Abstract
Background
Malnutrition affects large section of population worldwide. Vitamin A and protein deficiencies have emerged as the major global health-issue. Traditional shrunken2 (sh2)-based sweet corn is deficient in provitamin A (proA), lysine and tryptophan. Natural variant of β-carotene hydroxylase1 (crtRB1) and opaque2 (o2) enhances proA, lysine and tryptophan in maize. So far, no sweet corn hybrid rich in these nutrients has been released elsewhere. Development of biofortified sweet corn hybrids would help in providing the balanced nutrition.
Methods and Results
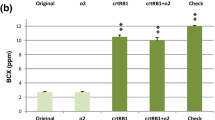
We targeted three sh2-based sweet corn inbreds (SWT-19, SWT-20 and SWT-21) for introgression of mutant crtRB1 and o2 genes using molecular breeding. The gene-based 3′TE-InDel and simple sequence repeat (SSR) (umc1066) markers specific to crtRB1 and o2, respectively were utilized in foreground selection in BC1F1, BC2F1 and BC2F2. Segregation distortion was observed for crtRB1 and o2 genes in majority of populations. Background selection using 91–100 SSRs revealed recovery of recurrent parent genome (RPG) up to 96%. The introgressed progenies possessed significantly higher proA (13.56 µg/g) as compared to the original versions (proA: 2.70 µg/g). Further, the introgressed progenies had accumulated moderately higher level of lysine (0.336%) and tryptophan (0.082%) over original versions (lysine: 0.154% and tryptophan: 0.038%). Kernel sweetness among introgressed progenies (17.3%) was comparable to original sweet corn (17.4%). The introgressed inbreds exhibited higher resemblance with their recurrent parents for yield and morphological characters.
Conclusion
These newly developed biofortified sweet corn genotypes hold immense promise to alleviate malnutrition.




Similar content being viewed by others
Data availability
All the data sets supporting the conclusion of this article are included within the article, and its supporting information files are provided as accompanying supplementary materials.
References
von Grebmer K, Bernstein J, Resnick D, Wiemers M, Reiner L, Bachmeier M, Hanano A, Towey O, Chéilleachair RN, Foley C, Gitter S, Larocque G, Fritschel H (2022) Global Hunger Index: Food Systems Transformation and Local Governance. Bonn: Welthungerhilfe; and Dublin: Concern Worldwide.
Global Nutrition Report (2022). Stronger commitments for greater action. Executive summary. www.globalnutritionreport.org.
World Health Organization (2021). Malnutrition; Available online: https://www.who.int/en/news-room/fact-sheets/detail/malnutrition Accessed on 8 December 2022
Hossain F, Zunjare RU, Muthusamy V, Kumar A, Madhavan J, Ikkurti G, Katral A, Talukder ZA, Chhabra R, Chand G, Bhatt V, Gul I, Mishra SJ, Duo H, Dutta S, Gain N, Chauhan P, Maman S, Reddappa SB, Kasana R (2023) Genetic improvement of specialty corn for nutritional quality traits. In: Wani SH et al (eds) Boo: maize improvement. Springer International Publishing, Chem, pp 235–257. https://doi.org/10.1007/978-3-031-21640-4_11
Lonzano-Alejo N, Carrillo VG, Pixley K, Rojas NP (2007) Physical properties and carotenoid content of maize kernels and its nixtamalized snacks. Innov Food Sci Emerg Technol 8:385–389. https://doi.org/10.1016/j.ifset.2007.03.015
Giuliano G (2017) Provitamin A biofortification of crop plants: a gold rush with many miners. Curr Opin Biotechnol 44:169–180. https://doi.org/10.1016/j.copbio.2017.02.001
Rice AL, West KP Jr, Black RE (2004) Vitamin A deficiency. Comp Quantification Health Risks 1:0211–0256
Caulfield LE, Richard SA, Rivera JA, Musgrove P, Black RE (2006) Stunting, wasting, and micronutrient deficiency disorders. Disease Control Priorities in Developing Countries. 2nd edition.
Bain LE, Awah PK, Geraldine N, Kindong NP, Siga Y, Bernard N, Tanjeko AT (2013) Malnutrition in sub-Saharan Africa: burden, causes and prospects. Pan Afr Med J 15:1–9. https://doi.org/10.11604/pamj.2013.15.120.2535]
Nyakurwa CS, Gasura E, Mabasa S (2017) Potential for quality protein maize for reducing protein energy undernutrition in maize dependent sub-saharan African countries: a review. Afr Crop Sci J 25:521–537. https://doi.org/10.4314/acsj.v25i4.9
Hossain F, Muthusamy V, Pandey N, Vishwakarma AK, Baveja A, Zunjare RU, Thirunavukkarasu N, Saha S, Manjaiah KM, Prasanna BM, Gupta HS (2018) Marker-assisted introgression of opaque2 allele for rapid conversion of elite hybrids into quality protein maize. J Genet 97:287–298. https://doi.org/10.1007/s12041-018-0914-z
Sarika K, Hossain F, Muthusamy V, Zunjare RU, Baveja A, Goswami R, Bhat JS, Saha S, Gupta HS (2018) Marker-assisted pyramiding of opaque2 and novel opaque16 genes for further enrichment of lysine and tryptophan in sub-tropical maize. Plant Sci J 272:142–152. https://doi.org/10.1016/j.plantsci.2018.04.014
Mehta BK, Muthusamy V, Baveja A, Chauhan HS, Chhabra R, Bhatt V, Chand G, Zunjare RU, Singh AK, Hossain F (2020) Composition analysis of lysine, tryptophan and provitamin-A during different stages of kernel development in biofortified sweet corn. J Food Compos Anal 94:103625. https://doi.org/10.1016/j.jfca.2020.103625
FAOSTAT (2020) https://www.fao.org/faostat/en/#data (data accessed on 12 December, 2022).
Lertrat K, Pulam T (2007) Breeding for increased sweetness in sweet corn. Int J Plant Breed Genet 1:27–30
Feng ZL, Liu J, Fu FL, Li WC (2008) Molecular mechanism of sweet and waxy in maize. Int J Plant Breed Genet 2:93–100. https://doi.org/10.3923/ijpbg.2008.93.100
Chhabra R, Muthusamy V, Baveja A, Katral A, Mehta B, Zunjare RU, Hossain F (2022) Allelic variation in shrunken2 gene affecting kernel sweetness in exotic-and indigenous-maize inbreds. PLoS ONE 17(9):e0274732. https://doi.org/10.1371/journal.pone.0274732
Baveja A, Muthusamy V, Panda KK, Zunjare RU, Das AK, Chhabra R, Mishra SJ, Mehta BK, Saha S, Hossain F (2021) Development of multinutrient-rich biofortified sweet corn hybrids through genomics-assisted selection of shrunken2, opaque2, lcyE and crtRB1 genes. J Appl Genet 62(3):419–429. https://doi.org/10.1007/s13353-021-00633-4
Feng F, Wang Q, Liang C, Yang R, Li X (2015) Enhancement of tocopherols in sweet corn by marker-assisted backcrossing of ZmVTE4. Euphytica 206:513–521. https://doi.org/10.1007/s10681-015-1519-8
Yan J, Kandianis BC, Harjes EC, Bai L, Kim HE, Yang X, Skinner DJ, Fu Z, Mitchell S, Li Q, Fernandez GSM, Zaharoeva M, Babu R, Fu Y, Palacios N, Li J, DallaPanna D, Brutnall T, Buckler SE, Warburton LM, Rocheford T (2010) Rare genetic variation at Zea mays crtRB1 increases β-carotene in maize grain. Nat Genet 42:322–327. https://doi.org/10.1038/ng.551
Mertz ET, Bates LS, Nelson OE (1964) Mutant genes that change protein composition and increase lysine content of maize endosperm. Science 145:279–280. https://doi.org/10.1126/science.145.3629.279
Gupta HS, Vignesh M, Hossain F, Nepolean T (2013) Enrichment of nutritional qualities in maize through marker-assisted selection. National Seminar on Genomics for Crop Improvement, IBAB, Bangalore, pp 69–70.
Chandrasekharan N, Ramanathan N, Pukalenthy B, Chandran S, Manickam D, Adhimoolam K, Natesan (2022) Development of β-carotene, lysine, and tryptophan-rich maize (Zea mays) inbreds through marker-assisted gene pyramiding. Sci Rep 12(1):8551
Zunjare RU, Hossain F, Muthusamy V, Baveja A, Chauhan HS, Bhat JS, Thirunavukkarasu N, Saha S, Gupta HS (2018) Development of biofortified maize hybrids through marker-assisted stacking of β-carotene hydroxylase, lycopene-ε-cyclase and opaque2 genes. Front Plant Sci 9:178. https://doi.org/10.1371/journal.pone.0113583
Neuffer MG, Coe EH, Wessler SR (1997) Mutants of maize. Cold Spring Harbor Laboratory Press, New York
Murray MG, Thompson WF (1980) Rapid isolation of high molecular weight plant DNA. Nucleic Acids Res 8:4321–4325. https://doi.org/10.1093/nar/8.19.4321
Muthusamy V, Hossain F, Thirunavakkarasu N, Choudhary M, Saha S, Bhat JS, Prasanna BM, Gupta HS (2014) Development of β-carotene rich maize hybrids through marker-assisted introgression of β-carotene hydroxylase Allele. PLoS ONE 9(12):e113583
Duo H, Hossain F, Muthusamy V, Zunjare RU, Goswami R, Chand G, Mishra SJ, Chhabra R, Gowda MM, Pal S, Baveja A, Bhat JS, Kamboj MC, Kumar B, Amalraj JJ, Khulbe R, Prakash B, Neeraja CN, Rakshit RS, Yadav OP (2021) Development of sub-tropically adapted diverse provitamin-A rich maize inbreds through marker assisted pedigree selection, their characterization and utilization in hybrid breeding. PLoS ONE 16(2):e0245497. https://doi.org/10.3382/ps.2011-01719
PPVFRA (2007) Guidelines for the conduct of test for distinctiveness, uniformity and stability on maize (Zea mays L.). Plant Var J India 1(1):1–13
Kurilich AC, Juvik JA (1999) Quantification of carotenoid and tocopherol antioxidants in Zea mays. J Agric Food Chem 47:1948–1955. https://doi.org/10.1021/jf981029d
Babu R, Rojas NP, Gao S, Yan J, Pixley K (2013) Validation of the effects of molecular marker polymorphisms in LcyE and CrtRB1 on provitamin A concentrations for 26 tropical maize populations. Theor Appl Genet 126:389–399. https://doi.org/10.1007/s00122-012-1987-3
Chand G, Muthusamy V, Allen T, Zunjare RU, Mishra SJ, Singh B, Mehta BK, Talukder ZA, Ismail MR, Sarika K, Kamboj MC (2022) Composition of lysine and tryptophan among biofortified-maize possessing novel combination of opaque2 and opaque16 genes. J Food Compos Anal 107:104376. https://doi.org/10.1016/j.jfca.2021.104376
Chauhan HS, Muthusamy V, Rashmi T, Basu S, Anand A, Mehta BK, Gain N, Zunjare RU, Singh AK, Gupta HS, Hossain F (2022) Characterization of crtRB1-and vte4-based biofortified sweet corn inbreds for seed vigour and physico-biochemical traits. J Appl Genet 63(4):651–662. https://doi.org/10.1007/s13353-022-00715-x
Kodrzycki R, Boston RS, Larkins BA (1989) The opaque-2 mutation of maize differentially reduces zein gene transcription. Plant Cell 1(1):105–114. https://doi.org/10.1105/tpc.1.1.105
Ueda T, Waverczak W, Ward K, Sher N, Ketudat M, Schmidt RJ, Messing J (1992) Mutations of the 22-and 27-kD zein promoters affect transactivation by the opaque-2 protein. Plant Cell 4(6):701–709. https://doi.org/10.1105/tpc.4.6.701
Acknowledgements
We thank WNC, IIMR, Hyderabad for providing the off-season nursery.
Funding
We are thankful to the ICAR-IARI, New Delhi for financial support.
Author information
Authors and Affiliations
Contributions
Conduct of the experiment: BS, Generation of backcross populations: RUZ, Recording of phenotypic data: GC and VB, Estimation of quality parameters: NG, Statistical analysis: VM, Writing of manuscript: BS, GC and FH, Designing of experiment: FH and SS.
Corresponding author
Ethics declarations
Conflict of interest
Authors declare that no conflict of interest exits.
Ethical approval
This article does not contain any studies with human participants or animals performed by any of the authors.
Additional information
Publisher's Note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary Information
Below is the link to the electronic supplementary material.
Rights and permissions
Springer Nature or its licensor (e.g. a society or other partner) holds exclusive rights to this article under a publishing agreement with the author(s) or other rightsholder(s); author self-archiving of the accepted manuscript version of this article is solely governed by the terms of such publishing agreement and applicable law.
About this article
Cite this article
Singh, B., Zunjare, R.U., Shrivastava, S. et al. Provitamin A, lysine and tryptophan enrichment in shrunken2-based sweet corn genotypes through genomics-assisted breeding for crtRB1 and opaque2 genes. Mol Biol Rep 50, 4965–4974 (2023). https://doi.org/10.1007/s11033-023-08446-w
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s11033-023-08446-w