Abstract
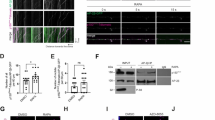
Kinesin-like calmodulin binding protein (KCBP) is a member of kinesin-14 subfamily with unconventional domains distinct from other kinesins. This unique kinesin has the myosin tail homology 4 domain (MyTH4) and band4.1, ezrin, radixin and moesin domain (FERM) at the N-terminal which interact with several cytoskeleton proteins. Although KCBP is implicated in several microtubule-related cellular processes, studies on the KCBP of Dunaliella salina (DsKCBP) have not been reported. In this study, the roles of DsKCBP in flagella and cytoskeleton were investigated and the results showed that DsKCBP was present in flagella and upregulated during flagellar assembly indicting that it may be a flagellar kinesin and plays a role in flagellar assembly. A MyTH4-FERM domain of the DsKCBP was identified as a microtubule and actin interacting site. The interaction of DsKCBP with both microtubules and actin microfilaments suggests that this kinesin may be employed to coordinate these two cytoskeleton elements in algal cells. To gain more insights into the cellular function of the kinesin, DsKCBP-interacting proteins were examined using yeast two-hybrid screen. A 26S proteasome subunit Rpn8 was identified as a novel interacting partner of DsKCBP and the MyTH4-FERM domain was necessary for the interaction of DsKCBP with Rpn8. Furthermore, the DsKCBP was polyubiquitinated and up-regulated by proteasome inhibitor and degraded by ubiquitin–proteasome system indicating that proteasome is related to kinesin degradation.







Similar content being viewed by others
References
Reddy AS, Safadi F, Narasimhulu SB et al (1996) A novel plant calmodulin-binding protein with a kinesin heavy chain motor domain. J Biol Chem 271:7052–7060
Abdel-Ghany SE, Day IS, Simmons MP et al (2005) Origin and evolution of kinesin-like calmodulin-binding protein. Plant Physiol 138:1711–1722
Preuss ML, Delmer DP, Liu B (2003) The cotton kinesin-like calmodulin-binding protein associates with cortical microtubules in cotton fibers. Plant Physiol 132:154–160
Oppenheimer DG, Pollock MA, Vacik J et al (1997) Essential role of a kinesin-like protein in Arabidopsis trichome morphogenesis. Proc Natl Acad Sci USA 94:6261–6266
Dymek EE, Goduti D, Kramer T et al (2006) A kinesin-like calmodulin-binding protein in Chlamydomonas: evidence for a role in cell division and flagellar functions. J Cell Sci 119:3107–3116
Deavours BE, Reddy AS, Walker RA (1998) Ca2+/calmodulin regulation of the Arabidopsis kinesin-like calmodulin-binding protein. Cell Motil Cytoskeleton 40:408–416
Reddy VS, Day IS, Thomas T et al (2004) KIC, a novel Ca2+ binding protein with one EF-hand motif, interacts with a microtubule motor protein and regulates trichome morphogenesis. Plant Cell 16:185–200
Liu L, Li J, Wang C et al (2010) Construction of cDNA library of Dunaliella salina and cloning of kinesin like calmodulin-binding protein gene. J Zhengzhou Univ 45:907–910
Gotesman M, Hosein RE, Gavin R (2011) MyTH4, independent of its companion FERM domain, affects the organization of an intramacronuclear microtubule array and is involved in elongation of the macronucleus in Tetrahymena thermophila. Cytoskeleton 68:220–236
Kao YL, Deavours B, Phelps K et al (2000) Bundling of microtubules by motor and tail domains of a kinesin-like calmodulin-binding protein from Arabidopsis: regulation by Ca2+/calmodulin. Biochem Biophys Res Commun 267:201–207
Cui L, Xue L, Li J et al (2010) Characterization of the glucose-6-phosphate isomerase (GPI) gene from the halotolerant alga Dunaliella salina. Mol Biol Rep 37:911–916
Reddy AS (2007) Analysis of calcium/calmodulin regulation of a plant kinesin using co-sedimentation and ATPase assays. Methods Mol Biol 392:23–36
Xu T, Qu Z, Yang X et al (2009) A cotton kinesin GhKCH2 interacts with both microtubules and microfilaments. Biochem J 421:171–180
Pfannenschmid F, Wimmer VC, Rios RM et al (2003) Chlamydomonas DIP13 and human NA14: a new class of proteins associated with microtubule structures is involved in cell division. J Cell Sci 116:1449–1462
Huang K, Diener DR, Rosenbaum JL (2009) The ubiquitin conjugation system is involved in the disassembly of cilia and flagella. J Cell Biol 186:601–613
Masuda T, Tanaka A, Melis A (2003) Chlorophyll antenna size adjustments by irradiance in Dunaliella salina involve coordinate regulation of chlorophyll a oxygenase (CAO) and Lhcb gene expression. Plant Mol Biol 51:757–771
Zhao C, Omori Y, Brodowska K et al (2012) Kinesin-2 family in vertebrate ciliogenesis. Proc Natl Acad Sci USA 109:2388–2393
Liem KF Jr, He M, Ocbina PJ et al (2009) Mouse Kif7/Costal2 is a cilia-associated protein that regulates Sonic hedgehog signaling. Proc Natl Acad Sci USA 106:13377–13382
Wang D, Zong C, Koag MC et al (2011) Proteome dynamics and proteome function of cardiac 19S proteasomes. Mol Cell Proteomics 10: M110.006122. doi:10.1074/mcp.M110.006122
Seong KM, Baek JH, Yu MH et al (2007) Rpn13p and Rpn14p are involved in the recognition of ubiquitinated Gcn4p by the 26S proteasome. FEBS Lett 581:2567–2573
Sharon M, Taverner T, Ambroggio XI et al (2006) Structural organization of the 19S proteasome lid: insights from MS of intact complexes. PLoS Biol 4:1314–1323
Li Z, Wang CC (2002) Functional characterization of the 11 non-ATPase subunit proteins in the trypanosome 19S proteasomal regulatory complex. J Biol Chem 277:42686–42693
Jenkins PM, McEwen DP, Martens JR (2009) Olfactory cilia: linking sensory cilia function and human disease. Chem Senses 34:451–464
Guo CW, Liu G, Xiong S et al (2011) The C-terminus of MIP-T3 protein is required for ubiquitin-proteasome-mediated degradation in human cells. FEBS Lett 585:1350–1356
Zhang Q, Seo S, Bugge K et al (2012) BBS proteins interact genetically with the IFT pathway to influence SHH-related phenotypes. Hum Mol Genet 21:1945–1953
Roy S (2012) Cilia and Hedgehog: when and how was their marriage solemnized? Differentiation 83:S43–S48
Moen RJ, Johnsrud DO, Thomas DD et al (2011) Characterization of a myosin VII MyTH/FERM domain. J Mol Biol 413:17–23
Acknowledgments
This study was supported by the grants from International Science and Technology Cooperation Program of the Ministry of Science and Technology of P.R. China (No. 2007DFA01240) and the National Natural Science Foundation of China (No. 30700014).
Conflict of interest
There no any conflicts of interest regarding this paper
Author information
Authors and Affiliations
Corresponding author
Rights and permissions
About this article
Cite this article
Shi, K., Li, J., Han, K. et al. The degradation of kinesin-like calmodulin binding protein of D. salina (DsKCBP) is mediated by the ubiquitin–proteasome system. Mol Biol Rep 40, 3113–3121 (2013). https://doi.org/10.1007/s11033-012-2385-2
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s11033-012-2385-2