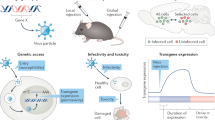
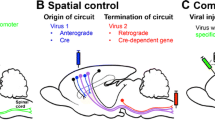
Methods based on genetically encoded molecular constructs such as antisense knockdown and RNA interference, which alter the expression of target genes, are widely used for analyzing the functions of the proteins encoded by these genes; they are also used in medical practice. Use of these methods has established, for example, that even short-term decreases in the expression of one of the noradrenaline receptors during the critical period of brain development leaves a long-lasting trace at the neurochemical and behavioral levels in subsequent life. Delivery to cells of viral constructs encoding any protein influencing cell function or small hairpin RNA (shRNA), which decrease the expression of a target gene, are also used in neurobiology. Optogenetics and chemogenetics provide clear demonstrations of the power of genetically encoded tools in studies of the central nervous system and are tools potentially suitable for controlling brain cell activity with therapeutic purposes. Both approaches are realized via expression in the target cell type of receptors novel for the organism and reacting to light of a specific wavelength or a chemical ligand molecule foreign to the organism. These approaches allow the functional consequences of changes in the activity of a specific population of neurons to be studied, and this has led to significant progress in deciphering the mechanisms of the central regulation of behavior. Thus, optogenetic studies have shown that activation of glutamatergic neurons in the dorsal hippocampus induce depression-like behavior, while the antidepressant effect of ketamine on this behavior is mediated by a direct action of this drug on NMDA receptors. Genome editing and gene expression control methods developed in recent years using bacterial CRISPR/Cas systems have already been used to study brain function. There are now optimistic expectations for the medical use of potential methods of opto- and chemogenetics, as well as CRISPR/Cas technologies, developed in model systems.
Similar content being viewed by others
References
R. Sprengel and M. T. Hasan, “Tetracycline-controlled genetic switches,” Handb. Exp. Pharmacol., 178, 49–72 (2007).
J. A. Harris, K. E. Hirokawa, S. A. Sorensen, et al., “Anatomical characterization of Cre driver mice for neural circuit mapping and manipulation,” Front. Neural Circuits, 8, 76 (2014).
K. M. Schoch and T. M. Miller, “Antisense oligonucleotides: translation from mouse models to human neurodegenerative diseases,” Neuron, 94, No. 6, 1056–1070 (2017).
R. L. Juliano, “The delivery of therapeutic oligonucleotides,” Nucleic Acids Res., 44, No. 14, 6518–6548 (2016).
N. N. Dygalo, T. S. Kalinina, and D. A. Lanshakov, “Translocation of oligonucleotide-oligosaccharide complexes into cells of the brain,” Dokl. Biochem. Biophys., 479, No. 1, 108–110 (2018).
A. W. Cwetsch, B. Pinto, A. Savardi, and L. Cancedda, “In vivo methods for acute modulation of gene expression in the central nervous system,” Prog. Neurobiol., 168, 69–85 (2018).
H. Tabata and K. Nakajima, “Efficient in utero gene transfer system to the developing mouse brain using electroporation: Visualization of neuronal migration in the developing cortex,” Neuroscience, 103, No. 4, 865–872 (2001).
S. Palfi, J. M. Gurruchaga, H. Lepetit, et al., “Long-term follow-up of a phase i/ii study of prosavin, a lentiviral vector gene therapy for Parkinson’s disease,” Hum. Gene Ther. Clin. Dev., 29, No. 3, 148–155 (2018).
N. N. Dygalo, T. S. Kalinina, and G. T. Shishkina, “Biological efficacy of antisense oligonucleotides complementary to over-lapping regions of the mRNA target,” Russ. Chem. Bull. (intern. ed.), 51, No. 7, 1031–1034 (2002).
N. K. Sahu, G. Shilakari, A. Nayak, and D. V. Kohli, “Antisense technology: A selective tool for gene expression regulation and gene targeting,” Curr. Pharm. Biotechnol., 8, No. 5, 291–304 (2007).
G. T. Shishkina, T. S. Kalinina, N. Y. Sournina, and N. N. Dygalo, “Effects of antisense to the (alpha)2A-adrenoceptors administered into the region of the locus ceruleus on behaviors in plus-maze and sexual behavior tests in sham-operated and castrated male rats,” J. Neurosci., 21, No. 2, 726–731 (2001).
R. S. Finkel, C. A. Chiriboga, J. Vajsar, et al., “Treatment of infantile- onset spinal muscular atrophy with nusinersen: A phase 2, open-label, dose-escalation study,” Lancet, 388, No. 10063, 3017–3026 (2016).
E. Mercuri, B. T. Darras, C. A. Chiriboga, et al., and the CHERISH Study Group, “Nusinersen versus sham control in later-onset spinal muscular atrophy,” New Engl. J. Med., 378, No. 7, 625–635 (2018).
S. M. Elbashir, J. Harborth, W. Lendeckel, et al., “Duplexes of 21-nucleotide RNAs mediate RNA interference in cultured mammalian cells,” Nature, 411, No. 6836, 494–498 (2001).
A. Reynolds, D. Leake, Q. Boese, et al., “Rational siRNA design for RNA interference,” Nat. Biotechnol., 22, No. 3, 326–330 (2004).
G. T. Shishkina, T. S. Kalinina, and N. N. Dygalo, “Attenuation of alpha2A-adrenergic receptor expression in neonatal rat brain by RNA interference or antisense oligonucleotide reduced anxiety in adulthood,” Neuroscience, 129, No. 3, 521–528 (2004).
N. N. Dygalo, T. S. Kalinina, and G. T. Shishkina, “Neonatal programming of rat behavior by downregulation of alpha2A-adrenoreceptor gene expression in the brain,” Ann. N. Y. Acad. Sci., 1148, 409–414 (2008).
E. Wyszko, K. Rolle, S. Nowak, et al., “A multivariate analysis of patients with brain tumors treated with ATN-RNA,” Acta Pol. Pharm., 65, No. 6, 677–684 (2008).
D. Adams, A. Gonzalez-Duarte, W. D. O’Riordan, et al., “Patisiran, an RNAi therapeutic, for hereditary transthyretin amyloidosis,” New Engl. J. Med., 379, No. 1, 11–21 (2018).
E. Sarno and A. J. Robison, “Emerging role of viral vectors for circuit- specific gene interrogation and manipulation in rodent brain,” Pharmacol. Biochem. Behav., 174, 2–8 (2018).
D. M. McCarty, S. M. Young, Jr., and R. J. Samulski, “Integration of adeno-associated virus (AAV) and recombinant AAV vectors,” Annu. Rev. Genet., 38, 819–845 (2004).
K. M. Tye and K. Deisseroth, “Optogenetic investigation of neural circuits underlying brain disease in animal models,” Nature Rev. Neurosci., 13, 251–266 (2012).
M. M. M. Verheij, C. Contet, P. Karel, et al., “Median and dorsal raphe serotonergic neurons control moderate versus compulsive cocaine intake,” Biol. Psychiatry, 83, 1024–1035 (2018).
S. Gong, M. Doughty, C. R. Harbaugh, et al., “Targeting Cre recombinase to specific neuron populations with bacterial artificial chromosome constructs,” J. Neurosci., 27, No. 37, 9817–9823 (2007).
M. S. Song and J. J. Rossi, “Molecular mechanisms of Dicer: Endonuclease and enzymatic activity,” Biochem. J., 474, No. 10, 1603–1618 (2017).
C. G. Toro and C. Mueller, “Design of shRNA and miRNA for delivery to the CNS,” Methods Mol. Biol., 1382, 67–80 (2016).
K. Albert, M. H. Voutilainen, A. Domanskyi, and M. Airavaara, “AAV vector-mediated gene delivery to substantia nigra dopamine neurons: Implications for gene therapy and disease models,” Genes (Basel), 8, No. 2, 63 (2017).
A. Chtarto, O. Bockstael, T. Tshibangu, et al., “A next step in adeno-associated virus-mediated gene therapy for neurological diseases: Regulation and targeting,” Br. J. Clin. Pharmacol., 76, No. 2, 217–232 (2013).
A. T. Das, L. Tenenbaum, and B. Berkhout, “Tet-on systems for doxycycline- inducible gene expression,” Curr. Gene Ther., 16, No. 3, 156–167 (2016).
G. T. Shishkina, D. A. Lanshakov, A. V. Bannova, et al., “Doxycycline used for control of transgene expression has its own effects on behaviors and Bcl-xl in the rat hippocampus,” Cell. Mol. Neurobiol., 38, No. 1, 281–288 (2018).
J. Nishiyama, “Genome editing in the mammalian brain using the CRISPR-Cas system,” Neurosci. Res., 141, 4–12 (2019).
E. Senis, C. Fatouros, S. Grosse, et al., “CRISPR/Cas9-mediated genome engineering: An adeno-associated viral (AAV) vector toolbox,” Biotechnol. J., 9, 1402–1412 (2014).
L. Swiech, M. Heidenreich, A. Banerjee, et al., “In vivo interrogation of gene function in the mammalian brain using CRISPR-Cas9,” Nat. Biotechnol., 33, 102–106 (2015).
J. D. Sander and J. K. Jong, “CRISPR-Cas systems for editing, regulating and targeting genomes,” Nat. Biotechnol., 32, 347–355 (2014).
C. Straub, A. J. Granger, J. L. Saulnier, and B. L. Sabatini, “CRISPR/ Cas9-mediated gene knock-down in post-mitotic neurons,” PLoS One, 9, e105584 (2014).
A. Chavez, J. Scheiman, S. Vora, et al., “Highly efficient Cas9- mediated transcriptional programming,” Nat. Methods, 12, 326–32 (2015).
C. H. Lau, J. W. Ho, P. K. Lo, and C. Tin, “Targeted transgene activation in the brain tissue by systemic delivery of engineered AAV1 expressing CRISPRa,” Mol. Ther. Nucl. Acids, 16, 637–649 (2019).
K. E. Savell and J. J. Day, “Applications of CRISPR/Cas9 in the mammalian central nervous system,” Yale J. Biol. Med., 90, No. 4, 567–581 (2017).
E. S. Boyden, F. Zhang, E. Bamberg, et al., “Millisecond timescale, genetically targeted optical control of neural activity,” Nat. Neurosci., 8, 1263–1268 (2005).
C. M. Gremel and R. M. Costa, “Orbitofrontal and striatal circuits dynamically encode the shift between goal-directed and habitual actions,” Nat. Commun., 4, 2264 (2013).
L. Fenno, O. Yizhar, and K. Deisseroth, “The development and application of optogenetics,” Annu. Rev. Neurosci., 34, 389–412 (2011).
S. M. Sternson and B. L. Roth, “Chemogenetic tools to interrogate brain functions,” Annu. Rev. Neurosci., 37, 387–407 (2014).
D. J. Urban and B. L. Roth, “DREADDs (designer receptors exclusively activated by designer drugs, chemogenetic tools with therapeutic utility,” Annu. Rev. Pharmacol. Toxicol., 55, 399–417 (2015).
N. N. Dygalo and G. T. Shishkina, “Optogenetic approach in investigations of pathophysiology and therapy of depression,” Zh. Vyssh. Nerv. Deyat., 67, No. 5, 32–40 (2017).
N. N. Dygalo, D. A. Lanshakov, U. S. Drozd, et al., “Optogenetic activation of the CA1 hippocampal pyramidal neurons induces a depressive-like behavioral phenotype,” Eur. Neuropsychopharmacol., 26, Supplement 2, S277–S278 (2016).
D. A. Lanshakov, U. S. Drozd, and N. N. Dygalo, “Optogenetic stimulation increases level of antiapoptotic protein Bcl-xL in neurons,” Biochemistry (Mosc.), 82, No. 3, 340–344 (2017).
S. M. Sternson and B. L. Roth, “Chemogenetic tools to interrogate brain functions,” Annu. Rev. Neurosci., 37, 387–407 (2014).
E. J. Campbell and N. J. Marchant, “The use of chemogenetics in behavioural neuroscience: Receptor variants, targeting approaches and caveats,” Br. J. Pharmacol., 175, No. 7, 994–1003 (2018).
S. Yun, R. P. Reynolds, I. Petrof, et al., “Stimulation of entorhinal cortex-dentate gyrus circuitry is antidepressive,” Nat. Med., 24, No. 5, 658–666 (2018).
R. S. Duman, G. Sanacora, and J. H. Krystal, “Altered connectivity in depression: GABA and glutamate neurotransmitter deficits and reversal by novel treatments,” Neuron, 102, No. 1, 75–90 (2019).
Author information
Authors and Affiliations
Corresponding author
Additional information
Translated from Rossiiskii Fiziologicheskii Zhurnal imeni I. M. Sechenova, Vol. 105, No. 11, pp. 1381–1391, November, 2019.
Rights and permissions
About this article
Cite this article
Dygalo, N.N. Investigation of Brain Functions Using Genetically Encoded Tools. Neurosci Behav Physi 50, 1051–1056 (2020). https://doi.org/10.1007/s11055-020-01004-y
Received:
Revised:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s11055-020-01004-y