Abstract
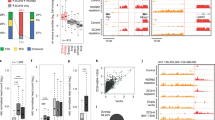
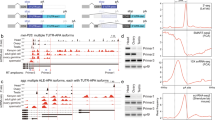
The termination of transcription is a complex process that substantially contributes to gene regulation in eukaryotes. Previously, it was noted that a single cytosine deletion at the position + 32 bp relative to the single polyadenylation signal AAUAAA (hereafter the dC mutation) causes a 2-fold increase in the transcription level of the upstream eGFP reporter in mouse embryonic stem cells. Here, we analyzed the conservation of this phenomenon in immortalized mouse, human and drosophila cell lines and the influence of the dC mutation on the choice of the pre-mRNA cleavage sites. We have constructed dual-reporter plasmids to accurately measure the effect of the dC and other nearby located mutations on eGFP mRNA level by RT-qPCR. In this way, we found that the dC mutation leads to a 2-fold increase in the expression level of the upstream eGFP reporter gene in cultured mouse and human, but not in drosophila cells. In addition, 3′ RACE analysis demonstrated that eGFP pre-mRNAs are cut at multiple positions between + 14 to + 31, and that the most proximal cleavage site becomes almost exclusively utilized in the presence of the dC mutation. We also identified new short sequence variations located within positions + 25.. + 40 and + 33.. + 48 that increase eGFP expression up to ~2-4-fold. Altogether, the positive effect of the dC mutation seems to be conserved in mouse embryonic stem cells, mouse embryonic 3T3 fibroblasts and human HEK293T cells. In the latter cells, the dC mutation appears to be involved in regulating pre-mRNA cleavage site selection. Finally, a multiplexed approach is proposed to identify motifs located downstream of cleavage site(s) that are essential for transcription termination.



Similar content being viewed by others
Data availability statement
All supporting data are included in the additional files. The materials generated during the current study are available upon request.
Data deposition
The complete nucleotide sequences of the plasmids pTTC-Dmel-WT, pTTC-Hsap-WT and pTTC-Mmus-WT were deposited in GenBank (accession numbers MN232102-MN232104).
References
Eisenberg E, Levanon EY (2013) Human housekeeping genes, revisited. Trends Genet 29(10):569–574
Hounkpe BW et al (2021) HRT Atlas v1.0 database: redefining human and mouse housekeeping genes and candidate reference transcripts by mining massive RNA-seq datasets. Nucleic Acids Res 49(D1):D947–D955
Mayr C, Bartel DP (2009) Widespread shortening of 3′UTRs by alternative cleavage and polyadenylation activates oncogenes in cancer cells. Cell 138(4):673–684
Elkon R, Ugalde AP, Agami R (2013) Alternative cleavage and polyadenylation: extent, regulation and function. Nat Rev Genet 14(7):496–506
Rehfeld A et al (2013) Alterations in polyadenylation and its implications for endocrine disease. Front Endocrinol (Lausanne) 4:53
Curinha A et al (2014) Implications of polyadenylation in health and disease. Nucleus 5(6):508–519
Hollerer I et al (2014) mRNA 3′end processing: a tale of the tail reaches the clinic. EMBO Mol Med 6(1):16–26
Masamha CP et al (2014) CFIm25 links alternative polyadenylation to glioblastoma tumour suppression. Nature 510(7505):412–416
Ogorodnikov A, Kargapolova Y, Danckwardt S (2016) Processing and transcriptome expansion at the mRNA 3′ end in health and disease: finding the right end. Pflugers Arch 468(6):993–1012
Proudfoot NJ (2016) Transcriptional termination in mammals: stopping the RNA polymerase II juggernaut. Science 352(6291):aad9926
Neve J et al (2017) Cleavage and polyadenylation: ending the message expands gene regulation. RNA Biol 14(7):865–890
Manning KS, Cooper TA (2017) The roles of RNA processing in translating genotype to phenotype. Nat Rev Mol Cell Biol 18(2):102–114
Veraldi KL et al (2000) Isolation and characterization of polyadenylation complexes assembled in vitro. RNA 6(5):768–777
Shi Y et al (2009) Molecular architecture of the human pre-mRNA 3′ processing complex. Mol Cell 33(3):365–376
Vallejos Baier R, Picao-Osorio J, Alonso CR (2017) Molecular regulation of alternative polyadenylation (APA) within the Drosophila nervous system. J Mol Biol 429(21):3290–3300
Hsin J-P, Manley JL (2012) The RNA polymerase II CTD coordinates transcription and RNA processing. Genes Dev 26(19):2119–2137
Andersen PK, Jensen TH, Lykke-Andersen S (2013) Making ends meet: coordination between RNA 3′-end processing and transcription initiation. Wiley Interdiscip Rev RNA 4(3):233–246
Fusby B et al (2016) Coordination of RNA polymerase II pausing and 3′ end processing factor recruitment with alternative polyadenylation. Mol Cell Biol 36(2):295–303
Tian B, Graber JH (2012) Signals for pre-mRNA cleavage and polyadenylation. Wiley Interdiscip Rev RNA 3(3):385–396
Shi Y, Manley JL (2015) The end of the message: multiple protein–RNA interactions define the mRNA polyadenylation site. Genes Dev 29(9):889–897
Gruber AJ et al (2016) A comprehensive analysis of 3′ end sequencing data sets reveals novel polyadenylation signals and the repressive role of heterogeneous ribonucleoprotein C on cleavage and polyadenylation. Genome Res 26(8):1145–1159
Gilmartin GM, Nevins JR (1991) Molecular analyses of two poly(A) site-processing factors that determine the recognition and efficiency of cleavage of the pre-mRNA. Mol Cell Biol 11(5):2432–2438
Brown KM, Gilmartin GM (2003) A mechanism for the regulation of pre-mRNA 3′ processing by human cleavage factor Im. Mol Cell 12(6):1467–1476
Yang Q, Gilmartin GM, Doublié S (2011) The structure of human cleavage factor Im hints at functions beyond UGUA-specific RNA binding: a role in alternative polyadenylation and a potential link to 5′ capping and splicing. RNA Biol 8(5):748–753
Beaudoing E et al (2000) Patterns of variant polyadenylation signal usage in human genes. Genome Res 10(7):1001–1010
Di Giammartino DC, Nishida K, Manley JL (2011) Mechanisms and consequences of alternative polyadenylation. Mol Cell 43(6):853–866
Shi Y (2012) Alternative polyadenylation: new insights from global analyses. RNA 18(12):2105–2117
Davis R, Shi Y (2014) The polyadenylation code: a unified model for the regulation of mRNA alternative polyadenylation. J Zhejiang Univ Sci B 15(5):429–437
Gruber AR et al (2014) Means to an end: mechanisms of alternative polyadenylation of messenger RNA precursors. Wiley Interdiscip Rev RNA 5(2):183–196
Tian B, Manley JL (2017) Alternative polyadenylation of mRNA precursors. Nat Rev Mol Cell Biol 18(1):18–30
Tian B et al (2005) A large-scale analysis of mRNA polyadenylation of human and mouse genes. Nucleic Acids Res 33(1):201–212
Xu C, Zhang J (2018) Alternative polyadenylation of mammalian transcripts is generally deleterious, not adaptive. Cell Syst 6(6):734–742
Fu Y et al (2018) Crosstalk between alternative polyadenylation and miRNAs in the regulation of protein translational efficiency. Genome Res 28(11):1656–1663
Wahle E, Rüegsegger U (1999) 3′-End processing of pre-mRNA in eukaryotes. FEMS Microbiol Rev 23(3):277–295
Zhang X, Virtanen A, Kleiman FE (2010) To polyadenylate or to deadenylate: that is the question. Cell Cycle 9(22):4437–4449
Gotic I, Schibler U (2012) The ticking tail: daily oscillations in mRNA poly(A) tail length drive circadian cycles in protein synthesis. Genes Dev 26(24):2669–2672
Akhtar W et al (2013) Chromatin position effects assayed by thousands of reporters integrated in parallel. Cell 154(4):914–927
Scotto-Lavino E, Du G, Frohman MA (2006) 3′ End cDNA amplification using classic RACE. Nat Protoc 1(6):2742–2745
Handler AM et al (1998) The lepidopteran transposon vector, piggyBac, mediates germ-line transformation in the Mediterranean fruit fly. Proc Natl Acad Sci U S A 95(13):7520–7525
Handler AM, Harrell RA II (1999) Germline transformation of Drosophila melanogaster with the piggyBac transposon vector. Insect Mol Biol 8(4):449–457
Shen D et al (2018) Enhancer trapping and annotation in zebrafish mediated with Sleeping Beauty, piggyBac and Tol2 transposons. Genes (Basel) 9(12):630
Marh J et al (2012) Hyperactive self-inactivating piggyBac for transposase-enhanced pronuclear microinjection transgenesis. Proc Natl Acad Sci U S A 109(47):19184–19189
Meir Y-J et al (2011) Genome-wide target profiling of piggyBac and Tol2 in HEK 293: pros and cons for gene discovery and gene therapy. BMC Biotechnol 11:28
Ochman H, Gerber AS, Hartl DL (1988) Genetic applications of an inverse polymerase chain reaction. Genetics 120(3):621–623
Cary LC et al (1989) Transposon mutagenesis of baculoviruses: analysis of Trichoplusia ni transposon IFP2 insertions within the FP-locus of nuclear polyhedrosis viruses. Virology 172(1):156–169
Salisbury J, Hutchison KW, Graber JH (2006) A multispecies comparison of the metazoan 3′-processing downstream elements and the CstF-64 RNA recognition motif. BMC Genomics 7:55
Pauws E et al (2001) Heterogeneity in polyadenylation cleavage sites in mammalian mRNA sequences: implications for SAGE analysis. Nucleic Acids Res 29(8):1690–1694
You L et al (2015) APASdb: a database describing alternative poly(A) sites and selection of heterogeneous cleavage sites downstream of poly(A) signals. Nucleic Acids Res 43(Database issue):D59–D67
Sheets MD, Ogg SC, Wickens MP (1990) Point mutations in AAUAAA and the poly (A) addition site: effects on the accuracy and efficiency of cleavage and polyadenylation in vitro. Nucleic Acids Res 18(19):5799–5805
Derti A et al (2012) A quantitative atlas of polyadenylation in five mammals. Genome Res 22(6):1173–1183
White E et al (2013) AT-rich sequence elements promote nascent transcript cleavage leading to RNA polymerase II termination. Nucleic Acids Res 41(3):1797–1806
Baejen C et al (2017) Genome-wide analysis of RNA polymerase II termination at protein-coding genes. Mol Cell 66(1):38–49
Schwalb B et al (2016) TT-seq maps the human transient transcriptome. Science 352(6290):1225–1228
Bogard N et al (2019) A deep neural network for predicting and engineering alternative polyadenylation. Cell 178(1):91–106
Haberle V, Lenhard B (2012) Dissecting genomic regulatory elements in vivo. Nat Biotechnol 30(6):504–506
Levo M, Segal E (2014) In pursuit of design principles of regulatory sequences. Nat Rev Genet 15(7):453–468
Ho SN et al (1989) Site-directed mutagenesis by overlap extension using the polymerase chain reaction. Gene 77(1):51–59
Kozhevnikova EN, Leshchenko AE, Pindyurin AV (2018) An inducible DamID system for profiling interactions of nuclear lamina protein component Lamin B1 with chromosomes in mouse cells. Biochemistry (Mosc) 83(5):586–594
Yang X et al (2013) Drosophila Vps36 regulates Smo trafficking in Hedgehog signaling. J Cell Sci 126(Pt 18):4230–4238
Litvinova EA et al (2018) Role of the Kaiso gene in the development of inflammation in Mucin-2 deficient mice. Vavilov J Genet Breed 22(8):1078–1083
Acknowledgments
We thank Bas van Steensel for providing Kc167 cells, Evgeniya N. Andreyeva and Tatiana D. Dubatolova for technical assistance with plasmid construction, Tatyana N. Belovezhets, Sergey V. Kulemzin and Evdokiya S. Reshetnikova for assistance with FACS, and Evgeniya N. Andreyeva for helpful suggestions. DNA sequencing, FACS and qPCR analyses were performed using resources provided by the Molecular and Cellular Biology core facility of the Institute of Molecular and Cellular Biology SB RAS.
Funding
This work was supported by the grant of Russian Science Foundation no. 16-14-10288.
Author information
Authors and Affiliations
Corresponding author
Ethics declarations
Conflict of interest
The authors declare that they have no conflict of interest.
Ethical approval
This article does not contain any studies with human participants or animals performed by any of the authors.
Additional information
Publisher's Note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary information
Below is the link to the electronic supplementary material.
Rights and permissions
About this article
Cite this article
Boldyreva, L.V., Yarinich, L.A., Kozhevnikova, E.N. et al. Fine gene expression regulation by minor sequence variations downstream of the polyadenylation signal. Mol Biol Rep 48, 1539–1547 (2021). https://doi.org/10.1007/s11033-021-06160-z
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s11033-021-06160-z