1050:54–7.
Niazi A, Leardi R. Genetic algorithms in chemometrics. J Chemom. 2012;26:345–51.
Leardi R. Genetic algorithms in chemometrics and chemistry: a review. J Chemom. 2001;15:559–69.
Lei B, Ma Y, Li J, Liu H, Yao X, Gramatica P. Prediction of the adsorption capability onto activated carbon of a large data set of chemicals by local lazy regression method. Atmos Environ. 2010;44:2954–60.
Li J, Lei B, Liu H, Li S, Yao X, Liu M, et al. QSAR study of malonyl-CoA decarboxylase inhibitors using GA-MLR and a new strategy of consensus modeling. J Comput Chem. 2008;29:2636–47.
Wang J, Krudy G, Xie X-Q, Wu C, Holland G. Genetic algorithm-optimized QSPR models for bioavailability, protein binding, and urinary excretion. J Chem Inf Model. 2006;46:2674–83.
Gramatica P, Giani E, Papa E. Statistical external validation and consensus modeling: a QSPR case study for Koc prediction. J Mol Graph Model. 2007;25:755–66.
Gramatica P, Pilutti P, Papa E. Validated QSAR prediction of OH tropospheric degradation of VOCs: splitting into training−test sets and consensus modeling. J Chem Inf Comput Sci. 2004;44:1794–802.
Guan P, Doytchinova IA, Walshe VA, Borrow P, Flower DR. Analysis of peptide−protein binding using amino acid descriptors: prediction and experimental verification for human histocompatibility complex HLA-A*0201. J Med Chem. 2005;48:7418–25.
Eriksson L, Jaworska J, Worth AP, Cronin MTD, McDowell RM, Gramatica P. Methods for reliability and uncertainty assessment and for applicability evaluations of classification- and regression-based QSARs. Environ Health Perspect. 2003;111:1361–75.
Tropsha A, Gramatica P, Gombar V. The importance of being Earnest: validation is the absolute essential for successful application and interpretation of QSPR models. QSAR Comb Sci. 2003;22:69–77.
Kiralj R, Ferreira MMC. Basic validation procedures for regression models in QSAR and QSPR studies: theory and application. J Braz Chem Soc. 2009;20:55.
Dearden JC, Cronin MTD, Kaiser KLE. How not to develop a quantitative structure–activity or structure–property relationship (QSAR/QSPR). SAR QSAR Environ Res. 2009;20:241–66.
Roy K, Kabir H. QSPR with extended topochemical atom (ETA) indices, 3: modeling of critical micelle concentration of cationic surfactants. Chem Eng Sci. 2012;81:169–78.
Rücker C, Rücker G, Meringer M. y-randomization and its variants in QSPR/QSAR. J Chem Inf Model. 2007;47:2345–57.
Wehrens R, Putter H, Buydens LM. The bootstrap: a tutorial. Chemom Intell Lab Syst. 2000;54:35–52.
Gharagheizi F, Eslamimanesh A, Ilani-Kashkouli P, Mohammadi AH, Richon D. QSPR molecular approach for representation/prediction of very large vapor pressure dataset. Chem Eng Sci. 2012;76:99–107.
Peppas NA, Sahlin JJ. A simple equation for the description of solute release. III. Coupling of diffusion and relaxation. Int J Pharm. 1989;57:169–72.
Author information
Authors and Affiliations
Corresponding author
Additional information
Publisher’s Note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Electronic Supplementary Material
ESM 1
(DOCX 60 kb)
Rights and permissions
About this article
Cite this article
Zhang, R., Wen, Ly., Wu, Ws. et al. Quantitative Structure-Property Relationship for pH-Triggered Drug Release Performance of Acid-Responsive Four/Six-Arms Star Polymeric Micelles. Pharm Res 36, 20 (2019). https://doi.org/10.1007/s11095-018-2549-4
Received:
Accepted:
Published:
DOI: https://doi.org/10.1007/s11095-018-2549-4