Abstract
Background
Metabolic dysregulation and excessive inflammation are implicated in the pathogenesis of the highly infectious disease of coronavirus disease 2019 (COVID-19), which is caused by a newly emerging coronavirus (i.e., severe acute respiratory syndrome-coronavirus 2; SARS-CoV-2). The adenosine 5′-monophosphate-activated protein kinase (AMPK), an energy sensor regulating the metabolic pathways in diverse cells, exerts a regulatory role in the immune system. This study aims to examine the mRNA expression level of AMPK and the plasma levels of interleukin-6 (IL-6) and IL-10 cytokines in patients with different grades of COVID-19.
Methods
Peripheral blood was collected from 60 patients with COVID-19 (Moderate, severe, and critical). The plasma levels of IL-6 and IL-10 were quantified by enzyme-linked immunosorbent assay (ELISA), and the mRNA expression level of AMPK was determined using real-time PCR.
Results
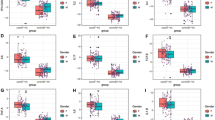
The results showed that the plasma levels of IL-6 increased significantly in critical and severe patients compared to moderate cases of COVID-19 (P < 0.001). Moreover, IL-10 plasma concentrations were significantly higher in critical and severe cases than in moderate cases of COVID-19 (P < 0.01 and P < 0.05, respectively). Also, the gene expression of AMPK was meaningfully enhanced in critical patients relative to moderate and severe cases of COVID-19, in order (P < 0.001 and P < 0.01, respectively). There was a positive association between AMPK gene expression and plasma levels of IL-6 and IL-10 (P = 0.006, r = 0.348, P = 0.028, r = 0.283, respectively).
Conclusion
Increasing AMPK gene expression is likely a necessary effort of the immune system to inhibit inflammation in critical COVID-19. However, this effort seems to be inadequate, probably due to factors that induce inflammation, like erythrocyte sedimentation rate (ESR) and IL-6.



Similar content being viewed by others
Data Availability
The datasets generated during and/or analyzed during the current study are available from the corresponding authors upon reasonable request.
Abbreviations
- AMPK:
-
Adenosine 5′-monophosphate-activated protein kinase
- COVID-19:
-
Coronavirus disease 2019
- SARS-CoV-2:
-
Severe acute respiratory syndrome-coronavirus 2
- IL-10:
-
Interleukin-10
- CRS:
-
Cytokine release syndrome
- ELISA:
-
Enzyme-linked immunosorbent assay
- qRT-PCR:
-
Quantitative real-time polymerase chain reaction
- ESR:
-
Erythrocyte sedimentation rate
References
Lotfi R, Kalmarzi RN, Roghani SA (2021) A review on the immune responses against novel emerging coronavirus (SARS-CoV-2). Immunol Res 69(3):213–224. https://doi.org/10.1007/s12026-021-09198-0
Siracusano G, Pastori C, Lopalco L (2020) Humoral Immune responses in COVID-19 patients: a window on the state of the art. Front Immunol 11:1049. https://doi.org/10.3389/fimmu.2020.01049
Deravi N, Ahsan E, Fathi M, Hosseini P, Yaghoobpoor S, Lotfi R, Pourbagheri-Sigaroodi A, Bashash D (2022) Complement inhibition: a possible therapeutic approach in the fight against Covid-19. Rev Med Virol 32(4):e2316. https://doi.org/10.1002/rmv.2316
Ciotti M, Ciccozzi M, Terrinoni A, Jiang WC, Wang CB, Bernardini S (2020) The COVID-19 pandemic. Crit Rev Clin Lab Sci 57(6):365–388. https://doi.org/10.1080/10408363.2020.1783198
Huang C, Wang Y, Li X, Ren L, Zhao J, Hu Y, Zhang L, Fan G, Xu J, Gu X, Cheng Z, Yu T, Xia J, Wei Y, Wu W, Xie X, Yin W, Li H, Liu M, Xiao Y, Gao H, Guo L, Xie J, Wang G, Jiang R, Gao Z, Jin Q, Wang J, Cao B (2020) Clinical features of patients infected with 2019 novel coronavirus in Wuhan, China. Lancet 395(10223):497–506. https://doi.org/10.1016/s0140-6736(20)30183-5
Kim J (2018) Regulation of Immune Cell Functions by Metabolic Reprogramming. J Immunol Res 2018:8605471. https://doi.org/10.1155/2018/8605471
Bettencourt IA, Powell JD (2017) Targeting metabolism as a Novel Therapeutic Approach to Autoimmunity, inflammation, and transplantation. J Immunol 198(3):999–1005. https://doi.org/10.4049/jimmunol.1601318
Gassen NC, Papies J, Bajaj T, Emanuel J, Dethloff F, Chua RL, Trimpert J, Heinemann N, Niemeyer C, Weege F, Hönzke K, Aschman T, Heinz DE, Weckmann K, Ebert T, Zellner A, Lennarz M, Wyler E, Schroeder S, Richter A, Niemeyer D, Hoffmann K, Meyer TF, Heppner FL, Corman VM, Landthaler M, Hocke AC, Morkel M, Osterrieder N, Conrad C, Eils R, Radbruch H, Giavalisco P, Drosten C, Müller MA (2021) SARS-CoV-2-mediated dysregulation of metabolism and autophagy uncovers host-targeting antivirals. Nat Commun 12(1):3818. https://doi.org/10.1038/s41467-021-24007-w
Wang T, Cao Y, Zhang H, Wang Z, Man CH, Yang Y, Chen L, Xu S, Yan X, Zheng Q, Wang YP (2022) COVID-19 metabolism: Mechanisms and therapeutic targets. MedComm (2020) 3 (3):e157. https://doi.org/10.1002/mco2.157
Oakhill JS, Chen ZP, Scott JW, Steel R, Castelli LA, Ling N, Macaulay SL, Kemp BE (2010) β-Subunit myristoylation is the gatekeeper for initiating metabolic stress sensing by AMP-activated protein kinase (AMPK). Proc Natl Acad Sci U S A 107(45):19237–19241. https://doi.org/10.1073/pnas.1009705107
Bhutta MS, Gallo ES, Borenstein R (2021) Multifaceted role of AMPK in viral infections. Cells 10(5):1118. https://doi.org/10.3390/cells10051118
Sag D, Carling D, Stout RD, Suttles J (2008) Adenosine 5’-monophosphate-activated protein kinase promotes macrophage polarization to an anti-inflammatory functional phenotype. J Immunol 181(12):8633–8641. https://doi.org/10.4049/jimmunol.181.12.8633
MacIver NJ, Blagih J, Saucillo DC, Tonelli L, Griss T, Rathmell JC, Jones RG (2011) The liver kinase B1 is a central regulator of T cell development, activation, and metabolism. J Immunol 187(8):4187–4198. https://doi.org/10.4049/jimmunol.1100367
Hardie DG (2011) AMP-activated protein kinase: an energy sensor that regulates all aspects of cell function. Genes Dev 25(18):1895–1908. https://doi.org/10.1101/gad.17420111
Mancini SJ, Salt IP (2018) Investigating the role of AMPK in inflammation. Methods Mol Biol 1732:307–319. https://doi.org/10.1007/978-1-4939-7598-3_20
Salminen A, Hyttinen JM, Kaarniranta K (2011) AMP-activated protein kinase inhibits NF-κB signaling and inflammation: impact on healthspan and lifespan. J Mol Med (Berl) 89(7):667–676. https://doi.org/10.1007/s00109-011-0748-0
Yin JX, Agbana YL, Sun ZS, Fei SW, Zhao HQ, Zhou XN, Chen JH, Kassegne K (2023) Increased interleukin-6 is associated with long COVID-19: a systematic review and meta-analysis. Infect Dis Poverty 12(1):43. https://doi.org/10.1186/s40249-023-01086-z
Nerstedt A, Johansson A, Andersson CX, Cansby E, Smith U, Mahlapuu M (2010) AMP-activated protein kinase inhibits IL-6-stimulated inflammatory response in human liver cells by suppressing phosphorylation of signal transducer and activator of transcription 3 (STAT3). Diabetologia 53(11):2406–2416. https://doi.org/10.1007/s00125-010-1856-z
Zhu YP, Brown JR, Sag D, Zhang L, Suttles J (2015) Adenosine 5’-monophosphate-activated protein kinase regulates IL-10-mediated anti-inflammatory signaling pathways in macrophages. J Immunol 194(2):584–594. https://doi.org/10.4049/jimmunol.1401024
Saraiva M, Vieira P, O’Garra A (2020) Biology and therapeutic potential of interleukin-10. J Exp Med 217(1):e20190418. https://doi.org/10.1084/jem.20190418
Lu L, Zhang H, Dauphars DJ, He YW (2021) A potential role of Interleukin 10 in COVID-19 pathogenesis. Trends Immunol 42(1):3–5. https://doi.org/10.1016/j.it.2020.10.012
Diao B, Wang C, Tan Y, Chen X, Liu Y, Ning L, Chen L, Li M, Liu Y, Wang G, Yuan Z, Feng Z, Zhang Y, Wu Y, Chen Y (2020) Reduction and functional exhaustion of T cells in patients with Coronavirus Disease 2019 (COVID-19). Front Immunol 11:827. https://doi.org/10.3389/fimmu.2020.00827
Han H, Ma Q, Li C, Liu R, Zhao L, Wang W, Zhang P, Liu X, Gao G, Liu F, Jiang Y, Cheng X, Zhu C, Xia Y (2020) Profiling serum cytokines in COVID-19 patients reveals IL-6 and IL-10 are disease severity predictors. Emerg Microbes Infect 9(1):1123–1130. https://doi.org/10.1080/22221751.2020.1770129
Pfaffl MW (2001) A new mathematical model for relative quantification in real-time RT-PCR. Nucleic Acids Res 29(9):e45. https://doi.org/10.1093/nar/29.9.e45
Szewczuk M, Boguszewska K, Kaźmierczak-Barańska J, Karwowski BT (2020) The role of AMPK in metabolism and its influence on DNA damage repair. Mol Biol Rep 47(11):9075–9086. https://doi.org/10.1007/s11033-020-05900-x
Juszczak F, Caron N, Mathew AV, Declèves AE (2020) Critical role for AMPK in Metabolic Disease-Induced chronic kidney disease. Int J Mol Sci 21(21):7994. https://doi.org/10.3390/ijms21217994
Haye A, Ansari MA, Rahman SO, Shamsi Y, Ahmed D, Sharma M (2020) Role of AMP-activated protein kinase on cardio-metabolic abnormalities in the development of diabetic cardiomyopathy: a molecular landscape. Eur J Pharmacol 888:173376. https://doi.org/10.1016/j.ejphar.2020.173376
Dasgupta B, Chhipa RR (2016) Evolving Lessons on the Complex Role of AMPK in Normal Physiology and Cancer. Trends Pharmacol Sci 37(3):192–206. https://doi.org/10.1016/j.tips.2015.11.007
Al-Kuraishy HM, Al-Gareeb AI, Alblihed M, Cruz-Martins N, Batiha GE (2021) COVID-19 and risk of Acute Ischemic Stroke and Acute Lung Injury in patients with type II diabetes Mellitus: the anti-inflammatory role of Metformin. Front Med (Lausanne) 8644295. https://doi.org/10.3389/fmed.2021.644295
Kratzel A, Kelly JN, V’Kovski P, Portmann J, Brüggemann Y, Todt D, Ebert N, Shrestha N, Plattet P, Staab-Weijnitz CA, von Brunn A, Steinmann E, Dijkman R, Zimmer G, Pfaender S, Thiel V (2021) A genome-wide CRISPR screen identifies interactors of the autophagy pathway as conserved coronavirus targets. PLoS Biol 19(12):e3001490. https://doi.org/10.1371/journal.pbio.3001490
Yim WW, Mizushima N (2021) Autophagosome maturation stymied by SARS-CoV-2. Dev Cell 56(4):400–402. https://doi.org/10.1016/j.devcel.2021.02.002
Zhao Z, Lu K, Mao B, Liu S, Trilling M, Huang A, Lu M, Lin Y (2021) The interplay between emerging human coronavirus infections and autophagy. Emerg Microbes Infect 10(1):196–205. https://doi.org/10.1080/22221751.2021.1872353
Qu Y, Wang X, Zhu Y, Wang W, Wang Y, Hu G, Liu C, Li J, Ren S, Xiao MZX, Liu Z, Wang C, Fu J, Zhang Y, Li P, Zhang R, Liang Q (2021) ORF3a-Mediated incomplete autophagy facilitates severe Acute Respiratory Syndrome Coronavirus-2 replication. Front Cell Dev Biol 9:716208. https://doi.org/10.3389/fcell.2021.716208
Miao G, Zhao H, Li Y, Ji M, Chen Y, Shi Y, Bi Y, Wang P, Zhang H (2021) ORF3a of the COVID-19 virus SARS-CoV-2 blocks HOPS complex-mediated assembly of the SNARE complex required for autolysosome formation. Dev Cell 56(4):427–442e425. https://doi.org/10.1016/j.devcel.2020.12.010
Bhaskar S, Sinha A, Banach M, Mittoo S, Weissert R, Kass JS, Rajagopal S, Pai AR, Kutty S (2020) Cytokine storm in COVID-19-Immunopathological mechanisms, clinical considerations, and therapeutic approaches: the REPROGRAM Consortium position paper. Front Immunol 11:1648. https://doi.org/10.3389/fimmu.2020.01648
Zhang C, Wu Z, Li JW, Zhao H, Wang GQ (2020) Cytokine release syndrome in severe COVID-19: interleukin-6 receptor antagonist tocilizumab may be the key to reduce mortality. Int J Antimicrob Agents 55(5):105954. https://doi.org/10.1016/j.ijantimicag.2020.105954
Tan L, Wang Q, Zhang D, Ding J, Huang Q, Tang YQ, Wang Q, Miao H (2020) Lymphopenia predicts disease severity of COVID-19: a descriptive and predictive study. Signal Transduct Target Ther 5(1):33. https://doi.org/10.1038/s41392-020-0148-4
Abbasifard M, Khorramdelazad H (2020) The bio-mission of interleukin-6 in the pathogenesis of COVID-19: a brief look at potential therapeutic tactics. Life Sci 257:118097. https://doi.org/10.1016/j.lfs.2020.118097
Zhang D, Zhou X, Yan S, Tian R, Su L, Ding X, Xiao M, Chen Y, Zhao H, Chen H, Zhang H, Li Z, Li Q, Xu Y, Yan X, Li Y, Zhang S (2020) Correlation between cytokines and coagulation-related parameters in patients with coronavirus disease 2019 admitted to ICU. Clin Chim Acta 510:47–53. https://doi.org/10.1016/j.cca.2020.07.002
Levi M, van der Poll T, ten Cate H, van Deventer SJ (1997) The cytokine-mediated imbalance between coagulant and anticoagulant mechanisms in sepsis and endotoxaemia. Eur J Clin Invest 27(1):3–9. https://doi.org/10.1046/j.1365-2362.1997.570614.x
Levi M, van der Poll T (2005) Two-way interactions between inflammation and coagulation. Trends Cardiovasc Med 15(7):254–259. https://doi.org/10.1016/j.tcm.2005.07.004
Du F, Liu B, Zhang S (2021) COVID-19: the role of excessive cytokine release and potential ACE2 down-regulation in promoting hypercoagulable state associated with severe illness. J Thromb Thrombolysis 51(2):313–329. https://doi.org/10.1007/s11239-020-02224-2
Que Y, Hu C, Wan K, Hu P, Wang R, Luo J, Li T, Ping R, Hu Q, Sun Y, Wu X, Tu L, Du Y, Chang C, Xu G (2022) Cytokine release syndrome in COVID-19: a major mechanism of morbidity and mortality. Int Rev Immunol 41(2):217–230. https://doi.org/10.1080/08830185.2021.1884248
Chen X, Zhao B, Qu Y, Chen Y, Xiong J, Feng Y, Men D, Huang Q, Liu Y, Yang B, Ding J, Li F (2020) Detectable serum severe Acute Respiratory Syndrome Coronavirus 2 viral load (RNAemia) is closely correlated with drastically elevated interleukin 6 level in critically ill patients with Coronavirus Disease 2019. Clin Infect Dis 71(8):1937–1942. https://doi.org/10.1093/cid/ciaa449
Zhou F, Yu T, Du R, Fan G, Liu Y, Liu Z, Xiang J, Wang Y, Song B, Gu X, Guan L, Wei Y, Li H, Wu X, Xu J, Tu S, Zhang Y, Chen H, Cao B (2020) Clinical course and risk factors for mortality of adult inpatients with COVID-19 in Wuhan, China: a retrospective cohort study. Lancet 395(10229):1054–1062. https://doi.org/10.1016/s0140-6736(20)30566-3
Gong J, Dong H, Xia QS, Huang ZY, Wang DK, Zhao Y, Liu WH, Tu SH, Zhang MM, Wang Q, Lu FE (2020) Correlation analysis between disease severity and inflammation-related parameters in patients with COVID-19: a retrospective study. BMC Infect Dis 20(1):963. https://doi.org/10.1186/s12879-020-05681-5
Liu X, Wang H, Shi S, Xiao J (2022) Association between IL-6 and severe disease and mortality in COVID-19 disease: a systematic review and meta-analysis. Postgrad Med J 98(1165):871–879. https://doi.org/10.1136/postgradmedj-2021-139939
Islam H, Chamberlain TC, Mui AL, Little JP (2021) Elevated Interleukin-10 levels in COVID-19: potentiation of pro-inflammatory responses or impaired anti-inflammatory action? Front Immunol 12:677008. https://doi.org/10.3389/fimmu.2021.677008
Zhao Y, Qin L, Zhang P, Li K, Liang L, Sun J, Xu B, Dai Y, Li X, Zhang C, Peng Y, Feng Y, Li A, Hu Z, Xiang H, Ogg G, Ho LP, McMichael A, Jin R, Knight JC, Dong T, Zhang Y (2020) Longitudinal COVID-19 profiling associates IL-1RA and IL-10 with disease severity and RANTES with mild disease. JCI Insight 5(13):e139834. https://doi.org/10.1172/jci.insight.139834
Lauw FN, Pajkrt D, Hack CE, Kurimoto M, van Deventer SJ, van der Poll T (2000) Proinflammatory effects of IL-10 during human endotoxemia. J Immunol 165(5):2783–2789. https://doi.org/10.4049/jimmunol.165.5.2783
Wang F, Hou H, Luo Y, Tang G, Wu S, Huang M, Liu W, Zhu Y, Lin Q, Mao L, Fang M, Zhang H, Sun Z (2020) The laboratory tests and host immunity of COVID-19 patients with different severity of illness. JCI Insight 5(10):e137799. https://doi.org/10.1172/jci.insight.137799
Majidpoor J, Mortezaee K (2022) Interleukin-6 in SARS-CoV-2 induced disease: interactions and therapeutic applications. Biomed Pharmacother 145:112419. https://doi.org/10.1016/j.biopha.2021.112419
Mishra D, Richard JE, Maric I, Porteiro B, Häring M, Kooijman S, Musovic S, Eerola K, López-Ferreras L, Peris E, Grycel K, Shevchouk OT, Micallef P, Olofsson CS, Wernstedt Asterholm I, Grill HJ, Nogueiras R, Skibicka KP (2019) Parabrachial Interleukin-6 reduces Body Weight and Food Intake and increases thermogenesis to regulate Energy Metabolism. Cell Rep 26(11):3011–3026e3015. https://doi.org/10.1016/j.celrep.2019.02.044
Li YS, Ren HC, Cao JH (2022) Roles of Interleukin-6-mediated immunometabolic reprogramming in COVID-19 and other viral infection-associated diseases. Int Immunopharmacol 110:109005. https://doi.org/10.1016/j.intimp.2022.109005
Rose-John S (2021) Therapeutic targeting of IL-6 trans-signaling. Cytokine 144:155577. https://doi.org/10.1016/j.cyto.2021.155577
Acknowledgments
The authors would like to thank all patients who generously participated in this study. Also, the authors acknowledge the Deputy of Research and Technology of the KUMS, Kermanshah, Iran for financial support of the current study.
Funding
This study was financially supported by the deputy of research and technology of the Kermanshah University of Medical Sciences (KUMS), Kermanshah, Iran [Grant Number: 4000648].
Author information
Authors and Affiliations
Contributions
All authors listed have made a substantial, direct, and intellectual contribution to the work and approved it for publication.
Corresponding authors
Ethics declarations
Competing Interests
The authors declare that they have no known competing financial interests or personal relationships that could have appeared to influence the work reported in this paper.
Ethics approval
This study was performed in line with the principles of the Declaration of Helsinki. Approval was granted by the Ethics Committee of the Kermanshah University of Medical Sciences (KUMS), Kermanshah, Iran (Ethical code: IR.KUMS.MED.REC.1400.058).
Consent to participate
Informed consent was obtained from all individual participants included in the study.
Consent to publish
The authors affirm that human research participants provided informed consent for the publication of the manuscript results.
Additional information
Publisher’s Note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Rights and permissions
Springer Nature or its licensor (e.g. a society or other partner) holds exclusive rights to this article under a publishing agreement with the author(s) or other rightsholder(s); author self-archiving of the accepted manuscript version of this article is solely governed by the terms of such publishing agreement and applicable law.
About this article
Cite this article
Assar, S., Dastbaz, M., Amini, K. et al. Assessing the gene expression of the adenosine 5′-monophosphate-activated protein kinase (AMPK) and its relation with the IL-6 and IL-10 plasma levels in COVID-19 patients. Mol Biol Rep 50, 9925–9933 (2023). https://doi.org/10.1007/s11033-023-08835-1
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s11033-023-08835-1