Abstract
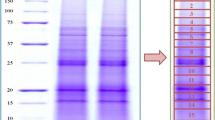
We utilized Percoll density gradient centrifugation to isolate and fractionate chloroplasts of Korean winter wheat cultivar cv. Kumgang (Triticum aestivum L.). The resulting protein fractions were separated by one dimensional polyacrylamide gel electrophoresis (1D-PAGE) coupled with LTQ-FTICR mass spectrometry. This enabled us to detect and identify 767 unique proteins. Our findings represent the most comprehensive exploration of a proteome to date. Based on annotation information from the UniProtKB/Swiss-Prot database and our analyses via WoLF PSORT and PSORT, these proteins are localized in the chloroplast (607 proteins), chloroplast stroma (145), thylakoid membrane (342), lumens (163), and integral membranes (166). In all, 67% were confirmed as chloroplast thylakoid proteins. Although nearly complete protein coverage (89% proteins) has been accomplished for the key chloroplast pathways in wheat, such as for photosynthesis, many other proteins are involved in regulating carbon metabolism. The identified proteins were assigned to 103 functional categories according to a classification system developed by the iProClass database and provided through Protein Information Resources. Those functions include electron transport, energy, cellular organization and biogenesis, transport, stress responses, and other metabolic processes. Whereas most of these proteins are associated with known complexes and metabolic pathways, about 13% of the proteins have unknown functions. The chloroplast proteome contains many proteins that are localized to the thylakoids but as yet have no known function. We propose that some of these familiar proteins participate in the photosynthetic pathway. Thus, our new and comprehensive protein profile may provide clues for better understanding that photosynthetic process in wheat.



Similar content being viewed by others
Abbreviations
- BSA:
-
Bovine serum albumin
- cTP:
-
Chloroplast transit peptide
- FT:
-
Fourier transform
- ICR:
-
Ion cyclotron resonance
- LTQ:
-
Linear quadruple trap
- PIR:
-
Protein information resources
- SP:
-
Signal peptide
- TMD:
-
Transmembrane domain
References
Bryant DA, Frigaard NU (2006) Prokaryotic photosynthesis and phototrophy illuminated. Trends Microbiol 14:488–496
Sheen J (1999) C-4 gene expression. Annu Rev Plant Physiol Plant Mol Biol 50:187–217
Ort DR, Yocum CF (1996) Electron transfer and energy transduction in photosynthesis: an overview. In: Ort DR, Yocum CF (eds) Oxygenic photosynthesis: the light reactions. Advances in photosynthesis and respiration. Springer, Dordrecht, pp 1–9
van Wijk KJ (2004) Plastid proteomics. Plant Physiol Biochem 42:963–977
Jung E, Heller M, Sanchez JC, Hochstrasser DF (2000) Proteomics meets cell biology: the establishment of subcellular proteomes. Electrophoresis 21:3369–3377
van Wijk KJ (2001) Challenges and prospects of plant proteomics. Plant Physiol 126:501–508
Bardel J, Louwagie M, Jaquinod M et al (2000) A survey of the plant mitochondrial proteome in relation to development. Proteomics 2:880–898
Millar AH, Sweetlove LJ, Giege P, Leaver CJ (2001) Analysis of the Arabidopsis mitochondrial proteome. Plant Physiol 127:1711–1727
van Wijk KJ (2000) Proteomics of the chloroplast: experimentation and prediction. Trends Plant Sci 5:420–425
Friso G, Giacomelli L, Ytterberg AJ, Peltier JB et al (2004) In-depth analysis of the thylakoid membrane proteome of Arabidopsis thaliana chloroplasts: new proteins, new functions, and a plastid proteome database. Plant Cell 16:478–499
Peltier JB, Ytterberg AJ, Sun Q, van Wijk KJ (2004) New functions of the thylakoid membrane proteome of Arabidopsis thaliana revealed by a simple, fast, and versatile fractionation strategy. J Biol Chem 279:49367–49383
Santoni V, Kieffer S, Desclaux D, Masson F, Rabilloud T (2000) Membrane proteomics: use of additive main effects with multiplicative interaction model to classify plasma membrane proteins according to their solubility and electrophoretic properties. Electrophoresis 21:3329–3344
Marmagne A, Rouet MA, Ferro M, Rolland N et al (2004) Identification of new intrinsic proteins in Arabidopsis plasma membrane proteome. Mol Cell Proteomics 3:675–691
Mitra SK, Gantt JA, Ruby JF, Clouse SD, Goshe MB (2007) Membrane proteomic analysis of Arabidopsis thaliana using alternative solubilization techniques. J Proteome Res 6:1933–1950
Fukao Y, Hayashi M, Nishimura M (2002) Proteomic analysis of leaf peroxisomal proteins in greening cotyledons of Arabidopsis thaliana. Plant Cell Physiol 43:689–696
Maltman DJ, Simon WJ, Wheeler CH, Dunn MJ et al (2002) Proteomic analysis of the endoplasmic reticulum from developing and germinating seed of castor (Ricinus communis). Electrophoresis 23:626–629
Chivasa S, Ndimba BK, Simon WJ, Robertson D (2002) Proteomic analysis of the Arabidopsis thaliana cell wall. Electrophoresis 23:1754–1765
Degenhardt RF, Bonham-Smith PC (2008) Arabidopsis ribosomal proteins RPL23aA and RPL23aB are differentially targeted to the nucleolus and are disparately required for normal development. Plant Physiol 147:128–142
Ferro M, Salvi D, Brugière S, Miras S, Kowalski S et al (2003) Proteomics of the chloroplast envelope membranes from Arabidopsis thaliana. Mol Cell Proteomics 2:325–345
Peltier JB, Friso G, Kalume DE, Roepstorff P et al (2000) Proteomics of the chloroplast: systematic identification and targeting analysis of lumenal and peripheral thylakoid proteins. Plant Cell 12:319–342
Schubert M, Petersson UA, Haas BJ, Funk C et al (2000) Proteome map of the chloroplast lumen of Arabidopsis thaliana. J Biol Chem 277:8354–8365
Kiselbach T, Hagman A, Anderson B, Schroder WP (1998) The thylakoid lumen of chloroplasts. J Biol Chem 20:6710–6716
D’Amici GM, Huber GC, Zolla L (2009) Separation of thylakoid membrane proteins by sucrose gradient ultracentrifuge or blue native-SDS-PAGE two-dimensional electrophoresis. In: Peirce MJ, Waits R (eds) Membrane proteomics: methods and protocols. Springer, New York, pp 61–70
Hippler M, Klein J, Fink A, Allinger T, Hoerth P (2001) Towards functional proteomics of membrane protein complexes: analysis of thylakoid membranes from Chlamydomonas reinharditii. Plant J 28:595–606
Kikuchi S, Hirohashi T, Nakai M (2006) Characterization of the preprotein translocon at the outer envelope membrane of chloroplasts by Blue Native PAGE. Plant Cell Physiol 47:363–371
Sugiyama N, Nakagami H, Mochida K, Daudi A (2008) Large-scale phosphorylation mapping reveals the extent of tyrosine phosphorylation in Arabidopsis. Mol Syst Biol 4:193
Krijgsveld J, Gauci S, Dormeyer W, Heck AJ (2006) In-gel isoelectric focusing of peptides as a tool for improved protein identification. J Proteome Res 5:1721–1730
Cao X, Nesvizhskii AI (2008) Improved sequence tag generation method for peptide identification in tandem mass spectrometry. J Proteome Res 7:4422–4434
Porra RJ, Thompson WA, Kriedemann PE (1989) Determination of accurate extinction coefficients and simultaneous equations for assaying chlorophylls a and b extracted with four different solvents; verification of the concentration of chlorophyll standards by atomic absorption spectroscopy. Biochim et Biophys Acta 975:384–394
Zorb C, Herbst R, Forreiter C, Schubert S (2009) Short-term effects of salt exposure on the maize chloroplast protein pattern. Proteomics 9:4209–4220
Schagger H, von Jagow G (1987) Tricine-sodium dodecyl sulfate-polyacrylamide gel electrophoresis for the separation of proteins in the range from 1 to 100 kDa. Anal Biochem 166:368–379
Kim JY, Lee JH, Park GW, Cho K, Kwon KH et al (2005) Utility of electrophoretically derived protein mass estimates as additional constraints in proteome analysis of human serum based on MS/MS analysis. Proteomics 5:3376–3385
Olsen JV, Mann M (2004) Improved peptide identification in proteomics by two consecutive stages of mass spectrometric fragmentation. Proc Natl Acad Sci USA 101:13417–13422
Horton P, Park KJ, Obayashi T, Fujita N et al (2007) WoLF PSORT: protein localization predictor. Nucleic Acids Res 35:585–587
Nakai K, Horton P (1999) PSORT: a program for detecting sorting signals in proteins and predicting their subcellular localization. Trends Biochem Sci 24:34–36
Emanuelsson O, Nielsen H, von Heijne G (1999) ChloroP, a neural network-based method for predicting chloroplast transit peptides and their cleavage sites. Protein Sci 8:978–984
Nielsen H, Engelbrecht J, Brunak S, von Heijne G (1997) Identification of prokaryotic and eukaryotic signal peptides and prediction of their cleavage sites. Protein Eng 10:1–6
Möller S, Croning MD, Apweiler R (2001) Evaluation of methods for the prediction of membrane spanning regions. Bioinformatics 17:646–653
Ishihama Y, Oda Y, Tabata T, Sato T et al (2005) Exponentially modified protein abundance index (emPAI) for estimation of absolute protein amount in proteomics by the number of sequenced peptides per protein. Mol Cell Proteomics 4:1265–1272
Matsuoka M (1990) Classification and characterization of cDNA that encodes the light-harvesting chlorophyll a/b binding protein of photosystem II from rice. Plant Cell Physiol 31:519–526
Lemieux C, Otis C, Turmel M (2000) Ancestral chloroplast genome in Mesostigma viride reveals an early branch of green plant evolution. Nature 403:649–652
Yuan HM, Li KL, Ni RJ, Guo WD et al (2010) A systemic proteomic analysis of Populus chloroplast by using shotgun method. Mol Biol Rep. doi:10.1007/s11033-010-9971-y
Ferro M, Salvi D, Rivière-Rolland H, Vermat T et al (2002) Integral membrane proteins of the chloroplast envelope: identification and subcellular localization of new transporters. Proc Natl Acad Sci USA 99:11487–11492
Syka JE, Marto JA, Bai DL, Horning S et al (2004) Novel linear quadrupole ion trap/FT mass spectrometer: performance characterization and use in the comparative analysis of histone H3 post-translational modifications. J Proteome Res 3:326–621
Peltier JB, Emanuelsson O, Kalume DE, Ytterberg J et al (2002) Central functions of the lumenal and peripheral thylakoid proteome of Arabidopsis determined by experimentation and genome-wide prediction. Plant Cell 14:211–236
Gómez SM, Bil KY, Aguilera R, Nishio JN et al (2003) Transit peptide cleavage sites of integral thylakoid membrane proteins. Mol Cell Proteomics 2:1068–1085
Schwartz R, Ting CS, King J (2001) Whole proteome pi values correlate with subcellular localizations of proteins for organisms within the three domains of life. Genome Res 11:703–709
Dalbey RE, Robinson C (1999) Protein translocation into and across the bacterial plasma membrane and the plant thylakoid membrane. Trends Biochem Sci 24:17–22
Keegstra K, Cline K (1999) Protein import and routing systems of chloroplasts. Plant Cell 11:557–570
Cristóbal S, de Gier JW, Nielsen H, von Heijne G (1999) Competition between Sec- and TAT-dependent protein translocation in Escherichia coli. EMBO J 18:2982–2990
Claros MG, von Heijne G (1994) TopPred II: an improved software for membrane protein structure predictions. Comput Appl Biosci 10:685–686
Krogh A, Larsson B, von Heijne G, Sonnhammer EL (2001) Predicting transmembrane protein topology with a hidden Markov model: application to complete genomes. J Mol Biol 305:567–580
Tanford C (1978) Hydrophobic effect and the organization of living matter. Science 200:1012–1018
Kruger NJ, von Schaewen A (2003) The oxidative pentose phosphate pathway: structure and organization. Curr Opin Plant Biol 6:236–246
Wall MK, Mitchenall LA, Maxwell A (2004) Arabidopsis thaliana DNA gyrase is targeted to chloroplasts and mitochondria. Proc Natl Acad Sci USA 101:7821–7826
Dupont FM (2008) Metabolic pathways of the wheat (Triticum aestivum) endosperm amyloplast revealed by proteomics. BMC Plant Biol 8:39
Huang H, Barker WC, Chen Y, Wu CH (2003) iProClass: an integrated database of protein family, function and structure information. Nucleic Acids Res 31:390–392
Martin W, Herrmann RG (1998) Gene transfer from organelles to the nucleus: how much, what happens, and why? Plant Physiol 118:9–17
Ogihara Y, Isono K, Kojima T, Endo A et al (2000) Chinese spring wheat (Triticum aestivum L.) chloroplast genome: complete sequence and contig clones. Plant Mol Biol Rep 18:243–253
Tang J, Xia H, Cao M, Zhang X et al (2004) A comparison of rice chloroplast genomes. Plant Physiol 135:412–420
Mallick P, Schirle M, Chen SS, Flory MR et al (2007) Computational prediction of proteotypic peptides for quantitative proteomics. Nat Biotechnol 25:125–131
MacKintosh C (2004) Dynamic interactions between 14-3-3 proteins and phosphoproteins regulate diverse cellular processes. Biochem J 381:329–342
Li J (2005) Brassinosteroid signaling: from receptor kinases to transcription factors. Curr Opin Plant Biol 8:526–531
Wu K, Rooney MF, Ferl RJ (1997) The Arabidopsis 14-3-3 multigene family. Plant Physiol 114:1421–1431
Acknowledgments
We would like to thank Prof. Shien Young Kang of the Department of Veterinary Medicine, College of Veterinary Medicine, Chungbuk National University, Korea, for conducting the ultra-centrifugation. Mainly, financial support for this study was obtained from the AGENDA (20090101036022), RDA, Korea to S. H. Woo, and also technically and financially supported by the Korea Basic Science Institute Grant (G30121) to Kun Cho and Korea Basic Science Institute K-MeP Project (T30110) to J.-S. Choi.
Author information
Authors and Affiliations
Corresponding author
Electronic supplementary material
Below is the link to the electronic supplementary material.
11033_2011_1302_MOESM1_ESM.xls
Supplementary Table 1 Detailed proteomic data for proteins detected in the chloroplast fraction. The table provides the protein accession number (UniProt_Sprot database), protein descriptions, taxonomy, gene name, subcellular location predicted by WoLF PSORT and PSORT, protein score, molecular weight (MW), protein length, protein matches, peptide queries, protein coverage, iso-electric point (pI), exponential modified protein abundance index (emPAI), chloroplast transit peptide (cTP), signal peptide (SP), transmembrane domains (TMD), and gene ontology-based functional classifications according to biological process using the iProClass database. (XLS 268 kb)
Rights and permissions
About this article
Cite this article
Kamal, A.H.M., Cho, K., Komatsu, S. et al. Towards an understanding of wheat chloroplasts: a methodical investigation of thylakoid proteome. Mol Biol Rep 39, 5069–5083 (2012). https://doi.org/10.1007/s11033-011-1302-4
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s11033-011-1302-4