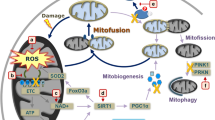
Abstract
Background
Rapamycin is hormetic in nature—it demonstrates contrasting effects at high and low doses. It is toxic at moderate/high doses, while it can restrain aging and extend lifespan at low doses. However, it is not fully understood how rapamycin governs cellular aging. On the other hand, aging is putatively correlated to mitochondrial dysregulation. Although previous studies have suggested that hormetic (low) doses of rapamycin can cause partial/incomplete inhibition of mTOR, the actual modus operandi of how such partial mTOR inhibition might modulate the mTOR-mitochondria cross-talk remained to be deciphered in the context of cellular aging. The present study was designed to understand the hormetic effects of rapamycin on cellular factors that govern aging-associated changes in mitochondrial facets, such as functional and metabolic homeostases, sustenance of membrane potential, biogenesis, mitophagy, and oxidative injury to mitochondrial macromolecules.
Methods and results
WRL-68 cells treated (24 h) with variable doses of rapamycin were studied for estimating their viability, apoptosis, senescence, mitochondrial density and Δψm. Expression levels of key functional proteins were estimated by immunofluorescence/immunoblots. Oxidative damage to mtDNA/mtRNA/proteins was measured in mitochondrial lysates. We demonstrated that hormetic doses (0.1 and 1 nM) of rapamycin can alleviate aging-associated mitochondrial dyshomeostasis in WRL-68 cells, such as oxidative injury to mitochondrial nucleic acids and proteins, as well as disequilibrium of mitochondrial density, membrane potential, biogenesis, mitophagy and overall metabolism.
Conclusions
We established that low doses of rapamycin can hormetically amend the mTOR-mitochondria cross-talk, and can consequently promote anti-aging outcome in cells.





Similar content being viewed by others
Data availability
Not applicable.
Code availability
Not applicable.
Abbreviations
- 4E-BP1:
-
Eukaryotic translation initiation factor 4E-binding protein 1
- 8-OH-dG:
-
8-Hydroxy-2’-deoxyguanosine
- 8-OH-g:
-
8-Hydroxyguanine
- 8-OH-G:
-
8-Hydroxyguanosine
- BECN1:
-
Beclin 1
- COXII:
-
Cytochrome c oxidase subunit II
- DAPI:
-
4′,6-Diamidino-2-phenylindole
- DHA:
-
Dinitrophenyl hydrazone adduct
- DHE:
-
Hydroethidine/dihydroethidium
- DNPH:
-
2,4-Dinitrophenylhydrazine
- Dox:
-
Doxorubicin
- eIF4E:
-
Eukaryotic translation initiation factor 4E
- FOXO1:
-
Forkhead box protein O1
- ITP:
-
Interventions Testing Program
- mtDNA:
-
Mitochondrial DNA
- mtOGR:
-
Mitochondrial OGR
- mTOR:
-
Mechanistic target of RAP
- mtRNA:
-
Mitochondrial RNA
- MTT:
-
3-(4,5- Dimethyl-2-thiazolyl)-2,5-diphenyl-2H-tetrazolium bromide
- OGR:
-
Oxidized guanine residues
- p70S6K:
-
70 kDa ribosomal protein S6 kinase
- PGC-1α:
-
Peroxisome proliferator-activated receptor gamma coactivator 1α
- PINK1:
-
PTEN-induced kinase 1
- RAP:
-
Rapamycin
- rTdT:
-
Recombinant terminal deoxynucleotidyl transferase
- SA-β-gal:
-
Senescence-associated β-galactosidase
- SDH:
-
Succinate dehydrogenase
- SDS:
-
Sodium dodecyl sulfate
- SIRT1:
-
Sirtuin 1
- TFAM:
-
Mitochondrial transcription factor A,
- TUNEL:
-
Terminal deoxynucleotidyl transferase dUTP nick-end labeling
- X-gal:
-
5-Bromo-4-chloro-3-indolyl β-D-galactopyranoside
- YY1:
-
Yin yang 1
- Δψm:
-
Mitochondrial membrane potential
References
Harrison DE, Strong R, Sharp ZD et al (2009) Rapamycin fed late in life extends lifespan in genetically heterogeneous mice. Nature 460:392–395. https://doi.org/10.1038/nature08221
Miller RA, Harrison DE, Astle CM et al (2011) Rapamycin, but not resveratrol or simvastatin, extends life span of genetically heterogeneous mice. J Gerontol A Biol Sci Med Sci 66:191–201. https://doi.org/10.1093/gerona/glq178
NIA (2020) National Institute on Aging - Interventions Testing Program (ITP). In: National Institute on Aging. http://www.nia.nih.gov/research/dab/interventions-testing-program-itp. Accessed 27 Dec 2020
Barlow AD, Nicholson ML, Herbert TP (2013) Evidence for rapamycin toxicity in pancreatic β-cells and a review of the underlying molecular mechanisms. Diabetes 62:2674–2682. https://doi.org/10.2337/db13-0106
Blagosklonny MV (2019) Fasting and rapamycin: diabetes versus benevolent glucose intolerance. Cell Death Dis 10:607. https://doi.org/10.1038/s41419-019-1822-8
Bhakta-Guha D, Efferth T (2015) Hormesis: decoding two sides of the same coin. Pharmaceuticals (Basel) 8:865–883. https://doi.org/10.3390/ph8040865
Law BK (2005) Rapamycin: an anti-cancer immunosuppressant? Crit Rev Oncol Hematol 56:47–60. https://doi.org/10.1016/j.critrevonc.2004.09.009
Sharp ZD, Bartke A (2005) Evidence for down-regulation of phosphoinositide 3-kinase/Akt/mammalian target of rapamycin (PI3K/Akt/mTOR)-dependent translation regulatory signaling pathways in Ames dwarf mice. J Gerontol A Biol Sci Med Sci 60:293–300
Sun L, Sadighi Akha AA, Miller RA, Harper JM (2009) Life-span extension in mice by preweaning food restriction and by methionine restriction in middle age. J Gerontol A Biol Sci Med Sci 64:711–722. https://doi.org/10.1093/gerona/glp051
Sciarretta S, Forte M, Frati G, Sadoshima J (2018) New insights into the role of mTOR signaling in the cardiovascular system. Circ Res 122:489–505. https://doi.org/10.1161/CIRCRESAHA.117.311147
Harman D (1972) The biologic clock: the mitochondria? J Am Geriatr Soc 20:145–147
Das P, Guha G (2011) Aging and mitochondrial DNA. J Sci Res 3:176–186. https://doi.org/10.3329/jsr.v3i1.5078
Barja G (2013) Updating the mitochondrial free radical theory of aging: an integrated view, key aspects, and confounding concepts. Antioxid Redox Signal 19:1420–1445. https://doi.org/10.1089/ars.2012.5148
Sanchez-Roman I, Gómez A, Pérez I et al (2012) Effects of aging and methionine restriction applied at old age on ROS generation and oxidative damage in rat liver mitochondria. Biogerontology 13:399–411. https://doi.org/10.1007/s10522-012-9384-5
Mahalakshmi R, Priyanga J, Vedha Hari BN et al (2020) Hexavalent chromium-induced autophagic death of WRL-68 cells is mitigated by aqueous extract of Cuminum cyminum L. seeds. 3 Biotech 10:191. https://doi.org/10.1007/s13205-020-02184-7
Schindelin J, Arganda-Carreras I, Frise E et al (2012) Fiji: an open-source platform for biological-image analysis. Nat Methods 9:676–682. https://doi.org/10.1038/nmeth.2019
Chiang GG, Abraham RT (2005) Phosphorylation of mammalian target of rapamycin (mTOR) at Ser-2448 Is mediated by p70S6 kinase. J Biol Chem 280:25485–25490. https://doi.org/10.1074/jbc.M501707200
Gelmetti V, De Rosa P, Torosantucci L et al (2017) PINK1 and BECN1 relocalize at mitochondria-associated membranes during mitophagy and promote ER-mitochondria tethering and autophagosome formation. Autophagy 13:654–669. https://doi.org/10.1080/15548627.2016.1277309
Ma X, Su P, Yin C et al (2020) The roles of FoxO transcription factors in regulation of bone cells function. Int J Mol Sci 21:692. https://doi.org/10.3390/ijms21030692
Ngo HB, Lovely GA, Phillips R, Chan DC (2014) Distinct structural features of TFAM drive mitochondrial DNA packaging versus transcriptional activation. Nat Commun 5:3077. https://doi.org/10.1038/ncomms4077
Nadon NL, Miller RA, Strong R, Harrison DE (2016) Chapter 10—NIA interventions testing program: a collaborative approach for investigating interventions to promote healthy aging. In: Kaeberlein MR, Martin GM (eds) Handbook of the biology of aging, 8th edn. Academic Press, San Diego, pp 287–303
van Deursen JM (2014) The role of senescent cells in ageing. Nature 509:439–446. https://doi.org/10.1038/nature13193
Guha G, Lu W, Li S et al (2015) Novel pactamycin analogs induce p53 dependent cell-cycle arrest at s-phase in human head and neck squamous cell carcinoma (HNSCC) cells. PLoS ONE 10:e0125322. https://doi.org/10.1371/journal.pone.0125322
Wu D, Prives C (2018) Relevance of the p53-MDM2 axis to aging. Cell Death Differ 25:169–179. https://doi.org/10.1038/cdd.2017.187
Rufini A, Tucci P, Celardo I, Melino G (2013) Senescence and aging: the critical roles of p53. Oncogene 32:5129–5143. https://doi.org/10.1038/onc.2012.640
Pearce LR, Alton GR, Richter DT et al (2010) Characterization of PF-4708671, a novel and highly specific inhibitor of p70 ribosomal S6 kinase (S6K1). Biochem J 431:245–255. https://doi.org/10.1042/BJ20101024
Katta A, Kakarla S, Wu M et al (2009) Altered regulation of contraction-induced Akt/mTOR/p70S6k pathway signaling in skeletal muscle of the obese Zucker rat. Exp Diabetes Res 2009:384683. https://doi.org/10.1155/2009/384683
Ayuso MI, Hernández-Jiménez M, Martín ME et al (2010) New hierarchical phosphorylation pathway of the translational repressor eIF4E-binding protein 1 (4E-BP1) in ischemia-reperfusion stress. J Biol Chem 285:34355–34363. https://doi.org/10.1074/jbc.M110.135103
Morita M, Gravel S-P, Hulea L et al (2015) mTOR coordinates protein synthesis, mitochondrial activity and proliferation. Cell Cycle 14:473–480. https://doi.org/10.4161/15384101.2014.991572
Ye Z, Li G, Kim C et al (2018) Regulation of miR-181a expression in T cell aging. Nat Commun 9:3060. https://doi.org/10.1038/s41467-018-05552-3
Anderson R, Prolla T (2009) PGC-1alpha in aging and anti-aging interventions. Biochim Biophys Acta 1790:1059–1066. https://doi.org/10.1016/j.bbagen.2009.04.005
Wenz T (2011) Mitochondria and PGC-1α in aging and age-associated diseases. J Aging Res 2011:e810619. https://doi.org/10.4061/2011/810619
Chen G, Kroemer G, Kepp O (2020) Mitophagy: an emerging role in aging and age-associated diseases. Front Cell Dev Biol 8:200. https://doi.org/10.3389/fcell.2020.00200
Mai S, Klinkenberg M, Auburger G et al (2010) Decreased expression of Drp1 and Fis1 mediates mitochondrial elongation in senescent cells and enhances resistance to oxidative stress through PINK1. J Cell Sci 123:917–926. https://doi.org/10.1242/jcs.059246
Zheng L, Bernard-Marissal N, Moullan N et al (2017) Parkin functionally interacts with PGC-1α to preserve mitochondria and protect dopaminergic neurons. Hum Mol Genet 26:582–598. https://doi.org/10.1093/hmg/ddw418
Xiang F, Ma S, Lv Y et al (2019) Tumor necrosis factor receptor-associated protein 1 regulates hypoxia-induced apoptosis through a mitochondria-dependent pathway mediated by cytochrome c oxidase subunit II. Burns Trauma 7:16. https://doi.org/10.1186/s41038-019-0154-3
Takahashi K, Miura Y, Ohsawa I et al (2018) In vitro rejuvenation of brain mitochondria by the inhibition of actin polymerization. Sci Rep 8:15585. https://doi.org/10.1038/s41598-018-34006-5
Calabrese V, Cornelius C, Dinkova-Kostova AT et al (2010) Cellular stress responses, the hormesis paradigm, and vitagenes: novel targets for therapeutic intervention in neurodegenerative disorders. Antioxid Redox Signal 13:1763–1811. https://doi.org/10.1089/ars.2009.3074
Siracusa R, Scuto M, Fusco R et al (2020) Anti-inflammatory and anti-oxidant activity of hidrox® in rotenone-induced Parkinson’s disease in mice. Antioxidants (Basel) 9:824. https://doi.org/10.3390/antiox9090824
Ristow M, Schmeisser K (2014) Mitohormesis: promoting health and lifespan by increased levels of reactive oxygen species (ROS). Dose Response 12:288–341. https://doi.org/10.2203/dose-response.13-035.Ristow
Herrero A, Barja G (2001) Effect of aging on mitochondrial and nuclear DNA oxidative damage in the heart and brain throughout the life-span of the rat. J Am Aging Assoc 24:45–50. https://doi.org/10.1007/s11357-001-0006-4
Sebastián D, Sorianello E, Segalés J et al (2016) Mfn2 deficiency links age-related sarcopenia and impaired autophagy to activation of an adaptive mitophagy pathway. EMBO J 35:1677–1693. https://doi.org/10.15252/embj.201593084
Gureev AP, Shaforostova EA, Popov VN (2019) Regulation of mitochondrial biogenesis as a way for active longevity: interaction between the Nrf2 and PGC-1α signaling pathways. Front Genet 10:435. https://doi.org/10.3389/fgene.2019.00435
Kitada M, Ogura Y, Koya D (2016) The protective role of Sirt1 in vascular tissue: its relationship to vascular aging and atherosclerosis. Aging 8:2290–2307. https://doi.org/10.18632/aging.101068
Salminen A, Kaarniranta K, Kauppinen A (2013) Crosstalk between oxidative stress and SIRT1: impact on the aging process. Int J Mol Sci 14:3834–3859. https://doi.org/10.3390/ijms14023834
Lee S-H, Lee J-H, Lee H-Y, Min K-J (2019) Sirtuin signaling in cellular senescence and aging. BMB Rep 52:24–34. https://doi.org/10.5483/BMBRep.2019.52.1.290
Yuan Y, Cruzat VF, Newsholme P et al (2016) Regulation of SIRT1 in aging: roles in mitochondrial function and biogenesis. Mech Ageing Dev 155:10–21. https://doi.org/10.1016/j.mad.2016.02.003
Zhang M, Zhang Q, Hu Y et al (2017) miR-181a increases FoxO1 acetylation and promotes granulosa cell apoptosis via SIRT1 downregulation. Cell Death Dis 8:e3088. https://doi.org/10.1038/cddis.2017.467
Liu T, Yang Q, Zhang X et al (2020) Quercetin alleviates kidney fibrosis by reducing renal tubular epithelial cell senescence through the SIRT1/PINK1/mitophagy axis. Life Sci 257:118116. https://doi.org/10.1016/j.lfs.2020.118116
Hu Q, Wang G, Peng J, et al (2017) Knockdown of SIRT1 Suppresses Bladder Cancer Cell Proliferation and Migration and Induces Cell Cycle Arrest and Antioxidant Response through FOXO3a-Mediated Pathways. In: BioMed Research International. https://www.hindawi.com/journals/bmri/2017/3781904/. Accessed 25 Dec 2020
Acknowledgements
The authors acknowledge Dr. Sudarshan Singh Rathore, Dr. M.R. Charan Raja, Dr. Sandeep Miryala and Ms. Nirekshana Krishnasagar for assistance with experiments and blinded confirmatory estimations. The authors also thank Dr. N. Saisubramanian, SCBT, SASTRA University, for his support. This study was funded by the Science and Engineering Research Board (SERB), Department of Science and Technology (DST), Government of India (Grant Nos. YSS/2014/000139 & YSS/2015/000025), Department of Science and Technology “Fund for Improvement of S&T Infrastructure in Universities and Higher Educational institutions” (DST-FIST), Government of India [Grant No. SR/FST/ETI-331/2013 (SASTRA)], and Department of Biotechnology (DBT), Government of India Grant No. BT/PR22434/MED/30/1901/2017.
Author information
Authors and Affiliations
Contributions
GG and DBG: Conception and design of the study; Acquisition of data: RM and JP; Analysis and interpretation of data: RM, JP, GG and DBG; Drafting of the article: GG and DBG; Critical revisions for intellectual content: GG and DBG; Final approval of the submitted version: GG, DBG, RM and JP.
Corresponding authors
Ethics declarations
Conflict of interest
R. Mahalakshmi, J. Priyanga, Dipita Bhakta-Guha and Gunjan Guha declare that there is no conflict of interest.
Ethical approval
Not applicable.
Consent to participate
Not applicable.
Consent for publication
Not applicable.
Additional information
Publisher's Note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Rights and permissions
About this article
Cite this article
Mahalakshmi, R., Priyanga, J., Bhakta-Guha, D. et al. Hormetic effect of low doses of rapamycin triggers anti-aging cascades in WRL-68 cells by modulating an mTOR-mitochondria cross-talk. Mol Biol Rep 49, 463–476 (2022). https://doi.org/10.1007/s11033-021-06898-6
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s11033-021-06898-6