Abstract
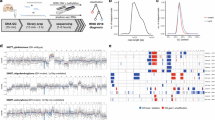
The MinION is a miniaturized high-throughput next generation sequencing platform of novel conception. The use of nucleic acids derived from formalin-fixed paraffin-embedded samples is highly desirable, but their adoption for molecular assays is hurdled by the high degree of fragmentation and by the chemical-induced mutations stemming from the fixation protocols. In order to investigate the suitability of MinION sequencing on formalin-fixed paraffin-embedded samples, the presence and frequency of BRAF c.1799T > A mutation was investigated in two archival tissue specimens of Hairy cell leukemia and Hairy cell leukemia Variant. Despite the poor quality of the starting DNA, BRAF mutation was successfully detected in the Hairy cell leukemia sample with around 50% of the reads obtained within 2 h of the sequencing start. Notably, the mutational burden of the Hairy cell leukemia sample as derived from nanopore sequencing proved to be comparable to a sensitive method for the detection of point mutations, namely the Digital PCR, using a validated assay. Nanopore sequencing can be adopted for targeted sequencing of genetic lesions on critical DNA samples such as those extracted from archival routine formalin-fixed paraffin-embedded samples. This result let speculating about the possibility that the nanopore sequencing could be trustably adopted for the real-time targeted sequencing of genetic lesions. Our report opens the window for the adoption of nanopore sequencing in molecular pathology for research and diagnostics.



Similar content being viewed by others
References
Norris AL, Workman RE, Fan Y, Eshleman JR, Timp W (2016) Nanopore sequencing detects structural variants in cancer. Cancer Biol Ther 17(3):246–253. https://doi.org/10.1080/15384047.2016.1139236
Wei S, Williams Z (2015) Rapid short-read sequencing and aneuploidy detection using MinION nanopore technology. Genetics 202(1):37–44. https://doi.org/10.1534/genetics.115.182311
Ip CL, Loose M, Tyson JR, de Cesare M, Brown BL, Jain M, Leggett RM, Eccles DA, Zalunin V, Urban JM, Piazza P, Bowden RJ, Paten B, Mwaigwisya S, Batty EM, Simpson JT, Snutch TP, Birney E, Buck D, Goodwin S, Jansen HJ, O’Grady J, Olsen HE (2015) MinION analysis and reference consortium. MinION analysis and reference consortium: phase 1 data release and analysis. F1000 Res 4:1075. https://doi.org/10.12688/f1000research.7201.1
Cherukuri Y, Janga SC (2016) Benchmarking of de novo assembly algorithms for nanopore data reveals optimal performance of OLC approaches. BMC Genomics 17(Suppl 7):507. https://doi.org/10.1186/s12864-016-2895-8
Hedegaard J, Thorsen K, Lund MK, Hein AM, Hamilton-Dutoit SJ, Vang S, Nordentoft I, Birkenkamp-Demtröder K, Kruhøffer M, Hager H, Knudsen B, Andersen CL, Sørensen KD, Pedersen JS, Ørntoft TF, Dyrskjøt L (2014) Next-generation sequencing of RNA and DNA isolated from paired fresh-frozen and formalin-fixed paraffin-embedded samples of human cancer and normal tissue. PLoS ONE 9(5):e98187. https://doi.org/10.1371/journal.pone.0098187
Swerdlow SH, Campo E, Harris NL, et al (2008) WHO classification of tumors of haematopoietic and lymphoid tissues. In: Bosman FT, Jaffe ES, Lakhani SR, OHgaki H (eds), World Health Organization Classification of tumors, Lyon, pp 188–193
Swerdlow SH, Campo E, Pileri SA, Harris NL, Stein H, Siebert R, Advani R, Ghielmini M, Salles GA, Zelenetz AD, Jaffe ES (2016) The 2016 revision of the World Health Organization classification of lymphoid neoplasms. Blood 127(20):2375–2390. https://doi.org/10.1182/blood-2016-01-643569
Tiacci E, Trifonov V, Schiavoni G, Holmes A, Kern W, Martelli MP, Pucciarini A, Bigerna B, Pacini R, Wells VA, Sportoletti P, Pettirossi V, Mannucci R, Elliott O, Liso A, Ambrosetti A, Pulsoni A, Forconi F, Trentin L, Semenzato G, Inghirami G, Capponi M, Di Raimondo F, Patti C, Arcaini L, Musto P, Pileri S, Haferlach C, Schnittger S, Pizzolo G, Foà R, Farinelli L, Haferlach T, Pasqualucci L, Rabadan R, Falini B (2011) BRAF mutations in hairy-cell leukemia. N Engl J Med 364(24):2305–2315. https://doi.org/10.1056/NEJMoa1014209
Verma S, Greaves WO, Ravandi F, Reddy N, Bueso-Ramos CE, O’Brien S, Thomas DA, Kantarjian H, Medeiros LJ, Luthra R, Patel KP (2012) Rapid detection and quantitation of BRAF mutations in hairy cell leukemia using a sensitive pyrosequencing assay. Am J Clin Pathol 138(1):153–156. https://doi.org/10.1309/AJCPL0OPXI9LZITV
Vitali C, Bassani C, Chiodoni C, Fellini E, Guarnotta C. Miotti S, Sangaletti S, Fuligni F, De Cecco L, Piccaluga PP, Colombo MP, Tripodo C (2015) SOCS2 controls proliferation and stemness of hematopoietic cells under stress conditions and its deregulation marks unfavorable acute leukemias. Cancer Res 75(11):2387–2399. https://doi.org/10.1158/0008-5472.CAN-14-3625
Willmore-Payne C, Holden JA, Tripp S, Layfield LJ (2005) Human malignant melanoma: detection of BRAF- and c-kit-activating mutations by high-resolution amplicon melting analysis. Hum Pathol 36(5):486–493
Loman NJ, Quinlan AR (2014) Poretools: a toolkit for analyzing nanopore sequence data. Bioinformatics 30(23):3399–3401. https://doi.org/10.1093/bioinformatics/btu555
Afgan E, Baker D, van den Beek M, Blankenberg D, Bouvier D, Čech M, Chilton J, Clements D, Coraor N, Eberhard C, Grüning B, Guerler A, Hillman-Jackson J, Von Kuster G, Rasche E, Soranzo N, Turaga N, Taylor J, Nekrutenko A, Goecks J (2016) The Galaxy platform for accessible, reproducible and collaborative biomedical analyses: 2016 update. Nucleic Acids Res 44(W1):W3–W10. https://doi.org/10.1093/nar/gkw343
Lamy PJ, Castan F, Lozano N, Montélion C, Audran P, Bibeau F, Roques S, Montels F, Laberenne AC (2015) Next-generation genotyping by digital PCR to detect and quantify the BRAF V600E mutation in melanoma biopsies. J Mol Diagn 17(4):366–373. https://doi.org/10.1016/j.jmoldx.2015.02.004
Acknowledgements
This work was supported by AIRC (Italian Association for Cancer Research) [Grant Numbers 12162, 15999].
Author information
Authors and Affiliations
Corresponding author
Ethics declarations
Conflict of interest
The authors declare that there are no conflicts of interest.
Electronic supplementary material
Below is the link to the electronic supplementary material.

11033_2017_4133_MOESM1_ESM.jpg
Online resource 1 Cumulative 2D basecalled yield versus the sequencing time using DNA extracted from archival Formalin Fixed Paraffin Embedded (FFPE) specimens: tissue sample 1, of Hairy cell leukemia variant, HCL-v. (JPG 30 KB)

11033_2017_4133_MOESM2_ESM.jpg
Online resource 2 Cumulative 2D basecalled yield versus the sequencing time using DNA extracted from archival Formalin Fixed Paraffin Embedded (FFPE) specimens: tissue sample 2, of Hairy cell leukemia, HCL. (JPG 35 KB)

11033_2017_4133_MOESM3_ESM.jpg
Online resource 3 All sequences have a length of about 250 nucleotides with a mean quality score of about 8 for HCL-v. (JPG 31 KB)

11033_2017_4133_MOESM4_ESM.jpg
Online resource 4 All sequences have a length of about 250 nucleotides with a mean quality score of about 7 for HCL. (JPG 36 KB)



Rights and permissions
About this article
Cite this article
Vacca, D., Cancila, V., Gulino, A. et al. Real-time detection of BRAF V600E mutation from archival hairy cell leukemia FFPE tissue by nanopore sequencing. Mol Biol Rep 45, 1–7 (2018). https://doi.org/10.1007/s11033-017-4133-0
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s11033-017-4133-0