Abstract
Background
Breast cancer (BRCA) is the most common and leading cause of cancer-related death in women. MicroRNAs (miRNAs) are short non-coding RNA fragments that play a role in regulating gene expression including the cancer-related pathways. Although dysregulation of miR-223 has been demonstrated in recent studies to have prognostic value in various cancers, its diagnostic and prognostic role in BRCA remains unknown.
Methods
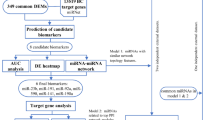
The expression and the prognostic value of miR-223 were evaluated using the TCGA data and verified by qRT-PCR. Subsequently, potential oncogenic targets of miR-223 were identified by using three different miRNA target prediction tools and the GEPIA database. In addition to these databases, protein-protein interaction network, molecular functions, prognostic value, and the expression level of miR-223 targets were included by using several other bioinformatics tools and databases; such as, UALCAN, GeneMANIA and Metascape.
Results
The bioinformatic results demonstrated that miR-223 downregulated in BRCA and associated with poor prognosis of patients. In vitro experiments validated that miR-223 significantly downregulated in BRCA cells, MCF-7, SK-BR3, MDA-MB-231 and HCC1500, compared to normal breast cell line hTERT-HME1. Furthermore, ANLN, DYNLT1, LRRC59, SLC12A8 and TPM3 genes were identified as the potential oncogenic target genes of miR-223 based on their expression and prognosis in BRCA. Additionally, protein-protein interaction network of these target genes was mainly enriched in dynein intermediate chain binding, cell division, regulation of cell cycle process, and positive regulation of cellular component biogenesis.
Conclusions
The results suggests that miR-223 and its targets, ANLN, DYNLT1, LRRC59, SLC12A8 and TPM3, might be reliable potential prognostic biomarkers in BRCA patients.






Similar content being viewed by others
Data Availability
The authors confirm that the data supporting the findings of this study are available within the article.
Code Availability
Not applicable.
References
Siegel RL, Miller KD, Fuchs HE, Jemal A (2022) Cancer statistics, 2022. CA Cancer J Clin 72:7–33. https://doi.org/10.3322/CAAC.21708
Jin X, Mu P (2015) Targeting breast Cancer metastasis. Breast Cancer (Auckl) 9:23. https://doi.org/10.4137/BCBCR.S25460
Cole M, Parajuli S, Laske D et al (2014) Peripheral primitive neuroectodermal tumor of the dura in a 51-year-old woman following intensive treatment for breast cancer. Am J Case Rep 15:294–299. https://doi.org/10.12659/AJCR.890656
de Moor CH, Meijer H, Lissenden S (2005) Mechanisms of translational control by the 3′ UTR in development and differentiation. Semin Cell Dev Biol 16:49–58. https://doi.org/10.1016/J.SEMCDB.2004.11.007
Bartel DP (2004) MicroRNAs: Genomics, Biogenesis, mechanism, and function. Cell 116:281–297. https://doi.org/10.1016/S0092-8674(04)00045-5
Sahin Y (2022) LncRNA H19 is a potential biomarker and correlated with immune infiltration in thyroid carcinoma. Clin Exp Med. https://doi.org/10.1007/S10238-022-00853-W
Liu H-T, Liu S, Liu L et al (2018) EGR1-mediated transcription of lncRNA-HNF1A-AS1 promotes cell-cycle progression in gastric cancer. Cancer Res 78:5877–5890
Liu B, Xiang W, Liu J et al (2021) The regulatory role of antisense lncRNAs in cancer. Cancer Cell International 2021 21:1 21:1–15. https://doi.org/10.1186/S12935-021-02168-4
Mathieu E-L, Belhocine M, Dao LT et al (2014) Functions of lncRNA in development and diseases. Med Sci (Paris) 30:790–796
Saliminejad K, Khorram Khorshid HR, Soleymani Fard S, Ghaffari SH (2019) An overview of microRNAs: Biology, functions, therapeutics, and analysis methods. J Cell Physiol 234:5451–5465. https://doi.org/10.1002/JCP.27486
Kabekkodu SP, Shukla V, Varghese VK et al (2018) Clustered miRNAs and their role in biological functions and diseases. Biol Rev 93:1955–1986. https://doi.org/10.1111/BRV.12428
Ardekani AM, Naeini MM (2010) The role of MicroRNAs in Human Diseases. Avicenna J Med Biotechnol 2:161
Seven M, Karatas OF, Duz MB, Ozen M (2014) The role of miRNAs in cancer: from pathogenesis to therapeutic implications. Future Oncol 10:1027–1048. https://doi.org/10.2217/FON.13.259
Li X, Zhang Y, Zhang H et al (2011) miRNA-223 promotes gastric cancer invasion and metastasis by targeting tumor suppressor EPB41L3. Mol Cancer Res 9:824–833. https://doi.org/10.1158/1541-7786.MCR-10-0529
Fazi F, Racanicchi S, Zardo G et al (2007) Epigenetic silencing of the myelopoiesis regulator microRNA-223 by the AML1/ETO oncoprotein. Cancer Cell 12:457–466. https://doi.org/10.1016/J.CCR.2007.09.020
Qu H, Zhu F, Dong H et al (2020) Upregulation of CCT-3 induces breast Cancer Cell Proliferation through miR-223 competition and Wnt/β-Catenin signaling pathway activation. https://doi.org/10.3389/FONC.2020.533176. Front Oncol 10:
Streppel MM, Pai S, Campbell NR et al (2013) MicroRNA 223 is upregulated in the multistep progression of Barrett’s esophagus and modulates sensitivity to chemotherapy by targeting PARP1. Clin Cancer Res 19:4067–4078. https://doi.org/10.1158/1078-0432.CCR-13-0601
Citron F, Segatto I, Vinciguerra GLR et al (2020) Downregulation of miR-223 expression is an early event during Mammary Transformation and confers resistance to CDK4/6 inhibitors in luminal breast Cancer. Cancer Res 80:1064–1077. https://doi.org/10.1158/0008-5472.CAN-19-1793
Sun X, Li Y, Zheng M et al (2016) MicroRNA-223 increases the sensitivity of Triple-Negative breast Cancer stem cells to TRAIL-Induced apoptosis by Targeting HAX-1. PLoS ONE 11:e0162754. https://doi.org/10.1371/JOURNAL.PONE.0162754
Catalanotto C, Cogoni C, Zardo G (2016) MicroRNA in Control of Gene expression: an overview of Nuclear Functions. Int J Mol Sci 17. https://doi.org/10.3390/IJMS17101712
Chandrashekar DS, Bashel B, Balasubramanya SAH et al (2017) UALCAN: a portal for facilitating Tumor Subgroup Gene expression and survival analyses. Neoplasia 19:649–658. https://doi.org/10.1016/J.NEO.2017.05.002
Zhou Y, Zhou B, Pache L et al (2019) Metascape provides a biologist-oriented resource for the analysis of systems-level datasets. Nature Communications 2019 10:1 10:1–10. https://doi.org/10.1038/s41467-019-09234-6
Ferlay J, Colombet M, Soerjomataram I et al (2021) Cancer statistics for the year 2020: an overview. Int J Cancer 149:778–789. https://doi.org/10.1002/IJC.33588
Carey LA, Perou CM, Livasy CA et al (2006) Race, breast cancer subtypes, and survival in the Carolina breast Cancer Study. JAMA 295:2492–2502. https://doi.org/10.1001/JAMA.295.21.2492
Koboldt DC, Fulton RS, McLellan MD et al (2012) Comprehensive molecular portraits of human breast tumors. Nature 490:61. https://doi.org/10.1038/NATURE11412
Lehmann BD, Bauer JA, Chen X et al (2011) Identification of human triple-negative breast cancer subtypes and preclinical models for selection of targeted therapies. J Clin Invest 121:2750–2767. https://doi.org/10.1172/JCI45014
Chang JTH, Wang F, Chapin W, Huang RS (2016) Identification of MicroRNAs as breast Cancer prognosis markers through the Cancer Genome Atlas. PLoS ONE 11:e0168284. https://doi.org/10.1371/JOURNAL.PONE.0168284
Ding HX, Lv Z, Yuan Y, Xu Q (2018) MiRNA polymorphisms and cancer prognosis: a systematic review and meta-analysis. Front Oncol 8:596. https://doi.org/10.3389/FONC.2018.00596/BIBTEX
Altan Z, Sahin Y (2022) miR-203 suppresses pancreatic cancer cell proliferation and migration by modulating DUSP5 expression. Mol Cell Probes 66:101866. https://doi.org/10.1016/J.MCP.2022.101866
Jia CY, Li HH, Zhu XC et al (2011) MiR-223 suppresses cell proliferation by targeting IGF-1R. https://doi.org/10.1371/JOURNAL.PONE.0027008. PLoS One 6:
Ji Q, Xu X, Song Q et al (2018) Mir-223-3p inhibits human Osteosarcoma Metastasis and Progression by directly targeting CDH6. Mol Ther 26:1299–1312. https://doi.org/10.1016/J.YMTHE.2018.03.009
Karakatsanis A, Papaconstantinou I, Gazouli M et al (2013) Expression of microRNAs, miR-21, miR-31, miR-122, miR-145, miR-146a, miR-200c, miR-221, miR-222, and miR-223 in patients with hepatocellular carcinoma or intrahepatic cholangiocarcinoma and its prognostic significance. Mol Carcinog 52:297–303. https://doi.org/10.1002/MC.21864
Yang Y, Jiang Z, Ma N et al (2018) MicroRNA-223 targeting STIM1 inhibits the Biological behavior of breast Cancer. Cell Physiol Biochem 45:856–866. https://doi.org/10.1159/000487180
Lv P, Zhang Z, Hou L et al (2020) Meta-analysis of the clinicopathological significance of miRNA-145 in breast cancer. Biosci Rep 40. https://doi.org/10.1042/BSR20193974
Nadeem F, Hanif M, Ahmed A et al (2017) Clinicopathological features associated to MiRNA-195 expression in patients with breast cancer: evidence of a potential biomarker. Pak J Med Sci 33:1242. https://doi.org/10.12669/PJMS.335.13008
Kangas R, Morsiani C, Pizza G et al (2018) Menopause and adipose tissue: miR-19a-3p is sensitive to hormonal replacement. Oncotarget 9:2279. https://doi.org/10.18632/ONCOTARGET.23406
Olivieri F, Ahtiainen M, Lazzarini R et al (2014) Hormone replacement therapy enhances IGF-1 signaling in skeletal muscle by diminishing miR-182 and miR-223 expressions: a study on postmenopausal monozygotic twin pairs. Aging Cell 13:850–861. https://doi.org/10.1111/ACEL.12245
Masciarelli S, Fontemaggi G, di Agostino S et al (2013) Gain-of-function mutant p53 downregulates miR-223 contributing to chemoresistance of cultured tumor cells. Oncogene 2014 33(12):1601–1608. https://doi.org/10.1038/onc.2013.106
Hirko KA, Rocque G, Reasor E et al (2022) The impact of race and ethnicity in breast cancer—disparities and implications for precision oncology. BMC Med 20. https://doi.org/10.1186/S12916-022-02260-0
Magnusson K, Gremel G, Rydén L et al (2016) ANLN is a prognostic biomarker independent of Ki-67 and essential for cell cycle progression in primary breast cancer. BMC Cancer 16. https://doi.org/10.1186/S12885-016-2923-8
Li LW, Xia J, Cui RT, Kong B (2021) Solute carrier family 12 member 8 impacts the biological behaviors of breast carcinoma cells by activating TLR/NLR signaling pathway. Cytotechnology 73:23–34. https://doi.org/10.1007/S10616-020-00439-Y
Yao B, Qu S, Hu R et al (2019) Delivery of platelet TPM3 mRNA into breast cancer cells via microvesicles enhances metastasis. FEBS Open Bio 9:2159. https://doi.org/10.1002/2211-5463.12759
Li D, Xing Y, Tian T et al (2020) Overexpression of LRRC59 is Associated with poor prognosis and promotes Cell Proliferation and Invasion in Lung Adenocarcinoma. Onco Targets Ther 13:6453. https://doi.org/10.2147/OTT.S245336
Funding
No funding was received.
Author information
Authors and Affiliations
Corresponding author
Ethics declarations
Conflict of interest
The authors declare that they have no known competing financial interests or personal relationships that could have appeared to influence the work reported in this paper.
Ethics approval and consent to participate
Not applicable.
Patient consent for publication
Not applicable.
Additional information
Publisher’s Note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Rights and permissions
Springer Nature or its licensor (e.g. a society or other partner) holds exclusive rights to this article under a publishing agreement with the author(s) or other rightsholder(s); author self-archiving of the accepted manuscript version of this article is solely governed by the terms of such publishing agreement and applicable law.
About this article
Cite this article
Sahin, Y., Altan, Z., Karabulut, A. et al. The role of miR-223 in breast cancer; an integrated analysis. Mol Biol Rep 50, 10179–10188 (2023). https://doi.org/10.1007/s11033-023-08850-2
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s11033-023-08850-2