Abstract
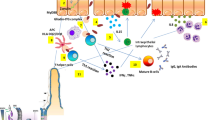
Celiac disease (CD) is the most common food-sensitive enteropathy in genetically susceptible individuals. The major genetic risk factors known are specific human leukocyte antigen (HLA)-DQ haplotypes, but other genetic factors are supposed to be involved. Interleukin-18 (IL-18) is a pro-inflammatory cytokine that has an important role in the immune defense and it has the potential to influence inflammatory disorders. IL-18 is able to promote Th1 cell development and it is expressed in the mucosa of the small intestine in celiac patients. Given the IL-18 biological role, and since a few studies have previously suggested its involvement in CD, in order to investigate the role of IL18 gene in the susceptibility to CD we have performed a case–control study, analyzing two IL18 gene promoter polymorphisms, previously reported to impair the transcriptional activity of the gene, (−137G > C and −607C > A, rs187238 and rs1946518 respectively). A total of 556 CD Italian patients and 582 controls, further stratified for HLA class II (DQ) CD risk haplotypes were enrolled. The −607A > C A allele and A/A genotype, as well as the combination of this allele with the −137G allele in the AG haplotype, were associated with an increased risk towards CD development, in particular in HLA-DQ2.2 patients. Although the association was very moderate, our results indicate the possible involvement of IL18 gene in the susceptibility to CD, and for this reason we do think it should deserve further investigation.
Similar content being viewed by others
References
Green PHR, Cellier C (2007) Celiac disease. N Engl J Med 357(17):1731–1743
Rostom A, Murray GA, Kagnoff MF (2006) American Gastroenterological Association (AGA) Institute technical review on the diagnosis and management of celiac disease. Gastroenterology 131(6):1981–2002
McCabe MA, Toughill EH, Parkhill AM, Bossett MS, Jevic MS, Nye ML (2012) Celiac disease: a medical puzzle. Am J Nurs 112(10):34–44
Klitz W, Maiers M, Spellman S, Spellman S, Baxter-Lowe LA, Schmeckpeper B, Williams TM, Fernandez-Vina M (2003) New HLA haplotype frequency reference standards: high-resolution and large sample typing of HLA DR–DQ haplotypes in a sample of European Americans. Tissue Antigens 62(4):296–307
Vader W, Stepniak D, Kooy Y, Mearin L, Thompson A, van Rood JJ, Spaenij L, Koning F (2003) The HLA-DQ2 gene dose effect in celiac disease is directly related to the magnitude and breadth of gluten-specific T cell response. PNAS 100(21):12390–12395
Karell K, Louka AS, Moodie SJ, Asher H, Clot F, Greco L, Ciclitira PJ, Sollid LM, Partanen J (2003) HLA types in celiac disease patients not carrying the DQA1*05-DQB1*02 (DQ2) heterodimer: results from the European genetics cluster on celiac disease. Hum Immunol 64(4):469–477
Van Heel DA, Franke L, Hunt KA, Gwilliam R, Zhernakova A, Inouye M, Wapenaar MC, Barnardo MCNM, Bethel G, Holmes GKT, Feighery C, Jewell D, Kelleher D, Kumar P, Travis S, Walters JRF, Sanders DS, Howdle P, Swift J, Playford RJ, McLaren WM, Mearin ML, Mulder CJ, McManus R, McGinnis R, Cardon LR, Deloukas P, Wijmenga C (2007) A genome-wide association study for celiac disease identifies risk variants in the region harboring IL2 and IL21. Nat Genet 39(7):827–829
Hunt KA, Zhernakova A, Turner G, Heap GAR, Franke L, Bruinenberg M, Romanos J, Lotte C, Dinesen LC, Ryan AW, Panesar D, Gwilliam R, Takeuchi F, McLaren WM, Holmes GKT, Peter D, Howdle PD, Walters JRF, Sanders DS, Playford RJ, Trynka G, Mulder CJJ, Mearin ML, Verbeek WHM, Trimble V, Stevens FM, O’Morain C, Kennedy NP, Kelleher D, Pennington DJ, Strachan DP, McArdle WL, Mein CA, Wapenaar MC, Deloukas P, McGinnis R, McManus R, Wijmenga C, van Heel DA (2008) Novel celiac disease genetic determinants related to the immune response. Nat Genet 40(4):395–402
Grandone E, Green PM, Groen HJM, Gwilliam R, Houwen RHJ, Hunt SE, Kaukinen K, Kelleher D, Korponay-Szabo I, Kurppa K, Padraic MacMathuna P, Mäki M, Mazzilli MC, McCann OT, Mearin ML, Mein CA, Mirza MM, Mistry V, Mora B, Morley BI, Mulder CJ, Murray JA, Núñez C, Oosterom E, Ophoff RA, Polanco I, Peltonen L, Platteel M, Rybak A, Salomaa V, Schweizer JJ, Sperandeo MP, Tack GJ, Turner G, Veldink JH, Verbeek WHM, Weersma RK, Wolters VM, Urcelay E, Cukrowska B, Greco L, Susan L, Neuhausen SL, McManus R, Barisani D, Deloukas P, Barrett JC, Saavalainen P, Wijmenga C, van Heel DA (2010) Multiple common variants for celiac disease influencing immune gene expression. Nat Genet 42(4):295–302
Trynka G, Zhernakova A, Romanos J, Trynka G, Zhernakova A, Romanos J, Franke L, Hunt KA, Turner G, Bruinenberg M, Heap GA, Platteel M, Ryan AW, de Kovel C, Holmes GKT, Howdle PD, Walters JRF, Sanders DS, Mulder CJJ, Mearin ML, Verbeek WHM, Trimble V, Stevens FM, Kelleher D, Barisani D, Bardella MT, McManus R, van Heel DA, Wijmenga C (2009) Coeliac disease-associated risk variants in TNFAIP3 and REL implicate altered NF-kappaB signalling. Gut 58(8):1078–1083
Trynka G, Hunt KA, Bockett NA, Romanos J, Mistry V, Szperl A, Bakker SF, Bardella MT, Bhaw-Rosun L, Castillejo G, de la Concha EG, de Almeida RC, Dias KR, van Diemen CC, Dubois PCA, Duerr RH, Edkins S, Franke L, Fransen K, Gutierrez J, Heap GAR, Hrdlickova B, Hunt S, Plaza Izurieta L, Izzo V, Joosten LAB, Langford C, Mazzilli MC, Mein CA, Midah V, Mitja Mitrovic M, Mora B, Morelli M, Nutland S, Núñez C, Onengut-Gumuscu S, Pearce K, Platteel M, Polanco I, Potter S, Ribes-Koninckx C, Ricaño-Ponce I, Rich SS, Rybak A, Santiago JL, Senapati S, Sood A, Szajewska H, Troncone R, Varadé J, Wallace C, Wolters VM, Zhernakova A, CEGEC (Spanish Consortium on the Genetics of Coeliac Disease), Prevent CD Study Group, Wellcome Trust Case Control Consortium, Thelma BK, Cukrowska B, Urcelay E, Bilbao JR, Mearin ML, Barisani D, Barrett JC, Plagnol V, Deloukas P, Wijmenga C, van Heel DA (2012) Dense genotyping identifies and localizes multiple common and rare variant association signals in celiac disease. Nat Genet 43(12):1193–1201
Greco L, Corazza G, Babron MC, Clot F, Fulchignoni-Lataud MC, Percopo S, Zavattari P, Bouguerra F, Dib C, Tosi R, Troncone R, Ventura A, Mantavoni W, Magazzù G, Gatti R, Lazzari R, Giunta A, Perri F, Iacono G, Cardi E, de Virgiliis S, Cataldo F, De Angelis G, Musumeci S, Clerget-Darpoux F (1998) Genome search in celiac disease. Am J Hum Genet 62(3):669–675
Holopainen P, Mustalahti K, Uimari P, Collin P, Maki M, Partanen J (2001) Candidate gene regions and genetic heterogeneity in gluten sensitivity. Gut 48(5):696–701
Naluai AT, Nilsson S, Gudjónsdóttir AH, Louka AS, Ascher H, Ek J, Hallberg B, Samuelsson L, Kristiansson B, Martinsson T, Nerman O, Sollid LM, Wahlström J (2001) Genome-wide linkage analysis of Scandinavian affected sib-pairs supports presence of susceptibility loci for celiac disease on chromosomes 5 and 11. Eur J Hum Genet 9(12):938–944
Nakanishi K, Yoshimoto T, Tsutsui H, Okamura H (2001) Interleukin-18 is a unique cytokine that stimulates both Th1 and Th2 responses depending on its cytokine milieu. Cytokine Growth Factor Rev 12(1):53–72
Ghayur T, Banerjee S, Hugunin M, Butler D, Herzog L, Carter A, Quintal L, Sekut L, Talanian R, Paskind M, Wong W, Kamen R, Tracey D, Alien H (1997) Caspase-1 processes IFN-gamma inducing factor and regulates LPS-induced IFN-gamma production. Nature 386(6625):619–623
Salvati VM, MacDonald TT, Bajaj-Elliott M, Borrelli M, Staiano A, Auricchio S, Troncone R, Monteleone G (2002) Interleukin 18 and associated markers of T helper cell type 1 activity in coeliac disease. Gut 50(2):186–190
Merendino RA, Di Pasquale G, Sturniolo GC, Ruello A, Albanese V, Minciullo PL, Di Mauro S, Gangemi S (2003) Relationship between IL-18 and sICAM-1 serum levels in patients affected by coeliac disease: preliminary considerations. Immunol Lett 85(3):257–260
Eaton AD, Damo X, Garside P (2003) Administration of exogenous interleukin-18 and interleukin-12 prevents the induction of oral tolerance. Immunol 108(2):196–203
Giedraitis V, He B, Huang WX, Hillert J (2001) Cloning and mutation analysis of the human IL-18 promoter: a possible role of polymorphisms in expression regulation. J Neuroimmunol 112(1–2):146-115
Dong GP, Yu ZS, Liang L, Zou CC, Fu JF, Wang CL (2007) IL-18 gene promoter −137C/G and −607C/A polymorphisms in Chinese Han children with type 1 diabetes mellitus. Int J Immunogenet 34(2):75–79
Song GG, Choi SJ, Ji JD, Lee YH (2013) Association between interleukin-18 polymorphisms and systemic lupus erythematosus: a meta-analysis. Mol Biol Rep 40(3):2581–2587
Ben Aleya W, Sfar I, Habibi I, Mouelhi L, Aouadi H, Makhlouf M, Ayed-Jendoubi S, Najjar T, Ben Abdallah T, Ayed K, Gorgi Y (2011) Interleukin-18 gene polymorphisms in tunisian patients with inflammatory bowel disease. Digestion 83(4):269–274
Brophy K, Ryan AW, turner G, Trimble V, Patel KD, O’Morain C, Kennedy NP, Egan B, Close E, Lawlor G, MacMathuna P, MacMathuna P, Stevens FM, Abuzakouk M, Feighery C, Kelleher D, McManus R (2010) Evaluation of 6 candidate genes on chromosome 11q23 for coeliac disease susceptibility: a case control study. BMC Med Genet 11:76
Walker-Smith JA, Guandalini S, Schmitz J, Shmerling DH, Visakorpi JK (1990) Revised criteria for diagnosis of celiac disease. Report of working group of European society of paediatric gastroenterology and nutrition. Arch Dis Child 65(8):909–911
Kaukinen K, Mäki M, Partanen J, Sievänen H, Collin P (2001) Celiac disease without villous atrophy: revision of criteria called for. Dig Dis Sci 46(4):879–888
Ja¨rvinen TT, Collin P, Rasmussen M, Kyrönpalo S, Mäki M, Partanen J, Reunala T, Kaukinen K (2004) Villous tip intraepithelial lymphocytes as markers of early-stage celiac disease. Scand J Gastroenterol 39(5):428–433
Kim SH, Hur GY, Jin HJ, Choi H, Park HS (2012) Effect of interleukin-18 gene polymorphisms on sensitization to wheat flour in bakery workers. J Korean Med Sci 27(4):382–387
Dziedziejko V, Kurzawski M, Paczkowska E, Machalinski B, Pawlik A (2012) The impact of IL18 gene polymorphisms on mRNA levels and interleukin-18 release by peripheral blood mononuclear cells. Postepy Hig Med Dosw (online) 66:409–414
Rueda B, Zhernakova A, López-Nevot MA, Martín J, Koeleman BPC (2005) Association study of functional genetic variants of innate immunity related genes in celiac disease. BMC Med Genet 6:29
Louka AS, Sollid LM (2003) HLA in coeliac disease: unravelling the complex genetics of a complex disorder. Tissue Antigens 61(2):105–117
Boniotto M, Braida L, Baldas V, Not T, Ventura A, Vatta S, Radillo O, Tedesco F, Percopo S, Montico M, Amoroso A, Crovella S (2005) Evidence of a correlation between mannose binding lectin and celiac disease: a model for other autoimmune diseases. J Mol Med 83(4):308–315
León AJ, Garrote JA, Blanco-Quirós A, Calvo C, Fernández-Salazar L, DelVillar A, Barrera A, Arranz E (2006) Interleukin 18 maintains a long-standing inflammation in coeliac disease patients. Clin Exp Immunol 146(3):479–485
Reuter BK, Pizarro Commentary TT (2004) The role of the IL-18 system and other members of the IL-1R/TLR superfamily in innate mucosal immunity and the pathogenesis of inflammatory bowel disease: friend or foe? Eur J Immunol 34(9):2347–2355
Coruh B, Marini M, Weber BD, Moskaluk CA, Pizarro TT (2001) Evidence that IL-18 plays a protective role in an in vivo inflammatory model of epithelial damage and repair. Gastroenterology 120(5):A-122 supplement 1
Takagi H, Kanai T, Okazawa A, Kishi Y, Sato T, Takaishi H, Inoue N, Ogata H, Iwao Y, Hoshino K, Takeda K, Akira S, Watanabe M, Hibi T (2003) Contrasting action of IL-12 and IL-18 in the development of dextran sodium sulphate colitis in mice. Scand J Gastroenterol 38(8):837–844
Acknowledgments
This work has been supported by RC13/12 Grants from IRCCS Burlo Garofolo Trieste (Italy).
Author information
Authors and Affiliations
Corresponding author
Rights and permissions
About this article
Cite this article
Zupin, L., Catamo, E., Polesello, V. et al. Interleukin-18 gene promoter polymorphisms and celiac disease in Italian patients. Mol Biol Rep 42, 525–533 (2015). https://doi.org/10.1007/s11033-014-3796-z
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s11033-014-3796-z