Abstract
One of the major limitations when attempting to obtain detailed biochemical, biophysical and immunological characterization of plant DNA mismatch repair proteins is their extremely low abundance in vivo under normal growth conditions. An initial analysis of PMS1 transcript level in various Arabidopsis thaliana tissues was carried out by quantitative real-time RT-PCR. For calli, flowers and seedlings, the corresponding cDNA copies per ng RNA were 66.9, 3.1 and 2.7, respectively. This suggests an important role of this gene in rapidly dividing tissues. In order to obtain a high level of PMS1 from Arabidopsis thaliana, the protein production was successfully optimized in an Escherichia coli host. The corresponding coding sequence of PMS1 was inserted into pET28a downstream a hexa-histidyl leader sequence. The pET28a–AtPMS1 plasmid was efficiently expressed in JM109(DE3)-pRIL strain probably due to the genotype features of the cells (endA1, recA1, relA1, Δ(lac-proAB), laqIqZΔM15) and the presence of extra copies of argU, ileY, and leuW tRNA genes, which encode the RIL codons. This strategy has allowed us to obtain His-tagged PMS1 at about 7% of the total soluble E. coli cell protein. The protein was purified by standard Ni+ affinity chromatography procedures and the electrophoretically homogeneous preparation was used as an antigen for antibody generation in rabbits. This approach provides effective tools for a further reconstitution of plant mismatch repair (MMR) system in vitro and for the analysis of protein expression and distribution of AtPMS1 in various tissues after different treatments (e.g. DNA mutagens).




Similar content being viewed by others
References
Li G-M (2008) Mechanisms and functions of DNA mismatch repair. Cell Res 18:85–98
Kunkel T, Erie D (2005) DNA mismatch repair. Annu Rev Biochem 74:681–710
Iyer R, Pluciennik A, Burdett V, Modrich P (2006) DNA mismatch repair: functions and mechanisms. Chem Rev 106:302–323
Jiricny J (2006) The multifaceted mismatch-repair system. Nat Rev Mol Cell Biol 7:335–346
Hsieh P, Yamane K (2008) DNA mismatch repair: molecular mechanism, cancer, and ageing. Mech Ageing Dev 129:391–407
Dzantiev L, Constantin N, Genschel J, Iyer R, Burgers P, Modrich P (2004) A defined human system that supports bidirectional mismatch-provoked excision. Mol Cell 15:31–41
Li G, Modrich P (1995) Restoration of mismatch repair to nuclear extracts of H6 colorectal tumor cells by a heterodimer of human MutL homologs. Proc Natl Acad Sci USA 92:1950–1954
Prolla T, Christie D, Liskay R (1994) Dual requirement in yeast DNA mismatch repair for MLH1 and PMS1, two homologs of the bacterial mutL gene. Mol Cell Biol 14:407–415
Wang T-F, Kleckner N, Hunter N (1999) Functional specificity of MutL homologs in yeast: Evidence for three Mlh1-based heterocomplexes with distinct roles during meiosis in recombination and mismatch correction. Proc Natl Acad Sci USA 96:13914–13919
Abdel-Rahman WM, Mecklin J-P, Peltomäki P (2006) The genetics of HNPCC: application to diagnosis and screening. Crit Rev Oncol Hematol 58:208–220
Bergerat A, de Massy B, Gadelle D, Varoutas PC, Nicolas A, Forterre P (1997) An atypical type II DNA topoisomerase from archaea with implication for meiotic recombination. Nature 386:414–417
Mushegian AR, Bassett DE Jr, Boguski MS, Bork P, Koonin EV (1997) Positionally cloned human disease genes: patterns of evolutionary conservation and functional motifs. Proc Natl Acad Sci USA 94:5831–5836
Dutta R, Inouye M (2000) GHKL, an emergent ATPase/kinase superfamily. Trends Biochem Sci 25:24–28
Hall M, Shcherbakova P, Fortune J et al (2003) DNA binding by yeast Mlh1 and Pms1: implications for DNA mismatch repair. Nucleic Acids Res 31:2025–2034
Ban C, Yang W (1998) Crystal structure and ATPase activity of MutL: implications for DNA repair and mutagenesis. Cell 95:541–552
Guarne A, Junop MS, Yang W (2001) Structure and function of the N-terminal 40 kDa fragment of human PMS2: a monomeric GHL ATPase. EMBO J 20:5521–5531
Hall M, Shcherbakova P, Kunkel T (2002) Differential ATP binding and intrinsic ATP hydrolysis by amino-terminal domains of the yeast Mlh1 and Pms1 proteins. J Biol Chem 277:3673–3679
Spampinato C, Modrich P (2000) The MutL ATPase is required for mismatch repair. J Biol Chem 275:9863–9869
Bende SM, Grafstrom RH (1991) The DNA binding properties of the MutL protein isolated from Escherichia coli. Nucleic Acids Res 19:1549–1555
Ban C, Junop M, Yang W (1999) Transformation of MutL by ATP binding and hydrolysis: a switch in DNA mismatch repair. Cell 97:85–97
Shcherbakova P, Hall M, Lewis M et al (2001) Inactivation of DNA mismatch repair by increased expression of yeast MLH1. Mol Cell Biol 21:940–951
Fukui J, Nishida M, Nakagawa T, Masui R, Kuramitsu S (2008) Bound nucleotide controls the endonuclease activity of mismatch repair enzyme MutL. J Biol Chem 283:12136–12145
Kadyrov FA, Dzantiev L, Constantin N, Modrich P (2006) Endonucleolytic function of MutLα in human mismatch repair. Cell 126:297–308
Kadyrov FA, Holmes SF, Arana ME et al (2007) Saccharomyces cerevisiae MutLα is a mismatch repair endonuclease. J Biol Chem 282:37181–37190
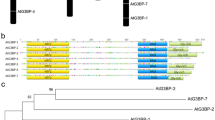
Alou AH, Jean M, Domingue O, Belzile FJ (2004) Structure and expression of AtPMS1, the Arabidopsis ortholog of the yeast DNA repair gene PMS1. Plant Sci 167:447–456
Jean M, Pelletier J, Hilpert M, Belzile F, Kunze R (1999) Isolation and characterization of AtMLH1, a MutL homologue from Arabidopsis thaliana. Mol Gen Genet 262:633–642
Jackson N, Sanchez-Moran E, Buckling E, Armstrong SJ, Jones GH, Franklin FCH (2006) Reduced meiotic crossovers and delayed prophase I progression in AtMLH3-deficient Arabidopsis. EMBO J 25:1315–1323
Murashige T, Skoog F (1962) A revised medium for rapid growth and bioassays with tobacco tissue cultures. Physiol Plant 15:473–497
Sambrook J, Russel DW (2001) Molecular cloning—a laboratory manual. Cold Spring Harbor Laboratory Press, Cold Spring Harbor
Plaxton WC (1989) Molecular and immunological characterization of plastid and cytosolic pyruvate kinase isozymes from castor-oil-plant endosperm and leaf. Eur J Biochem 181:443–451
Laemmli UK (1970) Cleavage of structural proteins during the assembly of the head of bacteriophage T4. Nature 227:680–685
Burnette WN (1981) Western Blotting: electrophoretic transfer of proteins from sodium dodecyl sulfate-polyacrylamide gels to unmodified nitrocellulose and radiographic detection with antibody and radioiodinated protein A. Anal Biochem 112:195–203
Bradford MM (1976) A rapid and sensitive method for the quantitation of microgram quantities of protein utilizing the principle of protein dye binding. Anal Biochem 72:248–254
Spampinato CP, Gomez RL, Galles C, Lario LD (2009) From bacteria to plants: a compendium of mismatch repair assays. Mutat Res 682:110–128
Dion E, Li L, Jean M, Belzile F (2007) An Arabidopsis MLH1 mutant exhibits reproductive defects and reveals a dual role for this gene in mitotic recombination. Plant J 51:431–440
Li L, Dion E, Richard G, Domingue O, Jean M, Belzile FJ (2009) The Arabidopsis DNA mismatch repair gene PMS1 restricts somatic recombination between homeologous sequences. Plant Mol Biol 69:675–684
Alou A, Azaiez A, Jean M, Belzile F (2004) Involvement of the Arabidopsis thaliana AtPMS1 gene in somatic repeat instability. Plant Mol Biol 56:339–349
Li L, Jean M, Belzile F (2006) The impact of sequence divergence and DNA mismatch repair on homeologous recombination in Arabidopsis. Plant J 45:908–916
Gustafson C, Govindarajan S, Minshull J (2004) Codon bias and heterologous protein expression. Trends Biotechnol 22:346–353
Räschle M, Marra G, Nyström-Lahti M, Schär P, Jiricny J (1999) Identification of hMutL, a heterodimer of hMLH1 and hPMS1. J Biol Chem 274:32368–32375
Cutalo JM, Darden TA, Kunkel TA, Tomer KB (2006) Mapping the dimer interface in the C-terminal domains of the yeast MLH1–PMS1 heterodimer. Biochemistry 45:15458–15467
Rosano GL, Ceccarelli EA (2009) Rare codon content affects the solubility of recombinant proteins in a codon bias-adjusted Escherichia coli strain. Microb Cell Fact 8:41
Acknowledgements
We acknowledge research support from Fundación Antorchas and CONICET. RLG and CG are fellows of the Consejo Nacional de Investigaciones Científicas y Técnicas (CONICET). CPS is a member of the Researcher Career of the same institution.
Author information
Authors and Affiliations
Corresponding author
Rights and permissions
About this article
Cite this article
Galles, C., Gomez, R.L. & Spampinato, C.P. PMS1 from Arabidopsis thaliana: optimization of protein overexpression in Escherichia coli . Mol Biol Rep 38, 1063–1070 (2011). https://doi.org/10.1007/s11033-010-0203-2
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s11033-010-0203-2