Abstract
FUS/TLS (fused in sarcoma/translocated in liposarcoma) encodes a multifunctional DNA/RNA binding protein with non-classical carboxy (C)-terminal nuclear localization signal (NLS). A variety of ALS-linked mutations are clustered in the C-terminal NLS, resulting in the cytoplasmic mislocalization and aggregation. Since the arginine methylations are implicated in the nuclear-cytoplasmic shuttling of FUS, a methylation inhibitor could be one of therapeutic targets for FUS-linked ALS. We here examined effects of methylation inhibitors on the cytoplasmic mislocalization and aggregates of ALS-linked C-terminal FUS mutant in a cell culture system. Treatment with adenosine dialdehyde (AdOx), a representative global methyltransferase inhibitor, remarkably mitigated the cytoplasmic mislocalization and aggregation of FUS mutant, which is consistent with previous reports. However, AdOx treatment of higher concentration and longer time period evoked the intranuclear aggregation of the ectopic expressed FUS protein. The pull down assay and the morphological analysis indicated the binding between FUS and Transportin could be potentiated by AdOx treatment through modulating methylation status in RGG domains of FUS. These findings indicated the treatment with a methylation inhibitor at the appropriate levels could alleviate the cytoplasmic mislocalization but in excess this could cause the intranuclear aggregation of FUS C-terminal mutant.



Similar content being viewed by others
References
Vance C, Rogelj B, Hortobagyi T, De Vos KJ, Nishimura AL, Sreedharan J, Hu X, Smith B, Ruddy D, Wright P, Ganesalingam J, Williams KL, Tripathi V, Al-Saraj S, Al-Chalabi A, Leigh PN, Blair IP, Nicholson G, de Belleroche J, Gallo JM, Miller CC, Shaw CE (2009) Mutations in FUS, an RNA processing protein, cause familial amyotrophic lateral sclerosis type 6. Science 323:1208–1211
Kwiatkowski TJ Jr, Bosco DA, Leclerc AL, Tamrazian E, Vanderburg CR, Russ C, Davis A, Gilchrist J, Kasarskis EJ, Munsat T, Valdmanis P, Rouleau GA, Hosler BA, Cortelli P, de Jong PJ, Yoshinaga Y, Haines JL, Pericak-Vance MA, Yan J, Ticozzi N, Siddique T, McKenna-Yasek D, Sapp PC, Horvitz HR, Landers JE, Brown RH Jr (2009) Mutations in the FUS/TLS gene on chromosome 16 cause familial amyotrophic lateral sclerosis. Science 323:1205–1208
Lagier-Tourenne C, Cleveland DW (2009) Rethinking ALS: the FUS about TDP-43. Cell 136:1001–1004
Lagier-Tourenne C, Polymenidou M, Cleveland DW (2010) TDP-43 and FUS/TLS: emerging roles in RNA processing and neurodegeneration. Hum Mol Genet 19:R46–R64
Dormann D, Rodde R, Edbauer D, Bentmann E, Fischer I, Hruscha A, Than ME, Mackenzie IR, Capell A, Schmid B, Neumann M, Haass C (2010) ALS-associated fused in sarcoma (FUS) mutations disrupt Transportin-mediated nuclear import. EMBO J 29:2841–2857
Dormann D, Madl T, Valori CF, Bentmann E, Tahirovic S, Abou-Ajram C, Kremmer E, Ansorge O, Mackenzie IR, Neumann M, Haass C (2012) Arginine methylation next to the PY-NLS modulates Transportin binding and nuclear import of FUS. EMBO J 31:4258–4275
Kato M, Han TW, Xie S, Shi K, Du X, Wu LC, Mirzaei H, Goldsmith EJ, Longgood J, Pei J, Grishin NV, Frantz DE, Schneider JW, Chen S, Li L, Sawaya MR, Eisenberg D, Tycko R, McKnight SL (2012) Cell-free formation of RNA granules: low complexity sequence domains form dynamic fibers within hydrogels. Cell 149:753–767
Kwon I, Kato M, Xiang S, Wu L, Theodoropoulos P, Mirzaei H, Han T, Xie S, Corden JL, McKnight SL (2013) Phosphorylation-regulated binding of RNA polymerase II to fibrous polymers of low-complexity domains. Cell 155:1049–1060
Mackenzie IR, Rademakers R, Neumann M (2010) TDP-43 and FUS in amyotrophic lateral sclerosis and frontotemporal dementia. Lancet Neurol 9:995–1007
Du K, Arai S, Kawamura T, Matsushita A, Kurokawa R (2011) TLS and PRMT1 synergistically coactivate transcription at the survivin promoter through TLS arginine methylation. Biochem Biophys Res Commun 404:991–996
Tradewell ML, Yu Z, Tibshirani M, Boulanger MC, Durham HD, Richard S (2012) Arginine methylation by PRMT1 regulates nuclear-cytoplasmic localization and toxicity of FUS/TLS harbouring ALS-linked mutations. Hum Mol Genet 21:136–149
Yamaguchi A, Kitajo K (2012) The effect of PRMT1-mediated arginine methylation on the subcellular localization, stress granules, and detergent-insoluble aggregates of FUS/TLS. PLoS One 7:e49267
Scaramuzzino C, Monaghan J, Milioto C, Lanson NA Jr, Maltare A, Aggarwal T, Casci I, Fackelmayer FO, Pennuto M, Pandey UB (2013) Protein arginine methyltransferase 1 and 8 interact with FUS to modify its sub-cellular distribution and toxicity in vitro and in vivo. PLoS One 8:e61576
Lee HW, Kim S, Paik WK (1977) S-adenosylmethionine: protein-arginine methyltransferase. Purification and mechanism of the enzyme. Biochemistry 16:78–85
Nicholson TB, Chen T, Richard S (2009) The physiological and pathophysiological role of PRMT1-mediated protein arginine methylation. Pharmacol Res 60:466–474
Bedford MT, Clarke SG (2009) Protein arginine methylation in mammals: who, what, and why. Mol Cell 33:1–13
Cheng D, Yadav N, King RW, Swanson MS, Weinstein EJ, Bedford MT (2004) Small molecule regulators of protein arginine methyltransferases. J Biol Chem 279:23892–23899
Wang J, Chen L, Sinha SH, Liang Z, Chai H, Muniyan S, Chou YW, Yang C, Yan L, Feng Y, Li KK, Lin MF, Jiang H, Zheng YG, Luo C (2012) Pharmacophore-based virtual screening and biological evaluation of small molecule inhibitors for protein arginine methylation. J Med Chem 55:7978–7987
Williams-Ashman HG, Seidenfeld J, Galletti P (1982) Trends in the biochemical pharmacology of 5′-deoxy-5′-methylthioadenosine. Biochem Pharmacol 31:277–288
Osborne TC, Obianyo O, Zhang X, Cheng X, Thompson PR (2007) Protein arginine methyltransferase 1: positively charged residues in substrate peptides distal to the site of methylation are important for substrate binding and catalysis. Biochemistry 46:13370–13381
Bonham K, Hemmers S, Lim YH, Hill DM, Finn MG, Mowen KA (2010) Effects of a novel arginine methyltransferase inhibitor on T-helper cell cytokine production. FEBS J 277:2096–2108
Endo M, Ohashi K, Mizuno K (2007) LIM kinase and slingshot are critical for neurite extension. J Biol Chem 282:13692–13702
Koga S, Kojima S, Kishimoto T, Kuwabara S, Yamaguchi A (2012) Over-expression of map kinase phosphatase-1 (MKP-1) suppresses neuronal death through regulating JNK signaling in hypoxia/re-oxygenation. Brain Res 1436:137–146
Chen Y, Zhang L, Jones KA (2011) SKIP counteracts p53-mediated apoptosis via selective regulation of p21Cip1 mRNA splicing. Genes Dev 25:701–716
Conte A, Lattante S, Zollino M, Marangi G, Luigetti M, Del Grande A, Servidei S, Trombetta F, Sabatelli M (2012) P525L FUS mutation is consistently associated with a severe form of juvenile amyotrophic lateral sclerosis. Neuromuscul Disord 22:73–75
Bosco DA, Lemay N, Ko HK, Zhou H, Burke C, Kwiatkowski TJ Jr, Sapp P, McKenna-Yasek D, Brown RH Jr, Hayward LJ (2010) Mutant FUS proteins that cause amyotrophic lateral sclerosis incorporate into stress granules. Hum Mol Genet 19:4160–4175
Takanashi K, Yamaguchi A (2014) Aggregation of ALS-linked FUS mutant sequesters RNA binding proteins and impairs RNA granules formation. Biochem Biophys Res Commun 452:600–607
Boisvert FM, Cote J, Boulanger MC, Cleroux P, Bachand F, Autexier C, Richard S (2002) Symmetrical dimethylarginine methylation is required for the localization of SMN in Cajal bodies and pre-mRNA splicing. J Cell Biol 159:957–969
Gal J, Zhang J, Kwinter DM, Zhai J, Jia H, Jia J, Zhu H (2011) Nuclear localization sequence of FUS and induction of stress granules by ALS mutants. Neurobiol Aging 32(12):2323.e27-40
Buratti E, Baralle FE (2012) TDP-43: gumming up neurons through protein-protein and protein-RNA interactions. Trends Biochem Sci 37:237–247
Zhou Y, Liu S, Liu G, Ozturk A, Hicks GG (2013) ALS-associated FUS mutations result in compromised FUS alternative splicing and autoregulation. PLoS Genet 9:e1003895
Schwartz JC, Podell ER, Han SS, Berry JD, Eggan KC, Cech TR (2014) FUS is sequestered in nuclear aggregates in ALS patient fibroblasts. Mol Biol Cell 25:2571–2578
Acknowledgments
This work was supported by research grants from the Japan Society for the Promotion of Science (#25461267) and Japan ALS Association (JALSA).
Author information
Authors and Affiliations
Corresponding author
Ethics declarations
Conflict of interest
The authors declare that they have no competing interests.
Electronic supplementary material
Below is the link to the electronic supplementary material.
11064_2015_1758_MOESM1_ESM.mp4
HEK293 cells, transfected with GFP-FUS P525L plasmid, were treated with 60 μM AdOx for 24 h initiating at 12 h after transfection. Then cells were fixed, stained with DAPI, and observed under confocal laser scanning microscopy (Olympus FV10i). 3D animation of the nucleus was reconstructed using Olympus FV10i software (MP4 675 kb)
Supplementary Fig. 1
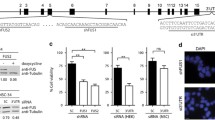
(A) HEK293 or SH-SY5Y cells, transfected with plasmid encoding HA-tagged FUS wt, FUS R521G, FUS P525L, or FUS dC, were fixed at 24 after transfection. Cells were immunostained with anti-HA antibody, and then incubated with DAPI. Images were obtained using fluorescence microscopy. Bars, 5 μm. (B) HEK293 cells, transfected with plasmids encoding GFP-FUS wt or each mutant, were lysed for Western blot analyses with anti-FUS and -Actin antibody at 24 h after transfection (upper panels). HEK293 cells, transfected with plasmid encoding GFP-FUS P525L, were lysed for Western blot analyses with anti-FUS and –Actin antibody at the indicated time points (lower panels). The bar graph indicates the relative fold of the band intensity for the ratio of ectopic GFP-FUS/endogenous FUS protein (n = 3, mean ± SE). (C) HEK293 cells, transiently transfected with GFP-FUS wt or HA-FUS wt plasmid, were treated with 60 μM AdOx for 24 h and then fixed with DAPI. Bars, 2 μm. Arrows indicate the intranuclear aggregates (PDF 506 kb)
Supplementary Fig. 2
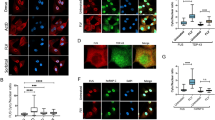
(A) (B) HEK293 cells, transiently transfected with GFP-control (A) or HA-tagged FUS P525L (B), were treated with (+) or without (−) 20 μM AdOx for 24 h initiating at 12 h after transfection. Cells were fixed and then stained with DAPI. Images were obtained using fluorescence microscopy. Bars, 5 μm. (C) HEK293 and SH-SY5Y cells, transiently transfected with GFP-FUS P525L plasmid, were treated with AMI-1(0, 100, 400 μM) for 24 h initiating at 12 h after transfection. Cells were fixed and then stained with DAPI. Bars, 5 μm. (D) HEK293 cells were treated with Mock, 5 or 20 μM AdOx, or 800 μM MTA for 24 h (upper panel). HEK293 cells were also treated with Mock, 20 μM AdOx or 400 μM AMI-1for 24 h (lower panel). Then FUS proteins, obtained by the immunoprecipitation with anti-FUS antibody in lysates of HEK293 cells, were then resolved by SDS-PAGE. The asymmetric arginine-methylation levels were measured by the immunoblot with anti-dimethyl-Arginine, asymmetric (ASYM24) Antibody (Millipore, #07-414). IgG HC, IgG heavy chain. “Relative ratio” means the relative levels of methylation compared with those of Mock sample (value = 1) (PDF 276 kb)
Rights and permissions
About this article
Cite this article
Fujii, S., Takanashi, K., Kitajo, K. et al. Treatment with a Global Methyltransferase Inhibitor Induces the Intranuclear Aggregation of ALS-Linked FUS Mutant In Vitro. Neurochem Res 41, 826–835 (2016). https://doi.org/10.1007/s11064-015-1758-z
Received:
Revised:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s11064-015-1758-z