Abstract
Objective
Free-range (FR) poultry production systems are associated with quality products and improved welfare. All the 19 diverse chicken breeds of India have evolved under the FR system and are adapted to different agro-climatic conditions. It is vital to explore indigenous germplasm with modern genomic tools to have insights into genomic characteristics of production traits and adaptation.
Methods
In this study, breast tissue transcriptome profiles were generated and analyzed from four biological replicates of two indigenous backyard poultry breeds of India-Ankaleshwar, a breed of the mainland, and Nicobari, a breed adapted to islands. The read quality of sequences was checked by FASTQC and processed reads were aligned to the reference genome (bGalGal1).
Results
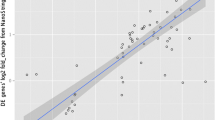
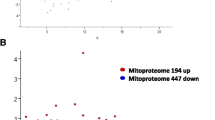
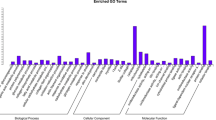
More than 94% mapping to the reference genome was observed for all samples. A total of 12,790 transcripts were common across both groups, while 657 were expressed only in Ankaleshwar and 169 in Nicobari. The highest expressed genes across both groups were associated mainly with muscle structure, contraction, and energy metabolism. The highly expressed genes identified in Ankaleshwar were involved in fatty acid catabolism and oxidative stress mitigation. Functional terms, pathways, and hub genes in Nicobari participated in muscle fiber growth, adipogenesis, and fatty acid anabolism. A key hub gene (RAC1) in Nicobari is a potential candidate affecting the laying rate in chickens. The qRT-PCR results also substantiate the RNA-seq results.
Conclusion
The findings provide a precious molecular resource to advance understanding of the genetic basis of adaptation, meat quality, and egg production in backyard chickens.





Similar content being viewed by others
References
Pellattiero E, Tasoniero G, Cullere M et al (2020) Are meat quality traits and sensory attributes in favor of slow-growing chickens? Animals (Basel) 10:960. https://doi.org/10.3390/ani10060960
Xiang H, Gao J, Yu B et al (2014) Early holocene chicken domestication in northern China. Proc Natl Acad Sci USA 111(49):17564–17569. https://doi.org/10.1073/pnas.141188211
Whitnall T, Pitts N (2019) Global trends in meat consumption. Agric Commod 9(1):96–99. https://doi.org/10.3316/informit.309517990386547
Zeuner FE (1963) A history of domesticated animals. Hutchinson & Co (Publishers) Ltd, London. https://doi.org/10.1017/S0003598X00068952
Singh NP, Bhatt N, Usman SM, Chaudhary P (2020) A detailed review on backyard poultry production and management in India. J Entomol Zool Stud 8(2):1411–1415
Rajkumar U, Rama Rao SV, Raju MVLN, Chatterjee RN (2021) Backyard poultry farming for sustained production and enhanced nutritional and livelihood security with special reference to India: a review. Trop Anim Health Prod 53(1):176. https://doi.org/10.1007/s11250-021-02621-6
Sehrawat R, Sharma R, Ahlawat S et al (2021) First report on better functional property of black chicken meat from India. Ind J Anim Res 55(6):723–733. https://doi.org/10.18805/IJAR.B-4014
Eleroglu H, Yildirim A, Sekero˘glu A, Çoksöyler FN, Duman M (2014) Comparison of growth curves by growth models in slow-growing chicken genotypes raised the organic system. Int J Agric Biol 16:529–535
Xiang H, Chen S, Zhang H et al (2018) Transcriptome changes provide genetic insights into the effects of rearing systems on chicken welfare and product quality. J Anim Sci 96(11):4552–4561. https://doi.org/10.1093/jas/sky314
De AK, Kundu A, Ruban VV, Kundu MS, Jeyakumar S, Sunder J (2013) Antibody response to goat erythrocytes in endangered Nicobari fowl Vanaraja and their various F1 and F2 crosses under hot humid climate of Andaman and Nicobar Islands of India. J Appl Anim Res 41(2):125–132
Mishra AK (2022) Poultry genetic resources of India and its role in rural poultry production. Indian J Plant Genet Resour 35(3):242–246. https://doi.org/10.5958/0976-1926.2022.00076.6
Jassal B, Matthews L, Viteri G et al (2020) The reactome pathway knowledge base. Nucleic Acids Res 48(D1):D498–D503. https://doi.org/10.1093/nar/gkz1031
Huang DW, Sherman BT, Lempicki RA (2009) Bioinformatics enrichment tools; paths toward the comprehensive functional analysis of large gene lists. Nucleic Acids Res 37(1):1–13. https://doi.org/10.1093/nar/gkn923
Huang DW, Sherman BT, Lempicki RA (2009) Systematic and integrative analysis of large gene lists using DAVID bioinformatics resources. Nat Protoc 4(1):44–57. https://doi.org/10.1038/nprot.2008.211
Shannon P, Markiel A, Ozier O et al (2003) Cytoscape: a software environment for integrated models of biomolecular interaction networks. Genome Res 13(11):2498–2504. https://doi.org/10.1101/gr.1239303
Chin CH, Chen SH, Wu HH, Ho CW, Ko MT, Lin CY (2014) cytoHubba: identifying hub objects and sub-networks from complex interactome. BMC Syst Biol 8(Suppl 4):S11. https://doi.org/10.1186/1752-0509-8-S4-S11
Untergasser A, Cutcutache I, Koressaar T et al (2012) Primer3—new capabilities and interfaces. Nucleic Acids Res 40(15):e115. https://doi.org/10.1093/nar/gks596
Chiang HI, Berghman LR, Zhou H (2009) Inhibition of NF-kB 1 (NF-kBp50) by RNA interference in chicken macrophage HD11 cell line challenged with Salmonella enteritidis. Genet Mol Biol 32(3):507–515. https://doi.org/10.1590/S1415-47572009000300013
Cogburn LA, Nares T, Chuming C et al (2018) Transcriptional profiling of liver during the critical embryo-to-hatchling transition period in the chicken (Gallus gallus). BMC Genomics 19(1):695. https://doi.org/10.1186/s12864-018-5080-4
Wu J, Lin Z, Chen G et al (2021) Characterization of chicken skin yellowness and exploration of genes involved in skin yellowness deposition in skin. Front Physiol. https://doi.org/10.3389/fphys.2021.58508
Sharma R, Sehrawat R, Ahlawat S et al (2022) An attempt to valorize the only black meat chicken breed of India by delineating superior functional attributes of its meat. Sci Rep 12:3555. https://doi.org/10.1038/s41598-022-07575-9
Liu J, Lei Q, Li F, Zhou Y, Gao J, Liu W, Han H, Cao D (2020) Dynamic transcriptomic analysis of breast muscle development from the embryonic to post-hatching periods in chickens. Front Genet 10(10):1308. https://doi.org/10.3389/fgene.2019.01308
Na W, Wang Y, Gong P, Zhang X, Zhang K, Zhang H, Wang N, Li H (2021) Screening of reference genes for RT-qPCR in chicken adipose tissue and adipocytes. Front Physiol 12:676864. https://doi.org/10.3389/fphys.2021.676864
Livak KJ, Schmittgen TD (2001) Analysis of relative gene expression data using real-time quantitative PCR and the 2(-delta delta C (T)) method. Methods 25:402–408. https://doi.org/10.1006/meth.2001.1262
Gregorich ZR, Cai W, Lin Z et al (2017) Distinct sequences and post-translational modifications in cardiac atrial and ventricular myosin light chains revealed by top-down mass spectrometry. J Mol Cell Cardiol 107:13–21. https://doi.org/10.1016/j.yjmcc.2017.04.002
Zhao D, Kogut MH, Genovese KJ, Hsu CY, Lee JT, Farnell YZ (2020) Altered expression of lactate dehydrogenase and monocarboxylate transporter involved in lactate metabolism in broiler wooden breast. Poult Sci 99(1):11–20. https://doi.org/10.3382/ps/pez572
Schell JC, Wisidagama DR, Bensard C et al (2017) Control of intestinal stem cell function and proliferation by mitochondrial pyruvate metabolism. Nat Cell Biol 19(9):1027–1036. https://doi.org/10.1038/ncb3593
Houten SM, Violante S, Ventura FV, Wanders RJ (2016) The biochemistry and physiology of mitochondrial fatty acid β-oxidation and its genetic disorders. Annu Rev Physiol 78:23–44. https://doi.org/10.1146/annurev-physiol-021115-105045
Li S, Lu CW, Diem EC et al (2022) Acetyl-CoA-carboxylase 1-mediated de novo fatty acid synthesis sustains Lgr5+ intestinal stem cell function. Nat Commun 13:3998. https://doi.org/10.1038/s41467-022-31725-2
Chen L, Toke NH, Luo S, Vasoya et al (2019) A reinforcing HNF4–SMAD4 feed-forward module stabilizes enterocyte identity. Nat Genet 51(5):777–785. https://doi.org/10.1038/s41588-019-0384-0
Quijano C, Trujillo M, Castro L, Trostchansky A (2016) Interplay between oxidant species and energy metabolism. Redox Biol 8:28–42. https://doi.org/10.1016/j.redox.2015.11.010
Morgan MJ, Liu ZG (2011) Crosstalk of reactive oxygen species and NF-κB signaling. Cell Res 21(1):103–115. https://doi.org/10.1038/cr.2010.178
Hager-Theodorides AL, Massouras T, Simitzis PE et al (2021) Hesperidin and naringin improve broiler meat fatty acid profile and modulate the expression of genes involved in fatty acid β-oxidation and antioxidant defense in a dose dependent manner. Foods. https://doi.org/10.3390/foods10040739
Murach KA, Liu Z, Jude B (2022) Multi-transcriptome analysis following an acute skeletal muscle growth stimulus yields tools for discerning global and MYC regulatory networks. J Biol Chem 298(11):102515. https://doi.org/10.1016/j.jbc.2022.102515
Aspenström P (2018) Activated rho GTPases in cancer—the beginning of a new paradigm. Int J Mol Sci 19(12):3949. https://doi.org/10.3390/ijms19123949
Ma Z, Jiang K, Wang D et al (2021) Comparative analysis of hypothalamus transcriptome between laying hens with different egg-laying rates. Poult Sci 100(7):101110. https://doi.org/10.1016/j.psj.2021.101110
Nematbakhsh S, Pei CP, Selamat J, Nordin N, Idris LH, Abdull Razis AF (2021) Molecular regulation of lipogenesis adipogenesis and fat deposition in chicken. Genes. https://doi.org/10.3390/genes12030414
Mir NA, Rafiq A, Kumar F, Singh V, Shukla V (2017) Determinants of broiler chicken meat quality and factors affecting them: a review. J Food Sci Technol 54(10):2997–3009. https://doi.org/10.1007/s13197-017-2789-z
Kong BW, Hudson N, Seo D et al (2017) RNA sequencing for global gene expression associated with muscle growth in a single male modern broiler line compared to a foundational barred Plymouth Rock chicken line. BMC Genomics 18(1):82. https://doi.org/10.1186/s12864-016-3471-y
Ren T, Li Z, Zhou Y et al (2018) Sequencing and characterization of lncRNAs in the breast muscle of Gushi and Arbor Acres chickens. Genome 61:337–347. https://doi.org/10.1139/gen-2017-0114
Kumar H, Choo H, Iskender AU et al (2020) RNA seq analyses of chicken reveals biological pathways involved in acclimation into different geographical locations. Sci Rep 10:19288. https://doi.org/10.1038/s41598-020-76234-8
Acknowledgements
This work was financially supported by ICAR-CABin Scheme, New Delhi. We are grateful to the Director, ICAR-National Bureau of Animal Genetic Resources (NBAGR), Karnal, and Indian Council of Agricultural Research (ICAR), New Delhi for providing the necessary facilities.
Author information
Authors and Affiliations
Contributions
All authors contributed to the study’s conception and design. Material preparation and data collection were performed by RS, SA, RA, PC, AK, MK, and data curation and analysis were done by SBL, DCM, and MSF. The first draft of the manuscript was written by RS, and all authors commented on and improved the manuscript. All authors read and approved the final manuscript.
Corresponding author
Ethics declarations
Conflict of interest
The authors declare that they have no known competing financial interests or personal relationships that could have appeared to influence the work reported in this paper.
Ethical approval
All protocols were carried out in compliance with Committee for the Purpose of Control and Supervision on Experiments on Animals (CPCSEA) rules. The animals were not experimented upon and the muscle samples were purchased in liaison with the butchers.
Informed consent
All authors consent to the submission of this manuscript to the journal Molecular Biology Reports.
Additional information
Publisher’s Note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary Information
Below is the link to the electronic supplementary material.
Rights and permissions
Springer Nature or its licensor (e.g. a society or other partner) holds exclusive rights to this article under a publishing agreement with the author(s) or other rightsholder(s); author self-archiving of the accepted manuscript version of this article is solely governed by the terms of such publishing agreement and applicable law.
About this article
Cite this article
Sharma, R., Arora, R., Ahlawat, S. et al. Study on the muscle transcriptome of two diverse Indian backyard poultry breeds acclimatized to different agro-ecological conditions. Mol Biol Rep 50, 2453–2461 (2023). https://doi.org/10.1007/s11033-022-08223-1
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s11033-022-08223-1