Abstract
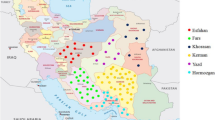
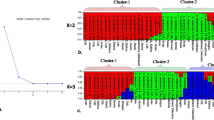
Assessment of genetic diversity is crucial for efficient selection genotypes in plant breeding and improvement programs. Studies of genetic diversity of S. persica are rare relative to the large species diversity in Saudi Arabia, despite its valuable importance as one of the most popular medicinal plants. We investigate the genetic variability and genetic differentiation among and within wild Salvadora persica populations distributed in four regions of Saudi Arabia. Twelve sequence-related amplified polymorphism (SRAP) primers combination generated 326 alleles, with an average of 27.2 alleles per primer. All primers showed 100 polymorphism percentage, and higher PIC values exceeded 0.90. Jaccard similarity values varied between 0.04 to 0.43, with an average of 0.31, which showed a weak relationship among the accessions and their origin. Based on UGPMA and principal coordinate analysis, accessions collected from the same region showed less aggregation. Genetic diversity parameters showed that both Aflaj and Joodah populations recorded the highest mean values for the effective number of alleles (1.26 and 1.24). Shannon index and genetic heterozygosity (0.23 and 0.15 for both populations), and percent of polymorphism 45.45% for Aflaj and 43.87 for Joodah population. Most of the genetic variation was because of differences within populations (77%) and 23% among populations. SRAP markers explored the genetic diversity among and within S. persica populations. In this work, genetic diversity within populations was high, and the population structure was weak. We detected no specific geographic structure, which may reveal an active movement of plants among populations.


Similar content being viewed by others
References
Ezmirly ST, Cheng JC, Wilson SR (1979) Saudi Arabia medicinal plants: Salvadora persica L. Planta Medica 35:191–192
Mathur S, Shekhawat GS, Batra A (2002) Micropropagation of Salvadora persica L via cotyledonary nodes. Indian. J Biotechnol 1:197–200
Murphy DJ (1995) New oil for old chemistry in Britain 31(4): 300-30.2
Sujata M (2015) Medicinally potent and highly salt tolerant plant of arid zone-Salvadora persica L (Meswak). A Rev J Plant Sci 3(1):45–49
Sharma DK, Shah KR, Dave RS (2018) A review on the pharmagnostic evaluation of Meswak. An International Peer Reviewed Open Access Journal for Rapid Publication, Salvadora persica, p 734
Monfared MA, Samsampour D, Sharifi-Sirchi GR, Sadeghi F (2018) Assessment of genetic diversity in Salvadora persica L based on inter simple sequence repeat (ISSR) genetic marker. J Genet Eng Biotechnol 16(2):661–667
Kumari S, Yadav K, Singh N (2017) Evaluation of genetic fidelity among micropropagated plants of Salvadora persica L using DNA-based markers. Meta Gene 14:129–33
Upendra JM, Rao SR, Dagla HR (2017) Genetic diversity analysis of Salvadora persica: an evergreen halo-xeric species of semi-arid and sub-humid regions of Rajasthan. India Ecol Genet Genom 2:35–41
Li G, Quiros CF (2001) Sequence-related amplified polymorphism (SRAP): a new marker system based on a simple PCR reaction, Its application to mapping and gene tagging in brassica. Theor Appl Genet 103:455–461
Anderson J, Churchill G, Autrique J, Tanksley S, Sorrells M (1993) Optimizing parental selection for genetic linkage maps. Genome 36(1):181–186
Jaccard P (1908) Nouvelles recherches sur la distribution florale. Bulletin de la Société vaudoise des sciences naturelles 44:223–270
Hammer Ø (2009) PAST Paleontological Statistics Reference manual Version 196. University of Oslo, Oslo, Natural History Museum
Peakall R, Smouse PG (2012) GenAlEx 6.5: genetic analysis in Excel. Population genetic software for teaching and research-an update. Bioinformatics 28:2537–2539
Hopley T, Byrne M (2019) Gene flow and genetic variation explain signatures of selection across a climate gradient in two riparian species. Genes 10(8):579–594
Saini S, Yadav JP (2013) Genetic variation in natural populations of Salvadora oleoides L: An important medicinal plant that needs conservation. Pelagia Res Lib 3:20–27
Yadav JP, Sandeep Kumar S, Yadav M, Kadyan S, Yadav S (2014) Assessment of genetic diversity using RAPD marker among different accessions of Salvadora oleoides of North-West India Bioresearch. Bulletin 4(1):1–7
Hegazi GA, El-Hanafy NA, Ahmed SE, Abu-Elkheir ZA, Hussein IA (2011) Genetic diversity and in vitro propagation of Salvadora persica L. Arab J Biotechnol 14:253–268
Hepsibha BT, Premalakshmi V, Sekar T (2010) Genetic diversity in Azima tetracantha (Lam) assessed through RAPD analysis. Indian J Sci Technol 3(2):170–173
Funding
This study did not receive funding from any organization.
Author information
Authors and Affiliations
Contributions
Conceptualization: AIA, and AMA. Data acquisition: MHM, and AAM. Design of method: AIA, and AAM. Writing and editing: MHM, and AAM. All authors reviewed, edited, and approved the final version of the manuscript.
Corresponding author
Ethics declarations
Conflict of interest
The authors declare that they have no conflict of interest.
Ethical approval
This manuscript does not imply human participants or studies on animals.
Additional information
Publisher's Note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Rights and permissions
About this article
Cite this article
ALmohisen, I.A., AL-muwayhi, M.A., Assaeed, A.M. et al. Evaluation of the genetic diversity of wild Salvadora persica ‘Arak’ from Saudi Arabia. Mol Biol Rep 47, 7843–7849 (2020). https://doi.org/10.1007/s11033-020-05860-2
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s11033-020-05860-2