Abstract
Background
Resolving genomic insults is essential for the survival of any species. In the case of eukaryotes, several pathways comprise the DNA damage repair network, and many components have high evolutionary conservation. These pathways ensure that DNA damage is resolved which prevents disease associated mutations from occurring in a de novo manner. In this study, we investigated the role of the Eyes Absent (EYA) homologue in Caenorhabditis elegans and its role in DNA damage repair. Current understanding of mammalian EYA1 suggests that EYA1 is recruited in response to H2AX signalling to dsDNA breaks. C. elegans do not possess a H2AX homologue, although they do possess homologues of the core DNA damage repair proteins. Due to this, we aimed to determine if eya-1 contributes to DNA damage repair independent of H2AX.
Methods and results
We used a putative null mutant for eya-1 in C. elegans and observed that absence of eya-1 results in abnormal chromosome morphology in anaphase embryos, including chromosomal bridges, missegregated chromosomes, and embryos with abnormal nuclei. Additionally, inducing different types of genomic insults, we show that eya-1 mutants are highly sensitive to induction of DNA damage, yet show little change to induced DNA replication stress and display a mortal germline resulting in sterility over successive generations.
Conclusions
Collectively, this study suggests that the EYA family of proteins may have a greater involvement in maintaining genomic integrity than previously thought and unveils novel roles of EYA associated DNA damage repair.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
Introduction
The eukaryotic genome is constantly under stress, which can lead to lesions in DNA in the form of single or double stranded breaks (DSBs) [1]. Unrepaired and unchecked DNA lesions can lead to multiple diseased states such as cancer and developmental defects. To resolve these constant insults, several pathways exist and form an extensive network of DNA damage repair mechanisms that maintain genomic integrity, or in the case of irreparable DSBs, can initiate apoptosis to prevent mutations from proliferating [2, 3]. The Eyes Absent family of proteins (EYA) are characterised by their role as transcriptional cofactors as tyrosine phosphatases and are multifunctional regarding their contribution to development [4,5,6]. Initially named due to the absence of eyes in eya mutants in Drosophila, the EYA family of proteins regulate several pathways that contribute to developmental processes [5, 7,8,9]. EYA proteins have been implemented in signalling pathways and various cellular processes and their breakdown has been associated with several diseased states [5]. Less understood is the role EYA proteins play regarding the maintenance of genomic integrity. However, studies have shown that EYA proteins are recruitment to the site of DNA DSBs by the histone variant H2AX, which functions as a scaffold for the recruitment of DNA damage repair proteins [10,11,12]. In apoptosis, EYA proteins also initiate apoptosis by interacting with the highly conserved homeobox SIX family proteins. Due to the multi-functional roles exhibited by EYA proteins, characterising their precise role is challenging.
One of the key models for exploring developmental processes is the roundworm, Caenorhabditis elegans, which has been a valuable model for deciphering highly conserved genomic protection mechanisms [13, 14]. In C. elegans EYA proteins are represented by one protein, EYA-1, which functions as a homologue for mammalian EYA1-4. Investigations into C. elegans EYA-1 have revealed its involvement in egg laying, apoptosis, and response to heat stress [15,16,17]. In mammals, EYA-1 is recruited to DSBs by H2AX which functions as a primary signal for DSB repair [12]. However, the C. elegans genome lacks a H2AX orthologue, despite possessing many of the core proteins that participate in DSB repair [13]. Due to this, any role of EYA-1 in DNA damage repair pathways in C. elegans is likely to be independent of H2AX signalling. In this study, we show that absence of eya-1 results in chromosomal abnormalities in anaphase embryos and results in a mortal germline phenotype. In addition to this, we also show that eya-1 mutants exhibit hypersensitivity to induction of DNA damage, but not to DNA replication stress. Given the absence of H2AX in C. elegans, our findings unveil potential novel contributions of EYA-1 and its role in maintaining genomic integrity.
Results and discussion
EYA-1 is highly conserved and required for normal reproduction
EYA family proteins possess an EYA domain (ED) which hosts a catalytic quintet of residues, which has previously been shown to exhibit high conservation of the EYA domain from plants, invertebrates and vertebrates [18]. The structure of EYA-1 has not been solved in either C. elegans or in higher eukaryotic species. To establish greater validity of EYA-1 as an ideal model for studying conserved features of EYA-1 homologues, structural models of C. elegans EYA-1 and homologues in D. melanogaster and H. sapiens were generated (Fig. 1). As expected, the structures of all three models exhibited similar overall features, with the notable difference of a smaller alpha-helix at the N-terminus in C. elegans compared to the N-terminus in H. sapiens and D. melanogaster. Despite this minor structural variation, the similarity suggests that C. elegans EYA-1 is a valid model for understanding roles conserved in higher eukaryotes.
Homology modelling and active site conservation of EYA-1: Homology models showing structure of C. elegans EYA-1, H. sapiens EYA2, and D. melanogaster EYA-1. N-terminus is represented as blue and C-terminus is red. Arrow indicates variance of the shorter N-terminus alpha helix in C. elegans compared to H. sapiens and D. melanogaster
Having established structural conservation of EYA-1, we next assessed the requirement of eya-1 for normal reproductive capacity. C. elegans hermaphrodites produce up to 300 progeny per animal with little genetic variability, making them an ideal model to assess the requirement of candidate proteins in reproductive tissue [14]. Therefore, we obtained an eya-1 putative null mutant eya-1(ok654), which has a large deletion of the EYA domain [15] and performed a brood size assay to determine if absence of eya-1 impedes normal reproductive capacity (Fig. 2). This was conducted at 20 °C and a less permissive temperature of 25 °C, and interestingly, brood size was significantly decreased in eya-1(ok654) mutants compared to wild-type animals at both temperatures (Fig. 2A). Moreover, eya-1(ok654) mutants displayed elevated levels of embryonic lethality (Fig. 2B). Absence of EYA proteins in Drosophila are embryonic lethal showing that eya-1(ok654) mutants show a similar, albeit less severe trend [19], although our results do differ from those previously reported which suggest that eya-1 mutants only display larval arrest [15]. Nonetheless, these findings suggest that eya-1 is required for maintaining normal reproductive capacity in C. elegans.
Reproductive capacity of eya-1 deletion mutants and mortal germline: (A) Brood size of eya-1(ok654) mutants showing decreased brood which is exacerbated at higher temperatures. *** p = < 0.0001, error bars = SEM, n = 10. (B) Percentage of embryonic lethality from the brood of eya-1(ok654) mutants. *** p = < 0.0001, error bars = SEM, n = 10. (C) Mortal germline assay represented as percentage of survival over successive generations. n = 20
eya-1 is required for embryonic chromosomal integrity and germline longevity
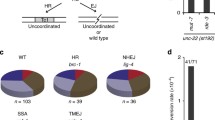
Abnormal reproductive capacity and embryonic lethality have been associated with chromosomal defects, which can be observed in anaphase stage embryos during meiosis II [20]. To investigate whether the germline and embryonic phenotypes observed in eya-1(ok654) mutants were due to any visible chromosome morphological defects, we examined embryos in the 1 to 2 cell stage of division. Strikingly, eya-1(ok654) mutants exhibit a broad range of adverse morphologies, including chromosomal bridges, embryos with missegregated chromosomes, and embryos with abnormal nuclei (Fig. 3A, B). We also assessed the germline nuclei of eya-1(ok654) mutants and interestingly found no abnormalities in pachytene nuclei or diakinetic oocytes (data not shown). However, we did observe a mild, low penetrance HIM phenotype (High Incidence of Males) at 25 °C which suggests that germline chromosomal segregation defects may be present (Fig. S1A). In addition to this, we also observed a mild increase in germ cell apoptosis when compared to wild-type animals (Fig. S1C). Nonetheless, we cannot determine if any unobservable defects are present in the preliminary stages of oocytes, or if these observed abnormal embryonic chromosomes are restricted to the anaphase stage events of meiosis II. Nonetheless, the reduced brood size of eya-1(ok654) mutants, coupled with an increase in germ cell apoptosis suggests that germline processes may be impeded in the absence of eya-1.
Abnormal chromosome morphology in eya-1(ok654) mutants: (A) Absence of eya-1 results in abnormal chromosomes in anaphase embryos. Scale bar = 5 μm (B) Percentage of embryos scored and binned based on morphology. Arrows indicate chromosomal bridges, missegregated chromosomes, and abnormal nuclei (top to bottom) n = 200. *** p = < 0.0001, error bars = SEM
To investigate this further, eya-1(ok654) mutants were subjected to a mortal brood size assay (Fig. 2C), where individual eya-1(ok654) mutants were transferred from each generation alongside wild-type animals until sterility occurred. The rationale behind is based on the accumulation of inherited germline mutations throughout generations, ultimately render the strain sterile [21, 22]. We observed a marked reduction in brood size that decreased with each generation. This trend persisted until the 10th generation, at which point eya-1(ok654) mutants were sterile, with no recorded change in the brood of wild-type animals. This suggests that eya-1(ok654) mutants accumulate germline defects that are transferred to each generation. This is interesting as eya-1 expression is typified from early embryogenesis and is most active during somatic development [15]. Unfortunately, we were unsuccessful in generating a functional EYA-1 germline antibody to investigate this further. However, it is possible that the absence of EYA-1 activity in the somatic gonad may have a generational accumulative influence of germline processes independent of eya-1 germline expression. Despite this, the absence of eya-1 in C. elegans results in sterility over successive generations, raising novel concerns regarding its influence of germ cell function and longevity.
eya-1(ok654) mutants are hypersensitive to DNA damage
The observation of embryonic lethality and abnormal anaphase chromosomes in eya-1(ok654) mutants suggests a role for eya-1 in DNA damage repair. To determine if these genomic insults were due to replication stress or DNA damage, we aimed to induce both insults in eya-1(ok654) mutants and assess the severity of each insult. Therefore, eya-1(ok654) mutants were subjected to hydroxyurea (HU) to induce replication stress and camptothecin (CPT), which induces single stranded breaks and indirectly leads to double stranded breaks [23] (Fig. 4). This was performed at both 20 °C and 25 °C, consistent with our prior results, and both temperatures showed the same trend upon exposure to each agent. HU exposure led to a minor increase in unhatched embryos in eya-1(ok654) mutants similar to the trend observed in wild-type animals (Fig. 4A). However, exposure to CPT led to a more noticeable increase in unhatched embryos (Fig. 4B). This hypersensitivity to CPT, coupled with observed embryonic chromosomal defects suggests a strong requirement of EYA-1 in single or double strand DNA break repair. To ensure that this trend was specific to eya-1 and not a background mutation we performed the same assay on wild-type animals that were subjected to RNAi of eya-1 and observed a similar trend (Fig S1B). Current understanding of EYA-1 and its role in DNA repair involves H2AX as a substrate for EYA-1 as part of the primary dsDNA break signal [11, 24]. Since C. elegans lacks H2AX, this might infer that EYA-1 has novel roles in DNA break repair beyond H2AX signalling.
Hypersensitivity of DNA damage in eya-1(ok654) mutants: (A) DNA replication stress induced with hydroxyurea (HU) showing little change from wild-type animals. n = 20, *** p = < 0.0001, error bars = SEM. (B) Hypersensitivity of eya-1(ok654) mutants to DNA breaks induced with camptothecin (CPT) with elevated embryonic lethality in eya-1(ok654) mutants. n = 20, *** p = < 0.0001, error bars = SEM. Replicated in triplicates
This correlation between EYA-1 and DNA damage repair may be more varied than first thought. Increased DNA damage in the absence of H2AX in C. elegans is a significant point of interest as it potentially adds a new role of EYA proteins in DNA repair. In mammals, after the occurrence of DSBs, EYA-1 recruits a series of proteins associated with DSB repair, and the stimulus for this association of EYA-1 is phosphorylation of H2AX. This allows EYA-1 to function as a scaffold for the core DSB machinery to assemble and facilitate DSB repair [5, 11, 24]. Although C. elegans do not possess H2AX, many of the proteins EYA-1 recruits in this process for DSB repair are conserved and therefore present in C. elegans [13]. Due to this, it is possible that EYA-1 may have a conserved role in DNA repair that goes beyond that of H2AX phosphorylation and may participate further upstream in DNA repair pathways.
EYA-1 is required for the initiation of apoptosis by its association with ceh-34 [17]. We therefore assessed germ cell apoptosis in eya-1(ok654) mutants and observed a mild increase in germ cells undergoing apoptosis (Fig. S1D, E). Due to this, we cannot rule out the possibility that this hypersensitivity to induced DNA damage in eya-1 mutants might be due to deficient induction of apoptosis as normally initiated by eya-1 and ceh-34. In this regard, it might well be that cells with DNA damage beyond repair that would otherwise undergo apoptosis might be unchecked and allowed to persist. Therefore, our observations may be due to an overaccumulation of DNA damage in cells that would otherwise be eradicated in wild-type animals. However, the specific sensitivity to induction of DNA damage in eya-1 mutants coupled with the lack of sensitivity to induction of replication stress suggests a correlation between eya-1 and single or double stranded DNA repair independent of H2AX signalling.
Conclusion
EYA proteins are multi-functional and participate in numerous pathways. Due to this, deciphering the precise role that EYA-1 plays within the network of DSB repair will enhance understanding of both EYA-1 and the overall repair process. Nonetheless, this study shows that EYA-1 likely has a role in DNA repair beyond its previously characterised role in association with H2AX signalling. Therefore, we conclude that the role of EYA-1 in resolving DNA damage may be more complex than current mechanisms assume, and deciphering its various contributions to this pathway will enhance understanding of DNA repair processes.
Materials and methods
Strains
Strains used in this study were maintained 20 and 25 °C under standard conditions [25]. The Bristol N2 strain was used for the wild-type strain and the eya-1(ok654) deletion mutant was obtained from the CGC [26] and backcrossed five times prior to performing all experiments. Genotyping of eya-1(ok654) animals was performed using the following primers. Forward: GCACGGCAAATTACGAAAGC and Reverse: GCCGTGCTTAACAAACTCCA.
Embryonic DNA analysis
Dissections were performed as described previously [27]. In short, 1-day old adult animals were isolated and anesthetized in 0.001% tetramisole in M9 buffer on 22 × 22 mm coverslips and dissected using two syringe needles. Coverslips were then placed onto microscopy slides precoated poly-L-lysine and snap frozen in liquid nitrogen then incubated in methanol for 20 min. Slides were washed twice with washing buffer (0.1% Tween-20 in PBS) for 10 min and covered with DAPI diluted in normal goat serum, then incubated at room temperature in a humidity chamber for 1 h. Slides were washed twice with wash buffer for 10 min and mounted on coverslips using Dako Fluorescent Mounting Media (Dako, Glostrup, Denmark). Slides were examined using a Zeiss Axiolab microscope with a pE-300 lite LED fluorescence adaptor (Zeiss, Oberkochen, Germany). Images were captured using a 1.4-MP (1,360 × 1,024) Tucsen 2/3 monochrome CCD camera (Tucson, Fujian, China). Anaphase embryos were binned as normal (two distinct sets of chromosomes), bridging (lagging strands of DNA between chromosomes) missegregation (chromosomal bodies not correctly separated), and abnormal (clumped or disordered yet did not clearly resemble the above descriptions). Analysis was conducted 3 times using the above protocol on wild-type and eya-1(ok654) animals with ∼ 100 L1 worms on NGM plates feeding on OP50 bacteria until reaching adulthood prior to dissection and DAPI staining. Counting of anaphase chromosomes occurred until n = 200 was achieved.
Brood and embryonic lethality
Brood and embryonic lethality counts were performed at 20 and 25 °C. Synchronized populations of each strain were grown at 20 °C until the fourth larval stage then placed onto NGM plates pre-seeded with OP50. Each worm was then transferred to new plate every 12 h and scored for progeny after 48 h. This was repeated until each worm failed to lay any new progeny. Total progeny was recorded as viable progeny and unhatched embryos. n = 10, performed in three replicates.
Induction of DNA damage
Inducing DNA damage was achieved by placing worms to plates prepared with 25 mM hydroxyurea (Sigma-Aldrich, Missouri, USA), or 50 µM camptothecin (Sigma-Aldrich, Missouri, USA) and repeated three times in triplicates as previously described [28]. In short, twenty L4 staged wild-type and eya-1 mutants were placed on NGM plates enriched with each reagent, seeded with OP50 and incubated at 20 °C and 25 °C for 16 h. Worms were then transferred to seeded NGM plates for 4 h for recovery, then transferred again to fresh seeded NGM plates. Plates with embryos were then incubated at 20–25 °C accordingly for 24 h, then scored for embryonic lethality and hatching rates.
C. elegans mortal germline assay
Mortal germline assays were conducted by placing 10 individual L1 wild-type and eya-1 mutants on seeded NGM plates at 25 °C until they laid progeny. One representative worm from the progeny of each plate were transferred to new plates until they matured and started to lay progeny. The process was then repeated until eya-1 mutants were sterile. Brood size and rates of embryonic lethality were recorded from the brood of each generation.
RNA interference
RNAi was performed by the plate feeding method and bacterial clones were sourced from the ORFeome library [29]. RNAi clones were grown in 2 x TY media plus 100 mg/mL ampicillin overnight at 37 °C on NGM plates (3% bacto-agar, 86 mM NaCl, 42 mM Na2HPO4, 22 mM KH2PO4, and 1 mM MgSO4) containing 100 mg/mL ampicillin plus 4 mM IPTG. Wild-type animals were fed an empty vector and grown alongside animals treated with RNAi. Approximately, 100 synchronized L1 wild-type animals were then transferred to seeded plates in duplicates and grown at 20 °C and 25 °C prior to analysis and performed in three separate replicates.
Acridine orange staining
Germ cells apoptosis used acridine orange as described previously [30]. In short, 20 day 1 adult worms were stained with 1 ml of 100µM acridine orange (Sigma-Aldrich, Missouri, USA) on NGM plates seeded with OP50 and for 1 h, then washed in M9 buffer and placed on 2% agarose gel pads in 0.03% tetramisole and scored using DIC and fluorescence microscopy. This was replicated for 20 °C and 25 °C in duplicates and performed in three separate replicates.
Statistical analysis and software
Generation of graphs and statistical analysis utilised the Prism 5 software package (GraphPad, San Diego, CA, USA). Statistical significance was determined using a student t-test and errors were represented as standard error of the mean. Images were processed using Adobe Photoshop, and Adobe Illustrator was used to assemble figures (Adobe, San Jose, CA, USA). Multiple sequence alignment was achieved using Clustral Omega [31] and protein homology models were generated using Swiss-Model [32].
Data availability
No datasets were generated or analysed during the current study.
References
Bournaka S et al (2024) The cell cycle revisited: DNA replication past S phase preserves genome integrity. Semin Cancer Biol 99:45–55
Chen Q, Gong B, Almasan A (2000) Distinct stages of cytochrome c release from mitochondria: evidence for a feedback amplification loop linking caspase activation to mitochondrial dysfunction in genotoxic stress induced apoptosis. Cell Death Differ 7(2):227–233
Gong B et al (1999) Ionizing radiation-induced, bax-mediated cell death is dependent on activation of cysteine and serine proteases. Cell Growth Differ 10(7):491–502
Sano T, Nagata S (2011) Characterization of the threonine-phosphatase of mouse eyes absent 3. FEBS Lett 585(17):2714–2719
Hegde RS, Roychoudhury K, Pandey RN (2020) The multi-functional eyes absent proteins. Crit Rev Biochem Mol Biol 55(4):372–385
Okabe Y, Sano T, Nagata S (2009) Regulation of the innate immune response by threonine-phosphatase of eyes absent. Nature 460(7254):520–524
Ohto H et al (1999) Cooperation of six and eya in activation of their target genes through nuclear translocation of Eya. Mol Cell Biol 19(10):6815–6824
Li X et al (2003) Eya protein phosphatase activity regulates Six1-Dach-eya transcriptional effects in mammalian organogenesis. Nature 426(6964):247–254
Soni UK, Roychoudhury K, Hegde RS (2021) The eyes absent proteins in development and in developmental disorders. Biochem Soc Trans 49(3):1397–1408
Ji JH et al (2019) De novo phosphorylation of H2AX by WSTF regulates transcription-coupled homologous recombination repair. Nucleic Acids Res 47(12):6299–6314
Cook PJ et al (2009) Tyrosine dephosphorylation of H2AX modulates apoptosis and survival decisions. Nature 458(7238):591–596
Nowsheen S et al (2018) ZNF506-dependent positive feedback loop regulates H2AX signaling after DNA damage. Nat Commun 9(1):2736
Gartner A, Engebrecht J (2022) DNA repair, recombination, and damage signaling. Genetics, 220(2)
Davis GM, Hipwell H, Boag PR (2023) Oogenesis in Caenorhabditis elegans. Sex Dev 17(2–3):73–83
Furuya M et al (2005) The C. Elegans eyes absent ortholog EYA-1 is required for tissue differentiation and plays partially redundant roles with PAX-6. Dev Biol 286(2):452–463
Amin NM et al (2009) A conserved six-eya cassette acts downstream of wnt signaling to direct non-myogenic versus myogenic fates in the C. Elegans postembryonic mesoderm. Dev Biol 331(2):350–360
Hirose T, Galvin BD, Horvitz HR (2010) Six and Eya promote apoptosis through direct transcriptional activation of the proapoptotic BH3-only gene egl-1 in Caenorhabditis elegans. Proc Natl Acad Sci U S A 107(35):15479–15484
Tootle TL et al (2003) The transcription factor eyes absent is a protein tyrosine phosphatase. Nature 426(6964):299–302
Bonini NM, Leiserson WM, Benzer S (1998) Multiple Roles eyes Absent gene Drosophila Dev Biol 196(1):42–57
Finardi A, Massari LF, Visintin R (2020) Anaphase Bridges: Not all Nat Fibers Are Healthy Genes (Basel), 11(8).
Smelick C, Ahmed S (2005) Achieving immortality in the C. Elegans germline. Ageing Res Rev 4(1):67–82
Ahmed S, Hodgkin J (2000) MRT-2 checkpoint protein is required for germline immortality and telomere replication in C. Elegans. Nature 403(6766):159–164
Craig AL et al (2012) Methods for studying the DNA damage response in the Caenorhabdatis Elegans germ line. Methods Cell Biol 107:321–352
Krishnan N et al (2009) Dephosphorylation of the C-terminal tyrosyl residue of the DNA damage-related histone H2A.X is mediated by the protein phosphatase eyes absent. J Biol Chem 284(24):16066–16070
Brenner S (1974) The genetics of Caenorhabditis elegans. Genetics 77(1):71–94
Consortium CeDM (2012) Large-scale screening for targeted knockouts in the Caenorhabditis elegans genome. G3 (Bethesda) 2(11):1415–1425
Dawson JA, Methven-Kelley C, Davis GM (2017) atz-1 influences meiosis to maintain germline chromosomal stability in Caenorhabditis elegans. Cell Biol Int 41(10):1160–1168
Dokshin GA et al (2020) GCNA interacts with Spartan and Topoisomerase II to Regulate Genome Stability. Dev Cell 52(1):53–68e6
Reboul J et al (2003) C. Elegans ORFeome version 1.1: experimental verification of the genome annotation and resource for proteome-scale protein expression. Nat Genet 34(1):35–41
Boag PR, Nakamura A, Blackwell TK (2005) A conserved RNA-protein complex component involved in physiological germline apoptosis regulation in C. Elegans. Development 132(22):4975–4986
Sievers F, Higgins DG (2018) Clustal Omega for making accurate alignments of many protein sequences. Protein Sci 27(1):135–145
Guex N, Peitsch MC, Schwede T (2009) Automated comparative protein structure modeling with SWISS-MODEL and Swiss-PdbViewer: a historical perspective. Electrophoresis 30(Suppl 1):S162–S173
Acknowledgements
We acknowledge Federation University for providing technical support and the facilities where this project was completed. Some strains were provided by the CGC, which is funded by NIH Office of Research Infrastructure Programs (P40 OD010440).
Funding
No funding bodies used for this research.
Open Access funding enabled and organized by CAUL and its Member Institutions
Author information
Authors and Affiliations
Contributions
HRT assisted with data collection, figure preparation and writing. SN contributed to data collection, PRB contributed to project design and GMD designed and assisted with data collection, figure preparation and writing.
Corresponding author
Ethics declarations
Ethical statements
No t applicable as all work was performed using C. elegans which is exempt from ethics approval.
Competing interests
The authors declare no competing interests.
Additional information
Publisher’s note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Electronic supplementary material
Below is the link to the electronic supplementary material.

11033_2024_9933_MOESM1_ESM.png
Supplementary Figure 1: Knockdown on eya-1 phenocopies eya-1(ok654) mutants and eya-1 mutants display enhanced germ cell apoptosis and a mild HIM phenotype: (A) eya-1(ok654) mutants show a low penetrance HIM phenotype compared to wild-type animals at 25°C. him-14(RNAi) represents positive control. *** p = < 0.0001, error bars = SEM, n = 700 replicated in triplicates. (B, C) Knockdown of eya-1 phenocopies DNA damage sensitivity to HU and CPT as observed in Fig. 4. n = 20, *** p = < 0.0001, error bars = SEM. Replicated in triplicates. (D, E) Germ cell death assessed via acridine orange (AO) staining in eya-1(ok654) mutants. Scale bar = 20 μm, n = 60, error bars = SEM
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution 4.0 International License, which permits use, sharing, adaptation, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons licence, and indicate if changes were made. The images or other third party material in this article are included in the article’s Creative Commons licence, unless indicated otherwise in a credit line to the material. If material is not included in the article’s Creative Commons licence and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this licence, visit http://creativecommons.org/licenses/by/4.0/.
About this article
Cite this article
Tatnell, H.R., Novakovic, S., Boag, P.R. et al. EYA-1 is required for genomic integrity independent of H2AX signalling in Caenorhabditis elegans. Mol Biol Rep 51, 1009 (2024). https://doi.org/10.1007/s11033-024-09933-4
Received:
Accepted:
Published:
DOI: https://doi.org/10.1007/s11033-024-09933-4