Abstract
Background
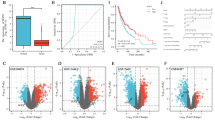
The PAX3 (paired box gene 3) gene is highly expressed in several cancer types. However, its underlying mechanism of action in skin cutaneous melanoma (SKCM) remains unknown.
Methods
In this study, we used the GEPIA database and western blotting to analyze the expression of PAX3. We performed the Kaplan-Meier survival analysis to evaluate the prognostic value of PAX3 in SKCM. Next, the Kyoto Encyclopedia of Genes and Genomes (KEGG) pathway enrichment analysis was performed to evaluate the function of PAX3-related co-expressed genes. Additionally, the function and potential mechanism of action of PAX3 in SKCM were studied through functional experiments. Western blotting was used to detect the changes in the levels of epithelial-mesenchymal transition (EMT)-related and MET (c-MET tyrosine kinase) proteins following PAX3 knockdown. Finally, we assessed the correlation between PAX3 expression and the infiltration of CD4+/CD8+ T cells using the TISIDB database.
Results
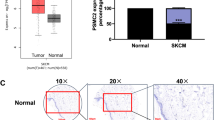
We found that PAX3 was overexpressed in the SKCM tissues and that these levels were indicative of a poor prognosis of SKCM. The KEGG pathway enrichment analysis showed that PAX3-related co-expressed genes were mainly associated with the oncogenic pathways. Knocking down PAX3 significantly inhibited the proliferation, invasion, and migration of SK-MEL-28 cells. The PAX3 expression was related significantly to the immune infiltration level of CD4+/CD8+ T cells.
Conclusions
Our findings demonstrated that PAX3 knockdown could reverse the EMT of tumor cells, inhibit the growth, and progression of SKCM cells. Therefore, PAX3 may have implications as a potential therapeutic target and promising prognostic biomarker for SKCM.






Similar content being viewed by others
Data Availability Statement
Part of the data is derived from the TCGA/GTEX data set and other original data are provided by the author and will not be retained.
References
Sung H, Ferlay J, Siegel RL, Laversanne M, Soerjomataram I, Jemal A et al (2021) Global Cancer Statistics 2020: GLOBOCAN Estimates of Incidence and Mortality Worldwide for 36 Cancers in 185 Countries. CA Cancer J Clin 71(3):209–249. https://doi.org/10.3322/caac.21660
Hamid O, Robert C, Daud A, Hodi FS, Hwu WJ, Kefford R et al (2019) Five-year survival outcomes for patients with advanced melanoma treated with pembrolizumab in KEYNOTE-001. Ann Oncol 30(4):582–588. https://doi.org/10.1093/annonc/mdz011
Tremblay P, Gruss P (1994) Pax: genes for mice and men. Pharmacol Ther 61(1–2):205–226. https://doi.org/10.1016/0163-7258(94)90063-9
Baldwin CT, Hoth CF, Macina RA, Milunsky A (1995) Mutations in PAX3 that cause Waardenburg syndrome type I: ten new mutations and review of the literature. Am J Med Genet 58(2):115–122. https://doi.org/10.1002/ajmg.1320580205
Rajan P, Panchision DM, Newell LF, Mckay RD (2003) BMPs signal alternately through a SMAD or FRAP-STAT pathway to regulate fate choice in CNS stem cells. J Cell Biol 161(5):911–921. https://doi.org/10.1083/jcb.200211021
Wu M, Li J, Engleka KA, Zhou B, Lu MM, Plotkin JB et al (2008) Persistent expression of Pax3 in the neural crest causes cleft palate and defective osteogenesis in mice. J Clin Invest 118(6):2076–2087. https://doi.org/10.1172/JCI33715
Wang Q, Fang WH, Krupinski J, Kumar S, Slevin M, Kumar P (2008) Pax genes in embryogenesis and oncogenesis. J Cell Mol Med 12(6A):2281–2294. https://doi.org/10.1111/j.1582-4934.2008.00427.x
Mascarenhas JB, Littlejohn EL, Wolsky RJ, Young KP, Nelson M, Salgia R et al (2010) PAX3 and SOX10 activate MET receptor expression in melanoma. Pigment Cell Melanoma Res 23(2):225–237. https://doi.org/10.1111/j.1755-148X.2010.00667.x
Tang Z, Li C, Kang B, Gao G, Li C, Zhang Z (2017) GEPIA: a web server for cancer and normal gene expression profiling and interactive analyses. Nucleic Acids Res 45(W1):W98–W102. https://doi.org/10.1093/nar/gkx247
Vasaikar SV, Straub P, Wang J, Zhang B (2018) LinkedOmics: analyzing multi-omics data within and across 32 cancer types. Nucleic Acids Res 46(D1):D956–D963. https://doi.org/10.1093/nar/gkx1090
Ru B, Wong CN, Tong Y, Zhong JY, Zhong S, Wu WC et al (2019) TISIDB: an integrated repository portal for tumor-immune system interactions. Bioinformatics 35(20):4200–4202. https://doi.org/10.1093/bioinformatics/btz210
Chen M, Zhang L, Liu X, Ma Z, Lv L (2021) PER1 Is a Prognostic Biomarker and Correlated With Immune Infiltrates in Ovarian Cancer. Front Genet 12:697471. https://doi.org/10.3389/fgene.2021.697471
Zhang K, Tian R, Wang G, Zhang J, Ma H, Hu X et al (2020) ADAMTS8 Inhibits Cell Proliferation and Invasion, and Induces Apoptosis in Breast Cancer. Onco Targets Ther 13:8373–8382. https://doi.org/10.2147/OTT.S248085
Stanton SE, Disis ML (2016) Clinical significance of tumor-infiltrating lymphocytes in breast cancer. J Immunother Cancer 4:59. https://doi.org/10.1186/s40425-016-0165-6
Clark WJ, Elder DE, Guerry DT, Braitman LE, Trock BJ, Schultz D et al (1989) Model predicting survival in stage I melanoma based on tumor progression. J Natl Cancer Inst 81(24):1893–1904. https://doi.org/10.1093/jnci/81.24.1893
Waardenburg PJ (1951) A new syndrome combining developmental anomalies of the eyelids, eyebrows and nose root with pigmentary defects of the iris and head hair and with congenital deafness. Am J Hum Genet 3(3):195–253
Wang-Wuu S, Soukup S, Ballard E, Gotwals B, Lampkin B (1988) Chromosomal analysis of sixteen human rhabdomyosarcomas. Cancer Res 48(4):983–987
Gryder BE, Yohe ME, Chou HC, Zhang X, Marques J, Wachtel M et al (2017) PAX3-FOXO1 Establishes Myogenic Super Enhancers and Confers BET Bromodomain Vulnerability. Cancer Discov 7(8):884–899. https://doi.org/10.1158/2159-8290.CD-16-1297
Gryder BE, Yohe ME, Chou HC, Zhang X, Marques J, Wachtel M et al (2017) PAX3-FOXO1 Establishes Myogenic Super Enhancers and Confers BET Bromodomain Vulnerability. Cancer Discov 7(8):884–899. https://doi.org/10.1158/2159-8290.CD-16-1297
Pandey PR, Chatterjee B, Olanich ME, Khan J, Miettinen MM, Hewitt SM et al (2017) PAX3-FOXO1 is essential for tumour initiation and maintenance but not recurrence in a human myoblast model of rhabdomyosarcoma. J Pathol 241(5):626–637. https://doi.org/10.1002/path.4867
Tonelli R, Mcintyre A, Camerin C, Walters ZS, Di Leo K, Selfe J et al (2012) Antitumor activity of sustained N-myc reduction in rhabdomyosarcomas and transcriptional block by antigene therapy. Clin Cancer Res 18(3):796–807. https://doi.org/10.1158/1078-0432.CCR-11-1981
Pandey PR, Chatterjee B, Olanich ME, Khan J, Miettinen MM, Hewitt SM et al (2017) PAX3-FOXO1 is essential for tumour initiation and maintenance but not recurrence in a human myoblast model of rhabdomyosarcoma. J Pathol 241(5):626–637. https://doi.org/10.1002/path.4867
Marshall AD, Lagutina I, Grosveld GC (2011) PAX3-FOXO1 induces cannabinoid receptor 1 to enhance cell invasion and metastasis. Cancer Res 71(24):7471–7480. https://doi.org/10.1158/0008-5472.CAN-11-0924
Libura J, Drukala J, Majka M, Tomescu O, Navenot JM, Kucia M et al (2002) CXCR4-SDF-1 signaling is active in rhabdomyosarcoma cells and regulates locomotion, chemotaxis, and adhesion. Blood 100(7):2597–2606. https://doi.org/10.1182/blood-2002-01-0031
Fang WH, Wang Q, Li HM, Ahmed M, Kumar P, Kumar S (2014) PAX3 in neuroblastoma: oncogenic potential, chemosensitivity and signalling pathways. J Cell Mol Med 18(1):38–48. https://doi.org/10.1111/jcmm.12155
Fujii R, Osaka E, Sato K, Tokuhashi Y (2019) MiR-1 Suppresses Proliferation of Osteosarcoma Cells by Up-regulating p21 via PAX3. Cancer Genomics Proteomics 16(1):71–79. https://doi.org/10.21873/cgp.20113
Medic S, Ziman M (2010) PAX3 expression in normal skin melanocytes and melanocytic lesions (naevi and melanomas). PLoS ONE 5(4):e9977. https://doi.org/10.1371/journal.pone.0009977
Bailey CM, Morrison JA, Kulesa PM (2012) Melanoma revives an embryonic migration program to promote plasticity and invasion. Pigment Cell Melanoma Res 25(5):573–583. https://doi.org/10.1111/j.1755-148X.2012.01025.x
Reid AL, Millward M, Pearce R, Lee M, Frank MH, Ireland A et al (2013) Markers of circulating tumour cells in the peripheral blood of patients with melanoma correlate with disease recurrence and progression. Br J Dermatol 168(1):85–92. https://doi.org/10.1111/bjd.12057
Liu F, Cao J, Lv J, Dong L, Pier E, Xu GX et al (2013) TBX2 expression is regulated by PAX3 in the melanocyte lineage. Pigment Cell Melanoma Res 26(1):67–77. https://doi.org/10.1111/pcmr.12029
Huang L, Zhai Y, La J, Lui JW, Moore S, Little EC et al (2021) Targeting Pan-ETS Factors Inhibits Melanoma Progression. Cancer Res 81(8):2071–2085. https://doi.org/10.1158/0008-5472.CAN-19-1668
Thiery JP, Sleeman JP (2006) Complex networks orchestrate epithelial-mesenchymal transitions. Nat Rev Mol Cell Biol 7(2):131–142. https://doi.org/10.1038/nrm1835
Kalluri R, Weinberg RA (2009) The basics of epithelial-mesenchymal transition. J Clin Invest 119(6):1420–1428. https://doi.org/10.1172/JCI39104
Lee JM, Dedhar S, Kalluri R, Thompson EW (2006) The epithelial-mesenchymal transition: new insights in signaling, development, and disease. J Cell Biol 172(7):973–981. https://doi.org/10.1083/jcb.200601018
Masuda T, Ueo H, Kai Y, Noda M, Hu Q, Sato K et al (2020) N-Cadherin mRNA Levels in Peripheral Blood Could Be a Potential Indicator of New Metastases in Breast Cancer: A Pilot Study. Int J Mol Sci 21(2). https://doi.org/10.3390/ijms21020511
Lamouille S, Xu J, Derynck R (2014) Molecular mechanisms of epithelial-mesenchymal transition. Nat Rev Mol Cell Biol 15(3):178–196. https://doi.org/10.1038/nrm3758
Kim HJ, Cantor H (2014) CD4 T-cell subsets and tumor immunity: the helpful and the not-so-helpful. Cancer Immunol Res 2(2):91–98. https://doi.org/10.1158/2326-6066.CIR-13-0216
Shankaran V, Ikeda H, Bruce AT, White JM, Swanson PE, Old LJ et al (2018) Pillars Article: IFNgamma and Lymphocytes Prevent Primary Tumour Development and Shape Tumour Immunogenicity. Nature. 2001. 410: 1107–1111. J Immunol 201(3):827–831
Gonzalez H, Hagerling C, Werb Z (2018) Roles of the immune system in cancer: from tumor initiation to metastatic progression. Genes Dev 32(19–20):1267–1284. https://doi.org/10.1101/gad.314617.118
Durgeau A, Virk Y, Corgnac S, Mami-Chouaib F (2018) Recent Advances in Targeting CD8 T-Cell Immunity for More Effective Cancer Immunotherapy. Front Immunol 9:14. https://doi.org/10.3389/fimmu.2018.00014
Martinez-Lostao L, Anel A, Pardo J (2015) How Do Cytotoxic Lymphocytes Kill Cancer Cells? Clin Cancer Res 21(22):5047–5056. https://doi.org/10.1158/1078-0432.CCR-15-0685
Acknowledgments
We thank Bullet Edits Limited for the linguistic editing and proofreading of our manuscript.
Funding
This research was supported by the Guangxi Provincial Department of Education (2021KY0521).
Author information
Authors and Affiliations
Contributions
All authors made substantial contributions to the conception and design, acquisition of data, or analysis and interpretation of data; aided in drafting the article or revising it critically for important intellectual content; provided final approval of the version to be published; and agreed to be accountable for all aspects of the work.
Corresponding authors
Ethics declarations
Competing interests
The author reports no conflicts of interest in this work.
Ethics approval and consent to participate
This study was approved by the Ethics Committee of the Second Affiliated Hospital of Shantou University Medical College (ethics number: 2021-117). Informed consent was obtained from all patients participating in this study.
Additional information
Publisher’s Note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Rights and permissions
Springer Nature or its licensor holds exclusive rights to this article under a publishing agreement with the author(s) or other rightsholder(s); author self-archiving of the accepted manuscript version of this article is solely governed by the terms of such publishing agreement and applicable law.
About this article
Cite this article
Zhang, K., Yu, C., Tian, R. et al. Downregulation of the paired box gene 3 inhibits the progression of skin cutaneous melanoma by inhibiting c-MET tyrosine kinase. Mol Biol Rep 49, 9137–9145 (2022). https://doi.org/10.1007/s11033-022-07706-5
Received:
Revised:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s11033-022-07706-5