ABSTRACT
Purpose
To describe a computational tool to calculate molecular descriptors of potential application in ADME virtual screening of antitumor Pt(II) drug candidates.
Methods
The multistep computational procedure consists in (a) building and optimization (PM3) of the 3D structures of the investigated complexes, (b) parametrization of Pt(II) and its implementation in GRID, (c) GRID calculations and extraction of the information content with VolSurf and BIOCUBE4mf, and (d) PLS analysis to look for the correlation between experimental data and the molecular descriptors.
Results
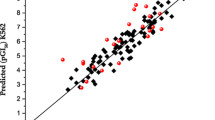
The following results were obtained: (a) the calibration of the GRID force field to take into account the platinum di-cation, (b) solid PLS models between log k30 and log kw with VolSurf descriptors which highlight the main structural differences between the two chromatographic parameters, (c) the prediction of virtual (of each conformer) log k30 and log kw, and (d) the identification of the main descriptors governing VDss of drugs in clinical use.
Conclusion
The study suggests a strategy to identify good Pt(II) complexes prior to their synthesis to eliminate as soon as possible drug candidates with unfavorable PK profile.






Similar content being viewed by others
REFERENCES
Kelland L. The resurgence of platinum-based cancer chemotherapy. Nat Rev Cancer. 2007;7(8):573–84.
Kelland L, Farrell N. Platinum-based drugs in cancer therapy. Totowa: Humana Pre; 2000.
Gabano E, Ravera M, Osella D. The drug targeting and delivery approach applied to Pt-antitumour complexes. A coordination point of view. Curr Med Chem. 2009;16(34):4544–80.
Gaviraghi G, Barnaby RJ, Pellegatti M. In: Testa B, van de Waterbeemd H, Folkers G, Guy R, editors. Pharmacokinetic challenges in lead optimization. Zurich: Wiley-VHCA; 2001. p. 3–14.
Tetko IV, Jaroszewicz I, Platts JA, Kuduk-Jaworska J. Calculation of lipophilicity for Pt(II) complexes: experimental comparison of several methods. J Inorg Biochem. 2008;102(7):1424–37.
Nasal A, Siluk D, Kaliszan R. Chromatographic retention parameters in medicinal chemistry and molecular pharmacology. Curr Med Chem. 2003;10:381–426.
Martel S, Guillarme D, Henchoz Y, Galland A, Veuthey J, Rudaz S, et al. Chromatographic approaches for measuring log P. In: Mannhold R, editor. Molecular drug properties. Measurement and prediction. Darmstadt: Wiley-VCH; 2008. p. 331–55.
Valko K. Application of high-performance liquid chromatography based measurements of lipophilicity to model biological distribution. J Chromatogr A. 2004;1037(1–2):299–310.
Boobbyer DN, Goodford PJ, Mcwhinnie PM, Wade RC. New hydrogen-bond potentials for use in determining energetically favorable binding sites on molecules of known structure. J Med Chem. 1989;32:1083–94.
Goodford PJ. A computational procedure for determining energetically favorable binding sites on biologically important macromolecules. J Med Chem. 1985;28:849–57.
Wade RC, Goodford PJ. Further development of hydrogen bond functions for use in determining energetically favorable binding sites on molecules of known structure. 2. Ligand probe groups with the ability to form more than two hydrogen bonds. J Med Chem. 1993;36:148–56.
Cruciani G, Mannhold R, Kubinyi H, Folkers G. Molecular interaction fields. Applications in drug discovery and ADME prediction. Zurich: Wiley-VCH; 2006.
Caron G, Ermondi G. Lipophilicity, polarity, and hydrophobicity. In: Testa B, van de Waterbeemd H, editors. Comprehensive medicinal chemistry, vol. 5. 2nd ed. Oxford: Elsevier; 2006. p. 425–52.
Caron G, Ermondi G. Calculating virtual log P in the alkane/water system \( \left( {{ \log }\,{\hbox{P}}_{\rm{alk}}^{\rm{N}}} \right) \) and its derived parameters \( \Delta { \log }\,{\hbox{P}}_{{{\rm{oct}} - {\rm{alk}}}}^{\rm{N}} \) and \( { \log }\,{\hbox{D}}_{\rm{alk}}^{\rm{pH}} \). J Med Chem. 2005;48(9):3269–79.
Vistoli G, Pedretti A, Testa B. Partition coefficient and molecular flexibility: the concept of lipophilicity space. Chem Biodivers. 2009;6:1152–69.
Obach RS, Lombardo F, Waters NJ. Trend analysis of a database of intravenous pharmacokinetic parameters in humans for 670 drug compounds. Drug Metab Dispos. 2008;36(7):1385–405.
Berellini G, Springer C, Waters NJ, Lombardo F. In silico prediction of volume of distribution in human using linear and nonlinear models on a 669 compound data set. J Med Chem. 2009;52(14):4488–95.
Caron G, Ermondi G, Gariboldi MB, Monti E, Gabano E, Ravera M, Osella D. The relevance of polar surface area (PSA) in rationalizing biological properties of several cis-diamminemalonatoplatinum(II) derivatives. ChemMedChem. 2009;4((10):1677–85.
Platts J, Oldfield SP, Reif MM, Palmucci A, Gabano E, Osella D. The RP-HPLC measurement and QSPR analysis of logP(o/w) values of several Pt(II) complexes. J Inorg Biochem. 2006;100(7):1199–207.
Heudi O, Mercier-Jobard S, Cailleux A, Allain P. Mechanisms of reaction of L-methionine with carboplatin and oxaliplatin in different media: a comparison with cisplatin. Biopharm Drug Dispos. 1999;20(2):107–16.
Coste F, Malinge JM, Serre L, Shepard W, Roth M, Leng M, et al. Crystal structure of a double-stranded DNA containing a cisplatin interstrand cross-link at 1.63Å resolution: hydration at the platinated site. Nucleic Acids Res. 1999;27(8):1837–46.
Coste F, Shepard W, Zelwer C. Description of ordered solvent molecules in a platinated decanucleotide duplex refined at 1.6Å resolution against experimental MAD phases. Acta Crystallogr Sect D. 2002;58:431–40.
Pastor M, Cruciani G, Watson KA. A strategy for the incorporation of water molecules present in a ligand binding site into a three-dimensional quantitative structure-activity relationship analysis. J Med Chem. 1997;40:4089–102.
Caron G, Nurisso A, Ermondi G. How to extend the use of grid-based interaction energy maps from chemistry to biotopics. ChemMedChem. 2009;4(1):29–36.
Pagliara A, Khamis E, Trinh A, Carrupt PA, Tsai RS, Testa B. Structural properties governing retention mechanisms on RP-HPLC stationary phase used for lipophilicity measurements. J Liq Chromatogr. 1995;18:1721–45.
McIntosh DP, Cooke RJ, McLachlan AJ, Daley-Yates PT, Rowland M. Pharmacokinetics and tissue distribution of cisplatin and conjugates of cisplatin with carboxymethyldextran and A5B7 monoclonal antibody in CD1 mice. J Pharm Sci. 1997;86(12):1478–83.
Welink J, Boven E, Vermorken JB, Gall HE, van der Vijgh WJ. Pharmacokinetics and pharmacodynamics of lobaplatin (D-19466) in patients with advanced solid tumors, including patients with impaired renal of liver function. Clin Cancer Res. 1999;5(9):2349–58.
Ishibashi T, Yano Y, Oguma T. Population pharmacokinetics of platinum after nedaplatin administration and model validation in adult patients. Br J Clin Pharmacol. 2003;56(2):205–13.
Ehrsson H, Wallin I, Yachnin J. Pharmacokinetics of oxaliplatin in humans. Med Oncol. 2002;19(4):261–5.
Lombardo F, Obach R, Shalaeva MY, Gao F. Prediction of volume of distribution values in humans for neutral and basic drugs using physicochemical measurements and plasma protein binding data. J Med Chem. 2002;45:2867–76.
Lombardo F, Obach R, Shalaeva MY, Gao F. Prediction of human volume of distribution values for neutral and basic drugs. 2. Extended data set and leave-class-out statistics. J Med Chem. 2004;47:1242–50.
Author information
Authors and Affiliations
Corresponding author
Electronic supplementary material
Below is the link to the electronic supplementary material.
ESM 1
Supporting Information: Chemical structures and experimental lipophilicity indexes, VIPs and PLS coefficients; VolSurf descriptors definitions; X-ray structures used for building models. (DOC 553 kb)
Rights and permissions
About this article
Cite this article
Caron, G., Ravera, M. & Ermondi, G. Molecular Interaction Fields (MIFs) to Predict Lipophilicity and ADME Profile of Antitumor Pt(II) Complexes. Pharm Res 28, 640–646 (2011). https://doi.org/10.1007/s11095-010-0317-1
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s11095-010-0317-1