Abstract
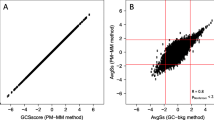
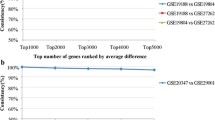
In gene expression analyses using a high-density oligonucleotide array in a rat ischemia model, two comparison methods, “pair-wise comparison” and “sample average comparison”, were evaluated based on statistical methods. The reliability of the elements screened with a 1.2 to 10-fold threshold was also evaluated. In pair-wise comparisons, most of the elements were significantly independent of the threshold value, with the percentage of significant elements remaining above 95%, when screened at 2.5-fold or higher threshold value. Pair-wise comparison structurally provided strict screening, which resulted in genes that were not selected despite significant alterations in expression. Screening by “sample average comparison” resulted in elements with low probability of significance, which suggested the necessity for increasing the reliability by additional statistical analyses after screening. When genes with altered expression were screened using an oligonucleotide array, marked differences in the numbers and reliability were proved to exist among elements screened by each sample comparison method.


Similar content being viewed by others
References
Lipshutz RJ, Fodor SP, Gingeras TR, Lockhart DJ (1999) High density synthetic oligonucleotide arrays. Nat Genet 21(1 Suppl):20–24
Kohane IS, Kho AT, Butte AJ (2003). Microarray Measurements to Analyses. In: Kohane IS, Kho AT, Butte AJ (eds) Microarrays for an integrative genomics. The MIT Press, London, PP 69–148
Mariani TJ, Budhraja V, Mecham BH, Gu CC, Watson MA, Sadovsky YA (2003) Variable fold change threshold determines significance for expression microarrays. FASEB J 17:321–323
Yang, IV, Chen E, Hasseman JP, Liang W, Frank BC, Wang S, Sharov V, Saeed AI, White J, Li J, Lee NH, Yeatman TJ, Quackenbush J (2002) Within the fold: assessing differential expression measures and reproducibility in microarray assays. Genome Biol 24:3, research 0062
Mutch DM, Berger A, Mansourian R, Rytz A, Roberts MA (2002) The limit fold change model: a practical approach for selecting differentially expressed genes from microarray data. BMC Bioinformatics 3:17
Jin W, Riley RM, Wolfinger RD, White KP, Passador-Gurgel G, Gibson G (2001) The contributions of sex, genotype and age to transcriptional variance in Drosophila melanogaster. Nat Genet 29:389–395
Yang P, Agapova O, Parker A, Shannon W, Pecen P, Duncan J, Salvador-Silva M, Hernandez MR (2004) DNA microarray analysis of gene expression in human optic nerve head astrocytes in response to hydrostatic pressure. Physiol Genomics 17:157–169
Jin K, Mao XO, Eshoo MW, Nagayama T, Minami M, Simon RP, Greenberg DA (2001) Microarray analysis of hippocampal gene expression in global cerebral ischemia. Ann Neurol 50:93–103
Lu A, Tang Y, Ran R, Clark JF, Aronow BJ, Sharp FR (2003) Genomics of the periinfarction cortex after focal cerebral ischemia. J Cereb Blood Flow Metab 23:786–810
Sergeev P, da Silva R, Lucchinetti E, Zaugg K, Pasch T, Schaub MC, Zaugg M (2004) Trigger-dependent gene expression profiles in cardiac preconditioning: evidence for distinct genetic programs in ischemic and anesthetic preconditioning. Anesthesiology 100:474–488
Raghavendra Rao VL, Bowen KK, Dhodda VK, Song G, Franklin JL, Gavva NR, Dempsey RJ (2002) Gene expression analysis of spontaneously hypertensive rat cerebral cortex following transient focal cerebral ischemia. J Neurochem 83:1072–1086
Iacobuzio-Donahue CA, Ashfaq R, Maitra A, Adsay NV, Shen-Ong GL, Berg K, Hollingsworth MA, Cameron JL, Yeo CJ, Kern SE, Goggins. M, Hruban RH (2003) Highly expressed genes in pancreatic ductal adenocarcinomas: a comprehensive characterization and comparison of the transcription profiles obtained from three major technologies. Cancer Res 63:8614–8622
Nagata T, Takahashi Y, Sugahara M, Murata A, Nishida Y, Ishikawa K, Asai S (2004) Profiling of genes associated with transcriptional responses in mouse hippocampus after transient forebrain ischemia using high-density oligonucleotide DNA array. Brain Res Mol Brain Res 121:1–11
Song G, Cechvala C, Resnick DK, Dempsey RJ, Rao VL (2001) GeneChip analysis after acute spinal cord injury in rat. J Neurochem 79:804–815
Tang Y, Lu A, Aronow BJ, Sharp FR (2001) Blood genomic responses differ after stroke, seizures, hypoglycemia, and hypoxia: blood genomic fingerprints of disease. Ann Neurol 50:699–707
Kobayashi MS, Takahashi Y, Nagata T, Nishida Y, Murata A, Ishikawa K, Asai S (2004) Screening for control genes in rat global cerebral ischemia using high-density oligonucleotide array. J Neurosci Res 76:512–518
Asai S, Zhao H, Kohno T, Takahashi Y, Nagata T, Ishikawa K (2000) Quantitative evaluation of extracellular glutamate concentration in postischemic glutamate re-uptake, dependent on brain temperature, in the rat following severe global brain ischemia. Brain Res 864:60–68
Geiss GK, Bumgarner RE, An MC, Agy MB, van ‘t Wout AB, Hammersmark E, Carter VS, Upchurch D, Mullins JI, Katze MG (2000) Large-scale monitoring of host cell gene expression during HIV-1 infection using cDNA microarrays. Virology 266:8–16
Incyte Pharmaceuticals Inc (1999) GEM microarray reproducibility study. Technical Report PN 99–0169
Nishida Y, Nagata T, Takahashi Y, Sugahara-Kobayashi M, Murata A, Asai S (2004) Alteration of serum/glucocorticoid regulated kinase-1 (sgk-1) gene expression in rat hippocampus after transient global ischemia. Brain Res Mol Brain Res 123(1–2):121–125
Acknowledgements
This work was supported by a Nihon University Research Grant for Assistants and Young Researchers (2004) (no. 04–038), a Nihon University Joint Research Grant for 2005 (no. 05-016), and by grants from the Ministry of Education, Culture, Sports, Science, and Technology of Japan to promote advanced scientific research.
Author information
Authors and Affiliations
Corresponding author
Rights and permissions
About this article
Cite this article
Kobayashi, M.S., Takahashi, Y., Nagata, T. et al. Statistical Validation of Two Sample Comparison Methods for Oligonucleotide Microarray in Rat Ischemia Model. Neurochem Res 31, 735–740 (2006). https://doi.org/10.1007/s11064-006-9074-2
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s11064-006-9074-2