Abstract
Purpose
Immunotherapy has gained traction in the treatment of solid tumors but the immunological landscape of pituitary adenomas is not well defined. We sought to investigate the immunological composition in pituitary adenomas using RNA deconvolution (CIBERSORTx) on an existing gene expression dataset for pituitary adenomas.
Methods
We applied an established computational approach (CIBERSORTx) on 134 pituitary adenomas from a previously published gene expression dataset to infer the proportions of 22 subsets of immune cells. We investigated associations between each immune cell type and tumor subtype.
Results
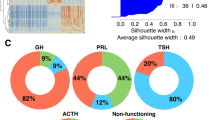
We found that the majority of infiltrating immune cells within pituitary adenomas were comprised of M2 macrophages followed by resting CD4+ memory T cells and mast cells. Silent pituitary tumors have higher M2 macrophage fractions when compared to other subtypes. In contrast, Cushing pituitary tumors, both overt and subclinical cases, had higher CD8+ T cells fractions than GH tumors, prolactinomas, hyperthyroid tumors, and silent tumors.
Conclusions
RNA deconvolution of the immune infiltrates of pituitary adenomas using CIBERSORTx suggests that most pituitary adenomas comprise of M2 macrophages, but each adenoma subtype has a unique immune landscape. This may have implications in targeting each adenoma subtype with different immunotherapies.

Similar content being viewed by others
References
Ostrom QT, Gittleman H, Xu J, Kromer C, Wolinsky Y, Kruchko C, Barnholtz-Sloan JS (2016) CBTRUS statistical report: primary brain and other central nervous system tumors diagnosed in the United States in 2009–2013. Neuro Oncol 18(5):1–75. https://doi.org/10.1093/neuonc/now207
Chen L, Han X (2015) Anti-PD-1/PD-L1 therapy of human cancer: past, present, and future. J Clin Invest 125(9):3384–3391. https://doi.org/10.1172/JCI80011
Lu JQ, Adam B, Jack AS, Lam A, Broad RW, Chik CL (2015) Immune cell infiltrates in pituitary adenomas: more macrophages in larger adenomas and more T cells in growth hormone adenomas. Endocr Pathol 26(3):263–272. https://doi.org/10.1007/s12022-015-9383-6
Mei Y, Bi WL, Greenwald NF, Du Z, Agar NY, Kaiser UB, Woodmansee WW, Reardon DA, Freeman GJ, Fecci PE, Laws ER Jr, Santagata S, Dunn GP, Dunn IF (2016) Increased expression of programmed death ligand 1 (PD-L1) in human pituitary tumors. Oncotarget 7(47):76565–76576. https://doi.org/10.18632/oncotarget.12088
Newman AM, Liu CL, Green MR, Gentles AJ, Feng W, Xu Y, Hoang CD, Diehn M, Alizadeh AA (2015) Robust enumeration of cell subsets from tissue expression profiles. Nat Methods 12(5):453–457. https://doi.org/10.1038/nmeth.3337
Athar A, Fullgrabe A, George N, Iqbal H, Huerta L, Ali A, Snow C, Fonseca NA, Petryszak R, Papatheodorou I, Sarkans U, Brazma A (2019) ArrayExpress update—from bulk to single-cell expression data. Nucleic Acids Res 47(D1):D711–D715. https://doi.org/10.1093/nar/gky964
Neou M, Villa C, Armignacco R, Jouinot A, RaffinSanson ML, Septier A, Letourneur F, Diry S, Diedisheim M, Izac B, Gaspar C, Perlemoine K, Verjus V, Bernier M, Boulin A, Emile JF, Bertagna X, Jaffrezic F, Laloe D, Baussart B, Bertherat J, Gaillard S, Assie G (2020) Pangenomic classification of pituitary neuroendocrine tumors. Cancer Cell 37(1):123–134. https://doi.org/10.1016/j.ccell.2019.11.002
Marques P, Barry S, Carlsen E, Collier D, Ronaldson A, Awad S, Dorward N, Grieve J, Mendoza N, Muquit S, Grossman AB, Balkwill F, Korbonits M (2019) Chemokines modulate the tumour microenvironment in pituitary neuroendocrine tumours. Acta Neuropathol Commun 7(1):172. https://doi.org/10.1186/s40478-019-0830-3
Sato M, Tamura R, Tamura H, Mase T, Kosugi K, Morimoto Y, Yoshida K, Toda M (2019) Analysis of Tumor Angiogenesis and Immune Microenvironment in NonFunctional Pituitary Endocrine Tumors. J Clin Med 8(5):695. https://doi.org/10.3390/jcm8050695
Yagnik G, Rutowski MJ, Shah SS, Aghi MK (2019) Stratifying nonfunctional pituitary adenomas into two groups distinguished by macrophage subtypes. Oncotarget 10(22):2212–2223. https://doi.org/10.18632/oncotarget.26775
Barry S, Carlsen E, Marques P, Stiles CE, Gadaleta E, Berney DM, Roncaroli F, Chelala C, Solomou A, Herincs M, Caimari F, Grossman AB, Crnogorac-Jurcevic T, Haworth O, Gaston-Massuet C, Korbonits M (2019) Tumor microenvironment defines the invasive phenotype of AIP-mutation-positive pituitary tumors. Oncogene 38(27):5381–5395. https://doi.org/10.1038/s41388-019-0779-5
Kemeny HR, Elsamadicy AA, Farber SH, Champion CD, Lorrey SJ, Chongsathidkiet P, Woroniecka KI, Cui X, Shen SH, Rhodin KE, Tsvankin V, Everitt J, Sanchez-Perez L, Healy P, McLendon RE, Codd PJ, Dunn IF, Fecci PE (2020) Targeting PD-L1 initiates effective antitumor immunity in a murine model of cushing disease. Clin Cancer Res 26(5):1141–1151. https://doi.org/10.1158/1078-0432.CCR-18-3486
Wang PF, Wang TJ, Yang YK, Yao K, Li Z, Li YM, Yan CX (2018) The expression profile of PD-L1 and CD8(+) lymphocyte in pituitary adenomas indicating for immunotherapy. J Neurooncol 139(1):89–95. https://doi.org/10.1007/s11060-018-2844-2
Zhang Y, Chen L (2016) Classification of advanced human cancers based on tumor immunity in the microenvironment (TIME) for cancer immunotherapy. JAMA Oncol 2(11):1403–1404. https://doi.org/10.1001/jamaoncol.2016.2450
Chen B, Khodadoust MS, Liu CL, Newman AM, Alizadeh AA (2018) Profiling tumor infiltrating immune cells with CIBERSORT. Methods Mol Biol 1711:243–259. https://doi.org/10.1007/978-1-4939-7493-1_12
Rohr-Udilova N, Klinglmuller F, Schulte-Hermann R, Stift J, Herac M, Salzmann M, Finotello F, Timelthaler G, Oberhuber G, Pinter M, Reiberger T, Jensen-Jarolim E, Eferl R, Trauner M (2018) Deviations of the immune cell landscape between healthy liver and hepatocellular carcinoma. Sci Rep 8(1):6220. https://doi.org/10.1038/s41598-018-24437-5
Yang S, Liu T, Cheng Y, Bai Y, Liang G (2019) Immune cell infiltration as a biomarker for the diagnosis and prognosis of digestive system cancer. Cancer Sci 110(12):3639–3649. https://doi.org/10.1111/cas.14216
Tang G, Yin W (2020) Development of an immune infiltration-related prognostic scoring system based on the genomic landscape analysis of glioblastoma multiforme. Front Oncol 10:154. https://doi.org/10.3389/fonc.2020.00154
Mony JT, Schuchert MJ (2018) Prognostic implications of heterogeneity in intra-tumoral immune composition for recurrence in early stage lung cancer. Front Immunol 9:2298. https://doi.org/10.3389/fimmu.2018.02298
Funding
None.
Author information
Authors and Affiliations
Corresponding author
Ethics declarations
Conflict of interest
None.
Ethical approval
This article does not contain any studies with human participants performed by any of the authors.
Additional information
Publisher's Note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Rights and permissions
About this article
Cite this article
Yeung, J.T., Vesely, M.D. & Miyagishima, D.F. In silico analysis of the immunological landscape of pituitary adenomas. J Neurooncol 147, 595–598 (2020). https://doi.org/10.1007/s11060-020-03476-x
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s11060-020-03476-x