Abstract
Background
Breast cancers exhibit genetic heterogeneity which causes differential responses to various chemotherapy agents. Given the unique demographic and genomic background in South Asia, genetic architecture in breast cancers is not fully explored.
Methods and results
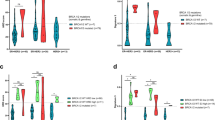
In this study, we determined the genetic landscape of our previously established luminal-A subtype breast cancer cell line (BC-PAK1), and compared it with a Caucasian origin MCF7 breast cancer cell line of the same molecular subtype. Deep whole-exome sequencing (100X) was performed from early passages of the primary cancer cells using the Illumina NextSeq500. Data analysis with in silico tools showed novel non-silent somatic mutations previously not described in breast cancers, including a frameshift insertion (p.Ala1591AlafsTer28) in CIC, and a frameshift deletion (p.Lys333LysfsTer21) in PABPC1. Five genes CDC27, PIK3CG, ARAP3, RAPGEF1, and EFNA3, related with cell cycle pathway (hsa04110), ErbB signaling pathway (hsa04012), Ras signaling pathway (hsa04014), and Rap1 signaling pathway (hsa04015) were found to have recurrent non-silent somatic mutations. Further, the major contribution of COSMIC signatures 3 (failure of DNA double-strand break repair by homologous recombination), and 12 (transcriptional strand-bias for T>C substitutions) was observed. Also, the somatic mutations landscape in BC-PAK1 was found to be different as compared to the MCF7 cell line. The unique genetic landscape of BC-PAK1 might be responsible for significantly reduced response to doxorubicin than the MCF7 cell line.
Conclusion
This study presents a distinct genetic architecture in luminal-A breast cancer potentially responsible for differential response to chemotherapy. Further studies on large cohorts of breast cancer patients are suggested for implementation in personalized medicine.




Similar content being viewed by others
Data availability
All the data presented herein has been included in the main manuscript and supplementary material.
Code availability
The bioinformatics tools/codes used in the analysis are freely available in the GitHub repository or at the tools’ websites.
References
Bray F, Ferlay J, Soerjomataram I, Siegel RL, Torre LA, Jemal A (2018) Global cancer statistics 2018: GLOBOCAN estimates of incidence and mortality worldwide for 36 cancers in 185 countries. CA: Cancer J Clin 68:394–424
Amir E, Freedman OC, Seruga B, Evans DG (2010) Assessing women at high risk of breast cancer: a review of risk assessment models. J Natl Cancer Inst 102:680–691
Gerlinger M, Rowan AJ, Horswell S, Math M, Larkin J et al (2012) Intratumor heterogeneity and branched evolution revealed by multiregion sequencing. N Engl J Med 366:883–892
Chin L, Hahn WC, Getz G, Meyerson M (2011) Making sense of cancer genomic data. Genes Dev 25:534–555
Hanahan D, Weinberg RA (2011) Hallmarks of cancer: the next generation. Cell 144:646–674
Schlabach MR, Luo J, Solimini NL, Hu G, Xu Q et al (2008) Cancer proliferation gene discovery through functional genomics. Science 319:620–624
Silva JM, Marran K, Parker JS, Silva J, Golding M et al (2008) Profiling essential genes in human mammary cells by multiplex RNAi screening. Science 319:617–620
Testa U, Castelli G, Pelosi E (2020) Breast cancer: a molecularly heterogenous disease needing subtype-specific treatments. Med Sci 8:18
Sorlie T, Tibshirani R, Parker J, Hastie T, Marron JS et al (2003) Repeated observation of breast tumor subtypes in independent gene expression data sets. Proc Natl Acad Sci USA 100:8418–8423
Dai X, Li T, Bai Z, Yang Y, Liu X et al (2015) Breast cancer intrinsic subtype classification, clinical use and future trends. Am J Cancer Res 5:2929–2943
Ferrari A, Vincent-Salomon A, Pivot X, Sertier A-S, Thomas E et al (2016) A whole-genome sequence and transcriptome perspective on HER2-positive breast cancers. Nat Commun 7:12222
Rakha EA, Pareja FG (2021) New advances in molecular breast cancer pathology. In: Proceedings of seminars in cancer biology, vol 72, pp 102–113. Elsevier, Amsterdam
Clusan L, Le Goff P, Flouriot G, Pakdel F (2021) A closer look at estrogen receptor mutations in breast cancer and their implications for estrogen and antiestrogen responses. Int J Mol Sci 22:756
Gillet JP, Varma S, Gottesman MM (2013) The clinical relevance of cancer cell lines. J Natl Cancer Inst 105:452–458
Ben-David U, Siranosian B, Ha G, Tang H, Oren Y et al (2018) Genetic and transcriptional evolution alters cancer cell line drug response. Nature 560:325–330
Begum N (2018) Breast cancer in Pakistan: a looming epidemic. J College Phys Surg 28:87–88
Malik SS, Baig M, Khan MB, Masood N (2019) Survival analysis of breast cancer patients with different treatments: a multicentric clinicopathological study. J Pak Med Assoc 69:976–980
Mughal AJ (2015) Establishment and characterization of human breast cancer cell line of Pakistani origin (thesis), University of Karachi, Karachi, Pakistan
Shakeel M, Irfan M, Khan IA (2018) Rare genetic mutations in Pakistani patients with dilated cardiomyopathy. Gene 673:134–139
Lek M, Karczewski KJ, Minikel EV, Samocha KE, Banks E et al (2016) Analysis of protein-coding genetic variation in 60,706 humans. Nature 536:285–291
Andrews S (2010) FastQC: a quality control tool for high throughput sequence data. Babraham Bioinformatics website: http://www.bioinformatics.bbsrc.ac.uk/projects/fastqc
Bolger AM, Lohse M, Usadel B (2014) Trimmomatic: a flexible trimmer for Illumina sequence data. Bioinformatics (Oxford, England) 30:2114–2120
Li H, Durbin R (2009) Fast and accurate short read alignment with Burrows-Wheeler transform. Bioinformatics 25:1754–1760
Li H, Handsaker B, Wysoker A, Fennell T, Ruan J et al (2009) The Sequence Alignment/Map format and SAMtools. Bioinformatics (Oxford, England) 25:2078–2079
McKenna A, Hanna M, Banks E, Sivachenko A, Cibulskis K et al (2010) The genome analysis Toolkit: a MapReduce framework for analyzing next-generation DNA sequencing data. Genome Res 20:1297–1303
Tyner JW, Tognon CE, Bottomly D, Wilmot B, Kurtz SE et al (2018) Functional genomic landscape of acute myeloid leukaemia. Nature 562:526–531
Yang H, Wang K (2015) Genomic variant annotation and prioritization with ANNOVAR and wANNOVAR. Nat Protoc 10:1556–1566
McLaren W, Gil L, Hunt SE, Riat HS, Ritchie GR et al (2016) The ensembl variant effect predictor. Genome Biol 17:1–14
Alexandrov LB, Nik-Zainal S, Wedge DC, Aparicio SA, Behjati S et al (2013) Signatures of mutational processes in human cancer. Nature 500:415–421
Landrum MJ, Lee JM, Riley GR, Jang W, Rubinstein WS et al (2014) ClinVar: public archive of relationships among sequence variation and human phenotype. Nucleic Acids Res 42:D980–D985
Hamosh A, Scott AF, Amberger JS, Bocchini CA, McKusick VA (2005) Online Mendelian Inheritance in Man (OMIM), a knowledgebase of human genes and genetic disorders. Nucleic Acids Res 33:D514–D517
Blokzijl F, Janssen R, van Boxtel R, Cuppen E (2018) Mutational patterns: comprehensive genome-wide analysis of mutational processes. Genome Med 10:33
Mularoni L, Sabarinathan R, Deu-Pons J, Gonzalez-Perez A, López-Bigas N (2016) OncodriveFML: a general framework to identify coding and non-coding regions with cancer driver mutations. Genome Biol 17:128
Heinrich V, Kamphans T, Stange J, Parkhomchuk D, Hecht J et al (2013) Estimating exome genotyping accuracy by comparing to data from large scale sequencing projects. Genome Med 5:69
Samuels DC, Han L, Li J, Quanghu S, Clark TA et al (2013) Finding the lost treasures in exome sequencing data. Trends Genet 29:593–599
Zhang X, Lin H, Zhao H, Hao Y, Mort M et al (2014) Impact of human pathogenic micro-insertions and micro-deletions on post-transcriptional regulation. Hum Mol Genet 23:3024–3034
Genomes Project Consortium (2015) A global reference for human genetic variation. Nature 526:68–74
Huang DW, Sherman BT, Tan Q, Collins JR, Alvord WG et al (2007) The DAVID Gene Functional Classification Tool: a novel biological module-centric algorithm to functionally analyze large gene lists. Genome Biol 8:R183–R183
Alexandrov LB, Nik-Zainal S, Wedge DC, Campbell PJ, Stratton MR (2013) Deciphering signatures of mutational processes operative in human cancer. Cell Rep 3:246–259
Thorn CF, Oshiro C, Marsh S, Hernandez-Boussard T, McLeod H et al (2011) Doxorubicin pathways: pharmacodynamics and adverse effects. Pharmacogenet Genom 21:440–446
Nik-Zainal S, Alexandrov LB, Wedge DC, Van Loo P, Greenman CD et al (2012) Mutational processes molding the genomes of 21 breast cancers. Cell 149:979–993
Alexandrov LB, Jones PH, Wedge DC, Sale JE, Campbell PJ et al (2015) Clock-like mutational processes in human somatic cells. Nat Genet 47:1402–1407
Jarvis MC, Ebrahimi D, Temiz NA, Harris RS (2018) Mutation signatures including APOBEC in cancer cell lines. JNCI Cancer Spectr 2:002
GeneCards (2019) GeneCards, The human gene database (PABPC1 Gene (Protein Coding)). https://www.genecards.org/cgi-bin/carddisp.pl?gene=PABPC1
GeneCards (2019) GeneCards, The human gene database (KMT2C Gene (Protein Coding)). https://www.genecards.org/cgi-bin/carddisp.pl?gene=KMT2C
Pareja F, Lee JY, Brown DN, Piscuoglio S, Gularte-Merida R et al (2019) The genomic landscape of mucinous breast cancer. J Natl Cancer Inst 111:216
Funding
The study was performed with funds granted to the institute, Dr. Panjwani Center for Molecular Medicine and Drug Research, University of Karachi, Karachi-75270, Pakistan, by the Searle Pharmaceuticals Pakistan Ltd. The funding agency had no role in the study design and presentation of the results.
Author information
Authors and Affiliations
Contributions
MS: performed NGS experimental work, bioinformatics analysis, data visualization, and manuscript preparation; SAK: conceived the study, performed experimental work, and manuscript preparation; AJM: performed experimental work to establish the BC-PAK1 cell line; MI: Performed data analysis, DCH: supervised in the establishment of the cell line, intellectual insights, and manuscript review; MIC: overall supervised the study, intellectual insights, and manuscript review; MA: performed DNA extraction from the cultured BC-PAK1 cells; IAK: intellectual insights, and manuscript review.
Corresponding authors
Ethics declarations
Conflict of interest
The authors declare that there is no conflict of interest.
Additional information
Publisher's Note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary Information
Below is the link to the electronic supplementary material.
Rights and permissions
About this article
Cite this article
Shakeel, M., Khan, S.A., Mughal, A.J. et al. Distinct genetic landscape and a low response to doxorubicin in a luminal-A breast cancer cell line of Pakistani origin. Mol Biol Rep 48, 6821–6829 (2021). https://doi.org/10.1007/s11033-021-06681-7
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s11033-021-06681-7