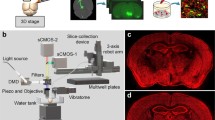
This review considers the current methods required for functional mapping of neural networks in 3D space with cellular resolution. Cyclic immunohistochemistry approaches and immunohistochemical staining methods for thick nervous tissue preparations are described in detail, along with approaches to clearing large brain formations to allow signals to be visualized at great depth. The main advantages and disadvantages of their use are considered. Perspectives for the development of molecular mapping for studies of the neural substrates of cognitive functions in health and disease are discussed.
Similar content being viewed by others
References
Anokhin, K. V., “Molecular scenarios for the consolidation of long-term memory,” Zh. Vyssh. Nerv. Deyat., 47, No. 2, 261–279 (1997).
Aoyagi, Y., Kawakami, R., Osanai, H., et al., “A rapid optical clearing protocol using 2,2’-thiodiethanol for microscopic observation of fixed mouse brain,” PLoS One, 10, No. 1, e0116280 (2015).
Barth, A. L., “Visualizing circuits and systems using transgenic reporters of neural activity,” Curr. Opin. Neurobiol., 17, No. 5, 567–571 (2007).
Becker, K., Jährling, N., Saghafi, S., et al., “Chemical clearing and dehydration of GFP expressing mouse brains,” PLoS One, 7, No. 3, e33916 (2012).
Chiang, A. S., Lin, W. Y., Liu, H. P., et al., “Insect NMDA receptors mediate juvenile hormone biosynthesis,” Proc. Natl. Acad. Sci. USA, 99, No. 1, 37–42 (2002).
Chung, K., Wallace, J., Kim, S. Y., et al., “Structural and molecular interrogation of intact biological systems,” Nature, 497, No. 7449, 332–337 (2013).
Cole, A. J., Saffen, D. W., Baraban, J. M., and Worley, P. F., “Rapid increase of an immediate early gene messenger RNA in hippocampal neurons by synaptic NMDA receptor activation,” Nature, 340, No. 6233, 474–476 (1989).
Denk, W. and Horstmann, H., “Serial block-face scanning electron microscopy to reconstruct three-dimensional tissue nanostructure,” PLoS Biol., 2, No. 11, e329 (2004).
Denk, W. and Svoboda, K., “Photon upmanship: why multiphoton imaging is more than a gimmick,” Neuron, 18, No. 3, 351–357 (1997).
Denk, W., Delaney, K. R., Gelperin, A., et al., “Anatomical and functional imaging of neurons using 2-photon laser scanning microscopy,” J. Neurosci. Meth, 54, No. 2, 151–162 (1994).
Dent, J. A., Polson, A. G., and Klymkowsky, M. W., “A whole-mount immunocytochemical analysis of the expression of the intermediate filament protein vimentin in Xenopus,” Development, 105, No. 1, 61–74 (1989).
Dodt, H. U., Leischner, U., Schierloh, A., et al., “Ultramicroscopy: three-dimensional visualization of neuronal networks in the whole mouse brain,” Nat. Methods, 4, No. 4, 331–336 (2007).
Ertürk, A., Becker, K., Jährling, N., et al., “Three-dimensional imaging of solvent-cleared organs using 3DISCO,” Nat. Protoc., 7, No. 11 1983–1995 (2012).
Ertürk, A., Mauch, C. P., Hellal, F., et al., “Three-dimensional imaging of the unsectioned adult spinal cord to assess axon regeneration and glial responses after injury,” Nat. Med., 18, No. 1, 166–171 (2011).
Farivar, R., Zangenehpour, and S. Chaudhuri, A., “Cellular-resolution activity mapping of the brain using immediate-early gene expression,” Front. Biosci., 9, 104–109 (2004).
Gendusa, R., Scalia, C. R., Buscone, S., and Cattoretti, G., “Elution of high-affinity (>10–9 kD) antibodies from tissue sections: clues to the molecular mechanism And use in sequential immunostaining,” J. Histochem. Cytochem., 62, No. 7, 519–531 (2014).
Gerdes, M. J., Sevinsky, C. J., Sood, A., et al., “Highly multiplexed single-cell analysis of formalin-fixed, paraffin-embedded cancer tissue,” Proc. Natl. Acad. Sci. USA, 110, No. 29, 11982–11987 (2013).
Glass, G., Papin, J. A., and Mandell, J. W., “SIMPLE: a sequential immunoperoxidase labeling and erasing method,” J. Histochem. Cytochem., 57, No. 10, 899–905 (2009).
Gleave, J. A., Lerch, J. P., Henkelman, R. M., and Nieman, B. J., “A method for 3D immunostaining and optical imaging of the mouse brain demonstrated in neural progenitor cells,” PLoS One, 8, No. 8, e72039 (2013).
Guzowski, J. F., Timlin, J. A., Roysam, B., et al., “Mapping behaviorally relevant neural circuits with immediate-early gene expression,” Curr. Opin. Neurobiol., 15, No. 5, 599–606 (2005).
Hama, H., Kurokawa, H., Kawano, H., et al., “Scale: a chemical approach for fluorescence imaging and reconstruction of transparent mouse brain,” Nat. Neurosci., 14, No. 11, 1481–1488 (2011).
Helmchen, F. and Denk, W., “Deep tissue two-photon microscopy,” Nat. Methods, 2, No. 12, 932–940 (2005).
Helmstaedter, M., Briggman, K. L., and Denk, W., “3D structural imaging of the brain with photons and electrons,” Curr. Opin. Neurobiol., 18, No. 6, 633–641 (2008).
Hou, B., Zhang, D., Zhao, S., et al., “Scalable and DiI-compatible optical clearance of the mammalian brain,” Front. Neuroanat., 9, 19 (2015).
Jahng, J. W. and Lee, J. H., “Activation of the hypothalamic-pituitary-adrenal axis in lithium-induced conditioned taste aversion learning,” Eur. J. Pharmacol., 768, 182–188 (2015).
Ke, M. T., Fujimoto, S., and Imai, T., “SeeDB: a simple and morphology- preserving optical clearing agent for neuronal circuit reconstruction,” Nat. Neurosci., 16, No. 8, 1154–1161 (2013).
Kolodziejczyk, E. and Baertschi, A. J., “Multiple immunolabeling in histology: a new method using thermo-inactivation of immunoglobulins,” J. Histochem. Cytochem., 34, No. 12, 1725–1729 (1986).
Kuwajima, T., Sitko, A. A., Bhansali, P., et al., “ClearT: a detergent-and solvent-free clearing method for neuronal and non-neuronal tissue,” Development, 140, No. 6, 1364–1368 (2013).
Lan, H. Y., Mu, W., Nikolic-Paterson, D. J., and Atkins, C., “A novel, simple, reliable and sensitive method for multiple immunoenzyme staining: use of microwave oven heating to block antibody cross-reactivity and retrieve antigens,” J. Histochem. Cytochem., 43, No. 1, 97–102 (1995).
Lanahan, A. and Worley, P., “Immediate-early genes and synaptic function,” Neurobiol. Learn. Mem., 70, No. 1–2, 37–43 (1998).
Li, J., Zhou, Y., and Gu, J., “Stain-Decolorize-stain (SDS, a new technique for multiple staining,” Histochem. Cell Biol., 141, No. 3, 251–262 (2014).
Lichtman, J. W., Livet, J., and Sanes, J. R., “A technicolour approach to the connectome,” Nat. Rev. Neurosci., 9, No. 6, 417–422 (2008).
Lin, H. H., Lai, J. S., Chin, A. L., et al., “A map of olfactory representation in the Drosophila mushroom body,” Cell, 128, No. 6, 1205–1217 (2007).
Liu, Y. A., Chen, Y., Chiang, A. S., et al., “Optical clearing improves the imaging depth and signal-to-noise ratio for digital analysis and three-dimensional projection of the human enteric nervous system,” Neurogastroenterol. Motil., 23, No. 10, e446–457 (2011).
Liu, Y. C. and Chiang, A. S., “High-resolution confocal imaging and three-dimensional rendering,” Methods, 30, No. 1, 86–93 (2003).
Livet, J., Weissman, T. A., Kang, H., et al., “Transgenic strategies for combinatorial expression of fluorescent proteins in the nervous system,” Nature, 450, No. 7166, 56–62 (2007).
Maleeva, H. E., Ivolgina, G. L., Anokhin, K. V., and Limborskaya, S. A., “Analysis of the expression of the c-fos oncogene in the rat cerebral cortex on learning,” Genetika, 25, 1119–1121 (1989).
Micheva, K. D. and Smith, S. J., “Array tomography: a new tool for imaging the molecular architecture and ultrastructure of neural circuits,” Neuron, 55, No. 1, 25–36 (2007).
Nakane, P. K., “Simultaneous localization of multiple tissue antigens using the peroxidase-labeled antibody method: a study on pituitary glands of the rat,” J. Histochem. Cytochem., 16, No. 9, 557–560 (1968).
Okuno, H., “Regulation and function of immediate-early genes in the brain: beyond neuronal activity markers,” Neurosci. Res., 69, No. 3, 175–186 (2011).
Parra, S. G., Chia, T. H., Zinter, J. P., and Levene, M. J., “Multiphoton microscopy of cleared mouse organs,” J. Biomed. Opt., 15, No. 3, 036017 (2010).
Parra, S. G., Vesuna, S. S., Murray, T. A., and Levene, M. J., “Multiphoton microscopy of cleared mouse brain expressing YFP,” J. Vis. Exp., 67, e3848 (2012).
Pavlova, I. P., Shipley, S. C., Lanio, M., et al., “Optimization of immunolabeling and clearing techniques for indelibly labeled memory traces,” Hippocampus, 28, No. 7, 523–535 (2018).
Pirici, D., Mogoanta, L., Kumar-Singh, S., et al., “Antibody elution method for multiple immunohistochemistry on primary antibodies raised in the same species and of the same subtype,” J. Histochem. Cytochem., 57, No. 6, 567–575 (2009).
Purkyně, J. E., Commentatio de Examine Physiologico Organi Visus et Systematis Cutanei, Univ. Breslau Press, Breslau, Prussia (1823).
Renier, N., Wu, Z., Simon, D. J., et al., “iDISCO: a simple, rapid method to immunolabel large tissue samples for volume imaging,” Cell, 159, No. 4, 896–910 (2014).
Roshchina, M. A., Ivashkina, O. I., and Anokhin, K. V., “Novel approaches to cognitive neurobiology: in vivo two-photon methods for visualization of cognitively active neurons,” Zh. Vyssh. Nerv. Deyat., 62, No. 2, 141–149 (2017).
Saidov, Kh. M. and Anokhin, K. V., “Novel approaches to cognitive neurobiology: molecular labeling methods and ex vivo visualization of cognitively active neurons,” Zh. Vyssh. Nerv. Deyat., 67, No. 3, 259–272 (2017).
Sillitoe, R. V. and Hawkes, R., “Whole-mount immunohistochemistry: a high-throughput screen for patterning defects in the mouse cerebellum,” J. Histochem. Cytochem., 50, No. 2, 235–244 (2002).
Susaki, E. A., Tainaka, K., Perrin, D., et al., “Whole-brain imaging with single-Cell resolution using chemical cocktails and computational analysis,” Cell, 157, No. 3, 726–739 (2014).
Tainaka, K., Kubota, S. I., Suyama, T. Q., et al., “Whole-body imaging with single-cell resolution by tissue decolorization,” Cell, 159, No. 4, 911–24 (2014).
Theer, P. and Denk, W., “On the fundamental imaging-depth limit in two-photon microscopy,” J. Opt. Soc. Am. A. Opt. Image Sci. Vis., 23, No. 12, 3139–3149 (2006).
Tomer, R., Ye, L., Hsueh, B., and Deisseroth, K., “Advanced CLARITY for rapid and high-resolution imaging of intact tissues,” Nat. Protoc., 9, 1682–1697 (2014).
Tornehave, D., Hougaard, D. M., and Larsson, L., “Microwaving for doubl indirect immunofluorescence with primary antibodies from the same species and for staining of mouse tissues with mouse monoclonal antibodies,” Histochem. Cell Biol., 113, No. 1, 19–23 (2000).
Tóth, Z. E. and Mezey, E., “Simultaneous visualization of multiple antigens with tyramide signal amplification using antibodies from the same species,” J. Histochem. Cytochem., 55, No. 6, 545–554 (2007).
Tsai, P. S., Kaufhold, J. P., Blinder, P., et al., “Correlations of neuronal and microvascular densities in murine cortex revealed by direct counting and colocalization of nuclei and vessels,” J. Neurosci., 29, No. 46, 1453–1470 (2009).
Van den Brand, M., Hoevenaars, B. M., Sigmans, J. H., et al., “Sequential immunohistochemistry: a promising new tool for the pathology laboratory,” Histopathology, 65, No. 5, 651–657 (2014).
Wählby, C., Erlandsson, F., Bengtsson, E., and Zetterberg, A., “Sequential immunofluorescence staining and image analysis for detection of large numbers of antigens in individual cell nuclei,” Cytometry, 47, No. 1, 32–41 (2002).
Yang, B., Treweek, J. B., Kulkarni, R. P., et al., “Single-cell phenotyping within transparent intact tissue through whole-body clearing,” Cell, 158, No. 4, 945–958 (2014).
Zipfel, W. R., Williams, R. M., and Webb, W. W., “Nonlinear magic: multiphoton microscopy in the biosciences,” Nat. Biotechnol, 21, No. 11, 1369–1377 (2003).
Zukor, K. A., Kent, D. T., and Odelberg, S. J., “Fluorescent whole-mount method for visualizing three-dimensional relationships in intact and regenerating adult newt spinal cords,” Dev. Dyn., 239, No. 11, 3048–3057 (2010).
Author information
Authors and Affiliations
Corresponding author
Additional information
Translated from Zhurnal Vysshei Nervnoi Deyatel’nosti imeni I. P. Pavlova, Vol. 68, No. 6, pp. 747–758, November–December, 2018.
Rights and permissions
About this article
Cite this article
Efimova, O.I., Balaban, P.M. & Khaitovich, F.E. Novel Approaches to the Molecular Mapping of the Brain: 3D Cyclic Immunohistochemistry and Optical Clearing. Neurosci Behav Physi 50, 73–80 (2020). https://doi.org/10.1007/s11055-019-00871-4
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s11055-019-00871-4