Abstract
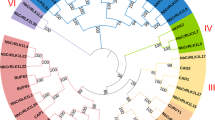
The receptor like kinases (RLKs) belong to the RLK/Pelle superfamily, one of the largest gene families in plants. RLKs play an important role in plant development, as well as in response to biotic and abiotic stresses. The lysine motif receptor like kinases (LysM-RLKs) are a subfamily of RLKs containing at least one lysine motif (LysM) that are involved in the perception of elicitors or pathogen-associated molecular patterns (PAMPs). In the present study, 77 putative RLKs genes and three receptor like proteins were identified in potato (Solanum tuberosum) genome, following a genome-wide search. The 77 potato RLK proteins are classified into two major phylogenetic groups based on their kinase domain amino acid sequence similarities. Out of 77 RLKs, 10 proteins had at least one LysM. Among them three RLP proteins were found in potato genome with either 2 or three tandem LysM but these lacked a cytoplasmic kinase domain. Expression analyses of a potato LysM-RLKs (StLysM-RLK05) was carried out by a Real time RT-PCR, following inoculation of potato leaves and immature tubers with late blight and common scab pathogens, respectively. The expression was significantly higher in resistant than in susceptible following S. scabies inoculation. The StLysM-RLK05 sequence was verified and it was polymorphic in scab susceptible cultivar. The present study provides an overview of the StLysM-RLKs gene family in potato genome. This information is helpful for future functional analysis of such an important protein family, in Solanaceae species.







Similar content being viewed by others
Abbreviations
- MAPK:
-
Mitogen activated protein kinases
- PAMPs:
-
Pathogen-associated molecular patterns
- RKs:
-
Receptor kinases
- RLKs:
-
Receptor-like kinases
- CEBiP:
-
Chitin-elicitor binding protein
- AUDPC:
-
Area under the disease progress curve
- PDB:
-
Protein Data Bank
References
Dangl JL, Jones JD (2001) Plant pathogens and integrated defence responses to infection. Nature 411:826–833
Kushalappa AC, Yogendra KN, Karre S (2016) Plant innate immune response: qualitative and quantitative resistance. Crit Rev Plant Sci 35:38–55
Cao Y, Liang Y, Tanaka K, Nguyen CT, Jedrzejczak RP, Joachimiak A, Stacey G (2014) The kinase LYK5 is a major chitin receptor in Arabidopsis and forms a chitin-induced complex with related kinase CERK1. Elife. https://doi.org/10.7554/eLife.03766
Zipfel C (2008) Pattern-recognition receptors in plant innate immunity. Curr Opin Immunol 20:10–16
Torii KU (2009) Transmembrane receptors in plants: receptor kinases and their ligands. Annu Plant Rev 33:1–29
Kanda Y, Yokotani N, Maeda S, Nishizawa Y, Kamakura T, Mori M (2017) The receptor-like cytoplasmic kinase BSR1 mediates chitin-induced defense signaling in rice cells. Biosci Biotechnol Biochem 81(8):1497–1500
Shiu S-H, Bleecker AB (2001) Plant receptor-like kinase gene family: diversity, function, and signaling. Sci STKE 2001:re22
Shiu S-H, Bleecker AB (2001) Receptor-like kinases from Arabidopsis form a monophyletic gene family related to animal receptor kinases. Proc Natl Acad Sci USA 98:10763–10768
Shiu S-H, Bleecker AB (2001) Plant receptor-like kinase gene family: diversity, function, and signaling. Sci STKE 113:re22
Gómez-Gómez L, Boller T (2000) FLS2: an LRR receptor-like kinase involved in the perception of the bacterial elicitor flagellin in Arabidopsis. Mol Cell 5:1003–1011
Li J, Chory J (1997) A putative leucine-rich repeat receptor kinase involved in brassinosteroid signal transduction. Cell 90:929–938
Clark SE, Williams RW, Meyerowitz EM (1997) The CLAVATA1gene encodes a putative receptor kinase that controls shoot and floral meristem size in Arabidopsis. Cell 89:575–585
Kaku H, Shibuya N (2016) Molecular mechanisms of chitin recognition and immune signaling by LysM-receptors. Physiol Mol Plant Pathol 95:60–65
Zhang X-C, Wu X, Findley S, Wan J, Libault M, Nguyen HT, Cannon SB, Stacey G (2007) Molecular evolution of lysin motif-type receptor-like kinases in plants. Plant Physiol 144:623–636
Buist G, Steen A, Kok J, Kuipers OP (2008) LysM, a widely distributed protein motif for binding to (peptido) glycans. Mol Microbiol 68:838–847
Hohmann U, Lau K, Hothorn M (2017) The structural basis of ligand perception and signal activation by receptor kinases. Annu Rev Plant Biol 68:109–137
Faulkner C, Petutschnig E, Benitez-Alfonso Y, Beck M, Robatzek S, Lipka V, Maule AJ (2013) LYM2-dependent chitin perception limits molecular flux via plasmodesmata. Proc Natl Acad Sci USA 110:9166–9170
Kaku H, Nishizawa Y, Ishii-Minami N, Akimoto-Tomiyama C, Dohmae N, Takio K, Minami E, Shibuya N (2006) Plant cells recognize chitin fragments for defense signaling through a plasma membrane receptor. Proc Natl Acad Sci USA 103:11086–11091
Liu B, Li JF, Ao Y, Qu J, Li Z, Su J, Zhang Y, Liu J, Feng D, Qi K, He Y, Wang J, Wang HB (2012) Lysin motif-containing proteins LYP4 and LYP6 play dual roles in peptidoglycan and chitin perception in rice innate immunity. Plant Cell 24:3406–3419
Wan J, Tanaka K, Zhang XC, Son GH, Brechenmacher L, Nguyen TH, Stacey G (2012) LYK4, a lysin motif receptor-like kinase, is important for chitin signaling and plant innate immunity in Arabidopsis. Plant Physiol 160:396–406
Espinoza C, Liang Y, Stacey G (2017) Chitin receptor CERK1 links salt stress and chitin-triggered innate immunity in Arabidopsis. Plant J 89:984–995
Carotenuto G, Chabaud M, Miyata K, Capozzi M, Takeda N, Kaku H, Shibuya N, Nakagawa T, Barker DG, Genre A (2017) The rice LysM receptor-like kinase OsCERK1 is required for the perception of short-chain chitin oligomers in arbuscular mycorrhizal signaling. New Phytol 214:1440–1446
Karre S, Kumar A, Dhokane D, Kushalappa AC (2017) Metabolo-transcriptome profiling of barley reveals induction of chitin elicitor receptor kinase gene (HvCERK1) conferring resistance against Fusarium graminearum. Plant Mol Biol 93:247–267
Bethke PC, Nassar AM, Kubow S, Leclerc YN, Li X-Q, Haroon M, Molen T, Bamberg J, Martin M, Donnelly DJ (2014) History and origin of Russet Burbank (Netted Gem) a sport of Burbank. Am J Potato Res 91:594–609
Yogendra KN, Kushalappa AC, Sarmiento F, Rodriguez E, Mosquera T (2015) Metabolomics deciphers quantitative resistance mechanisms in diploid potato clones against late blight. Funct Plant Biol 42:284–298
Pushpa D, Yogendra KN, Gunnaiah R, Kushalappa AC, Murphy A (2014) Identification of late blight resistance-related metabolites and genes in potato through nontargeted metabolomics. Plant Mol Biol Report 32:584–595
Yogendra KN, Kushalappa AC (2016) Integrated transcriptomics and metabolomics reveal induction of hierarchies of resistance genes in potato against late blight. Funct Plant Biol 43:766–782
Shirling ET, Gottlieb D (1966) Methods for characterization of Streptomyces species. Int J Syst Bacteriol 16:313–340
Goodstein DM, Shu S, Howson R, Neupane R, Hayes RD, Fazo J, Mitros T, Dirks W, Hellsten U, Putnam N (2011) Phytozome: a comparative platform for green plant genomics. Nucleic Acids Res 40:D1178–D1186
Miya A, Albert P, Shinya T, Desaki Y, Ichimura K, Shirasu K, Narusaka Y, Kawakami N, Kaku H, Shibuya N (2007) CERK1, a LysM receptor kinase, is essential for chitin elicitor signaling in Arabidopsis. Proc Natl Acad Sci USA 104:19613–19618
Dhaliwal AK, Mohan A, Gill KS (2014) Comparative analysis of ABCB1 reveals novel structural and functional conservation between monocots and dicots. Front Firm Knowl Creat Entity. https://doi.org/10.3389/fpls.2014.00657
Voorrips R (2002) MapChart: software for the graphical presentation of linkage maps and QTLs. J Hered 93:77–78
Kumar S, Stecher G, Tamura K (2016) MEGA7: molecular evolutionary genetics analysis version 7.0 for bigger datasets. Mol Biol Evol 33:1870–1874
Livak KJ, Schmittgen TD (2001) Analysis of relative gene expression data using real-time quantitative PCR and the 2−ΔΔCT method. Methods 25:402–408
Zimmermann L, Stephens A, Nam S-Z, Rau D, Kübler J, Lozajic M, Gabler F, Söding J, Lupas AN, Alva V (2017) A completely reimplemented mpi bioinformatics toolkit with a new HHpred server at its core. J Mol Biol 430(15):2237–2243
Webb B, Sali A (2016) Comparative protein structure modeling using MODELLER. Curr Protoc Protein Sci 86:2.9.1–2.9.37
Irwin JJ, Sterling T, Mysinger MM, Bolstad ES, Coleman RG (2012) ZINC: a free tool to discover chemistry for biology. J Chem Inf Model 52:1757–1768
Morris GM, Huey R, Lindstrom W, Sanner MF, Belew RK, Goodsell DS, Olson AJ (2009) AutoDock4 and AutoDockTools4: automated docking with selective receptor flexibility. J Comput Chem 30:2785–2791
Trott O, Olson AJ (2010) AutoDock Vina: improving the speed and accuracy of docking with a new scoring function, efficient optimization, and multithreading. J Comput Chem 31:455–461
Bouaziz D, Charfeddine M, Jbir R, Saidi MN, Pirrello J, Charfeddine S, Bouzayen M, Gargouri-Bouzid R (2015) Identification and functional characterization of ten AP2/ERF genes in potato. Plant Cell Tissue Organ Cult 123:155–172
Charfeddine M, Saïdi MN, Charfeddine S, Hammami A, Bouzid RG (2015) Genome-wide analysis and expression profiling of the ERF transcription factor family in potato (Solanum tuberosum L.). Mol Biotechnol 57:348–358
Gromadka R, Cieśla J, Olszak K, Szczegielniak J, Muszyńska G, Polkowska-Kowalczyk L (2018) Genome-wide analysis and expression profiling of calcium-dependent protein kinases in potato (Solanum tuberosum). Plant Growth Regul 84:303–315
Jupe F, Pritchard L, Etherington GJ, MacKenzie K, Cock PJ, Wright F, Sharma SK, Bolser D, Bryan GJ, Jones JD (2012) Identification and localisation of the NB-LRR gene family within the potato genome. BMC Genom 13:75
Ma H, Cao X, Shi S, Li S, Gao J, Ma Y, Zhao Q, Chen Q (2016) Genome-wide survey and expression analysis of the amino acid transporter superfamily in potato (Solanum tuberosum L.). Plant Physiol Biochem 107:164–177
Singh AK, Sharma V, Pal AK, Acharya V, Ahuja PS (2013) Genome-wide organization and expression profiling of the NAC transcription factor family in potato (Solanum tuberosum L.). DNA Res 20:403–423
Zhao P, Wang D, Wang R, Kong N, Zhang C, Yang C, Wu W, Ma H, Chen Q (2018) Genome-wide analysis of the potato Hsp20 gene family: identification, genomic organization and expression profiles in response to heat stress. BMC Genom 19:61
Schmutz J, Cannon SB, Schlueter J, Ma J, Mitros T, Nelson W, Hyten DL, Song Q, Thelen JJ, Cheng J (2010) Genome sequence of the palaeopolyploid soybean. Nature 463:178
Yang X, Tuskan GA (2006) Divergence of the Dof gene families in poplar, Arabidopsis, and rice suggests multiple modes of gene evolution after duplication. Plant Physiol 142:820–830
Consortium PGS (2011) Genome sequence and analysis of the tuber crop potato. Nature 475:189
Buendia L, Wang T, Girardin A, Lefebvre B (2016) The LysM receptor-like kinase SlLYK10 regulates the arbuscular mycorrhizal symbiosis in tomato. New Phytol 210:184–195
Zeng L, Velásquez AC, Munkvold KR, Zhang J, Martin GB (2012) A tomato LysM receptor-like kinase promotes immunity and its kinase activity is inhibited by AvrPtoB. Plant J 69:92–103
Zhang X-C, Cannon SB, Stacey G (2009) Evolutionary genomics of LysM genes in land plants. BMC Evol Biol 9:183
Smit P, Limpens E, Geurts R, Fedorova E, Dolgikh E, Gough C, Bisseling T (2007) Medicago LYK3, an entry receptor in rhizobial nodulation factor signaling. Plant Physiol 145:183–191
Radutoiu S, Madsen LH, Madsen EB, Jurkiewicz A, Fukai E, Quistgaard EM, Albrektsen AS, James EK, Thirup S, Stougaard J (2007) LysM domains mediate lipochitin–oligosaccharide recognition and Nfr genes extend the symbiotic host range. EMBO J 26(17):3923–3935
Lohmann GV, Shimoda Y, Nielsen MW, Jørgensen FG, Grossmann C, Sandal N, Sørensen K, Thirup S, Madsen LH, Tabata S (2010) Evolution and regulation of the Lotus japonicus LysM receptor gene family. Mol Plant Microbe Interact 23:510–521
Rairdan G, Moffett P (2007) Brothers in arms? Common and contrasting themes in pathogen perception by plant NB-LRR and animal NACHT-LRR proteins. Microbes Infect 9:677–686
Massa AN, Childs KL, Lin H, Bryan GJ, Giuliano G, Buell CR (2011) The transcriptome of the reference potato genome Solanum tuberosum Group Phureja clone DM1-3 516R44. PLoS ONE 6:e26801
Peng K-C, Wang C-W, Wu C-H, Huang C-T, Liou R-F (2015) Tomato SOBIR1/EVR homologs are involved in elicitin perception and plant defense against the oomycete pathogen Phytophthora parasitica. Mol Plant Microbe Interact 28:913–926
Wan J, Zhang X-C, Neece D, Ramonell KM, Clough S, Kim S-y, Stacey MG, Stacey G (2008) A LysM receptor-like kinase plays a critical role in chitin signaling and fungal resistance in Arabidopsis. Plant Cell 20:471–481
Kushalappa AC, Yogendra KN, Sarkar K, Kage U, Karre S (2016) Gene discovery and genome editing to develop cisgenic crops with improved resistance against pathogen infection. Can J Plant Path 38:279–295
Funding
This work was carried out with the aid of a grant from The Natural Sciences and Engineering Research Council of Canada (NSERC). SJ acknowledges MITACS for funding and Progest2001 Inc. for providing the genotype AG704.10.
Author information
Authors and Affiliations
Contributions
FN conducted the experiments, bioinformatics analyses and wrote the manuscript; SJ conducted inoculation studies and edited the manuscript; HX conducted scab inoculation studies; AK conceived the idea and edited the manuscript.
Corresponding author
Ethics declarations
Conflict of interest
Authors declare no conflicts of interest.
Additional information
Publisher's Note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Rights and permissions
About this article
Cite this article
Nazarian-Firouzabadi, F., Joshi, S., Xue, H. et al. Genome-wide in silico identification of LysM-RLK genes in potato (Solanum tuberosum L.). Mol Biol Rep 46, 5005–5017 (2019). https://doi.org/10.1007/s11033-019-04951-z
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s11033-019-04951-z