Abstract
Channa marulius (Hamilton, 1822) is a commercially important freshwater fish and a potential candidate species for aquaculture. The present study evaluated partial Cytochrome b gene sequence of mtDNA for determining the genetic variation in wild populations of C. marulius. Genomic DNA extracted from C. marulius samples (n = 23) belonging to 3 distant rivers; Mahanadi, Teesta and Yamuna was analyzed. Sequencing of 307 bp Cytochrome b mtDNA fragment revealed the presence of 5 haplotypes with haplotype diversity value of 0.763 and nucleotide diversity value of 0.0128. Single population specific haplotype was observed in Mahanadi and Yamuna samples and 3 haplotypes in Teesta samples. The analysis of data demonstrated the suitability of partial Cytochrome b sequence in determining the genetic diversity in C. marulius population.

Similar content being viewed by others
References
Munro ISR (1955) The marine and freshwater fishes of Ceylon. Dept. Ext. Affairs, Canberra, 348 pp, 56 pls
Bardach JE, Ryther JH, McLarney W (1972) Aquaculture: the farming and husbandry of freshwater and marine organisms. Wiley, New York, 867p
Talwar PK, Jhingran AG (1992) Inland fishes of India. Rec Ind J 3:19–24
CAMP (1998) Report of the workshop conservation assessment and management plan (CAMP) for freshwater fishes of India. Organized by Zoo Outreach Activity and National Bureau of Fish Genetic Resources, Lucknow
Lecomte F, Grant WS, Dodson JJ, Rodriguez-Sanchez R, Bowen BW (2004) Living with uncertainty: genetic imprints of climate shifts in east Pacific anchovy (Engraulis mordax) and sardine (Sardinops sagax). Mol Ecol 13:2169–2182
Fontana F, Conterio F, Gandolfi G, Tagliavini J, Rosenthal H, Bronzi P, McKenzie DJ (2007) Mitochondrial DNA sequences of 6 sturgeon species and phylogenetic relationships within Acipenseridae. J Appl Ichthyol 15(4–5):17–22
Pages M, Desse-Berset N, Tongard C, Brosse L, Hänni C, Berrebi P (2009) Historical presence of the sturgeon Acipenser sturio in the Rhone Basin determined by the analysis of ancient DNA cytochrome b sequences. Conserv Genet 10:217–224
Murray WB, Wang JY, Yang SC, Stevens JD, Fisk A, Svavarsson J (2008) Mitochondrial cytochrome b variation in sleeper sharks (Squaliformes: Somniosidae). Mar Biol 153:1015–1022
Oleinik AG, Skurikhina LA, Brykov VA (2007) Divergence of Salvelinus sp from north eastern Asia based on mitochondrial DNA. Ecol Freshw 16:87–98
Bouza C, Vilas R, Castro J (2008) Mitochondrial haplotype variability of brown trout populations from north-western Iberian Peninsula, a secondary contact area between lineages. Conserv Genet 9:917–920
Daemen E, Cross T, Ollevier F, Volckaert FAM (2001) Analysis of genetic structure of European eel (Anguilla anguilla) using microsatellite DNA and mtDNA markers. Mar Biol 139:755–764
Fayazi J, Moradi M, Rahimi G, Ashtyani R, Galledari H (2006) Genetic differentiation and phylogenetic relationships among Barbus xanthopterus (Cyprinidae) populations in south west of Iran using mitochondrial DNA markers. Pak J Biol Sci 9(12):2249–2254
So N, VanHoudt JKJ, Volckaert FAM (2006) Genetic diversity and population history of the migratory catfishes Pangasianodon hypophthalmus and Pangasius bocourti in the Cambodian Mekong River. Fish Sci 72:469–476
Hsu KC, Shih NT, Ni HI, Tsao Shao K (2007) Genetic variation in Trichiurus lepturus (Perciformes: Trichiuridae) in water of Taiwan: several species or cohort distribution? Raffles Bull Zool 14:215–220
Brown J, Stapien CA (2008) Ancient divisions, recent expansions: phylogeography and population genetics of the round Gobi Appolonia melanostoma. Mol Ecol 17:2598–2615
Abol-Munafi AB, Ambok MA, Ismail P, MinhTam B (2007) Molecular data from the cytochrome b for the phylogeny of Channidae (Channa sp) in Malaysia. Biotechnology 6(1):22–27
Li X, Musikasinthorn P, Kumazawa Y (2006) Molecular phylogenetic analysis of snakeheads (Perciformes: Channidae) using mitochondrial DNA sequences. Ichthyol Res 53:148–159
Haniffa MA, Nagarajan M, Gopalakrishnan A, Basheer VS, Muneer A (2006) Genetic variability of Channa punctatus populations using randomly amplified polymorphic DNA. Aquac Res 37:1151–1155
Hanniffa MA, Nagarajan M, Gopalakrishnan A, Musammilu KK (2007) Allozyme variation in threatened freshwater fish spotted Murrel (Channa punctatus) in south Indian river system. Biochem Genet 45(3/4):363–373
Ponniah AG, Sarkar UK (2000) Fish biodiversity of north east India. NBFGR-NATP Publ 2:24–25
Abdul Muneer PM, Gopalakrishnan A, Musammilu KK, Mohindra V, Lal KK, Basheer VS, Lakra WS (2009) Genetic variation and population structure of endemic yellow catfish, Horabagrus brachysoma (Bagridae) among three populations of Western Ghat region using RAPD and microsatellite markers. Mol Biol Rep 36:1779–1791
Ruzzante DE, Taggart CT, Cook D (1996) Spatial and temporal variation in the genetic composition of a larval cod (Gadus morhua) aggregation: cohort contribution and genetic stability. Can J Fish Aquat Sci 53:2695–2705
Kocher TD, Thomas WK, Meyer A, Edwards SV, Pabo S, Villablanca FX, Wilson AC (1989) Dynamics of mitochondrial DNA evolution in animals: amplification and sequencing with conserved primers. Proc Natl Acad Sci USA 86:6196–6200
Thompson JD, Higgins DG, Gibson TJ (1994) CLUSTAL W: improving the sensitivity of progressive multiple sequence alignment through sequence weighting, position-specific gap penalties and weight matrix choice. Nucleic acids research advance access. Nucleic Acids Res 22:4673–4680
Tamura K, Dudley J, Nei M, Kumar S (2007) MEGA 4: molecular evolutionary genetics analysis (MEGA) software version 4.0. Mol Biol Evol 24:1596–1599
Excoffier LG, Schneider LS (2005) Arlequin ver. 3.0: an integrated software package for population genetics data analysis. Evol Bioinformatics Online 1:47–50
Rozas J, Sánchez-Delbarrio JC, Messeguer X, Rozas R (2003) DnaSP, DNA polymorphism analyses by the coalescent and other methods. Bioinformatics 19:2496–2497
Kocher TD, Conroy JA, McKaye KR, Stauffer JR (1993) Similar morphologies of cichlid fish in Lakes Tanganyika and evolution, edited by M. H. A. Keenleyside. Chapman and Hall, Malawi are due to convergence. Mol Phylogenet Evol 2:158–165
Song CB, Near TJ, Page LM (1998) Phylogenetic relationships among Percid fishes as inferred from mitochondrial cytochrome b DNA sequence data. Mol Phylogenet Evol 10:343–353
Abol-Munafi AB, Ambok MA, Ismail P, MinhTam B (2007) Molecular data from the cytochrome b for the phylogeny of Channidae (Channa sp) in Malaysia. Biotechnology 6(1):22–27
Apostolidis AP, Triantaphyllidis C, Kouvatsi A, Economidis PS (1997) Mitochondrial DNA sequence variation and phylogeography among Salmo trutta L (Greek brown trout) populations. Mol Ecol 6(6):531–542
Finne KL (2001) Phylogeographic structure of the Atlantic pupfish, Cyprinodon variegatus (Cyprinodontidae), along the eastern coast of North America. Unpublished M.S. Thesis, Virginia Polytechnic Institute and State University, Blacksburg, Virginia
Tinti F, di Nunno C, Guarniero I, Talenti M, Tommasini S, Fabbri E, Piccinetti C (2002) Mitochondrial DNA sequence variation suggests the lack of genetic heterogeneity in the adriatic and ionian stocks of Sardina pilchardus. Mar Biotechnol 4:163–172
Marshall CRE (2005) Evolutionary genetics of barramundi (Lates calcarifer) in the Australian region. Unpublished Ph.D Thesis, School of Biological Sciences and Biotechnology, Murdoch University, Perth, Western Australia
Vrijenhoek RC (1998) Conservation genetics of freshwater fish. J Fish Biol 53(Suppl A):394–412
Chondar SL (1999) Biology of fin fishes and shellfishes. SCSC Publishers, Howrah, India
Avise JC (1994) Molecular markers, natural history, and evolution. Chapman and Hall, New York
Lakra WS, Goswami M, Gopalakrishnan A (2009) Molecular identification and phylogenetic relationships of seven Indian Sciaenids (Pisces: Perciformes, Sciaenidae) based on 16S rRNA and cytochrome c oxidase subunit I mitochondrial genes. Mol Biol Rep 36:831–839
Bowen BW, Meylan AB, Perran R (1992) Global population structure and natural history of Green Turtle (Chelonia mydas) in terms of matriarchal phylogeny. Evolution 46:865–881
Acknowledgments
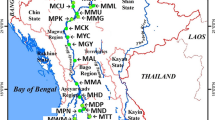
The authors thank Sh. A.K. Pathak (Scientist and OIC ARIS Cell, NBFGR) for providing the locality map. Excellent technical cooperation from Sh. Akhilesh Mishra, Sh. Rajesh Kumar and Sh. R.S Sah is duly acknowledged.
Author information
Authors and Affiliations
Corresponding author
Rights and permissions
About this article
Cite this article
Habib, M., Lakra, W.S., Mohindra, V. et al. Evaluation of cytochrome b mtDNA sequences in genetic diversity studies of Channa marulius (Channidae: Perciformes). Mol Biol Rep 38, 841–846 (2011). https://doi.org/10.1007/s11033-010-0175-2
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s11033-010-0175-2