Abstract
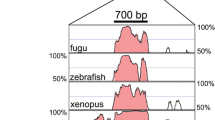
The intermediate filament protein vimentin is involved in a variety of cellular functions both during the normal developmental processes and in human malignancies. We here describe the identification of an alternative vimentin transcript initiating upstream for the canonical vimentin gene promoter and spliced using the vimentin promoter sequence as an intron. Expression analysis showed that the alternative vimentin promoter had the same expression profile as the canonical vimentin gene promoter. The presented data suggest that alternative promoter usage and alternative splicing could be regulatory mechanisms participating in vimentin gene regulation.


Similar content being viewed by others
References
Parry DA, Steinert PM (1999) Intermediate filaments: molecular architecture, assembly, dynamics and polymorphism. Q Rev Biophys 32(2):99–187
Herrmann H, Aebi U (2004) Intermediate filaments: molecular structure, assembly mechanism, and integration into functionally distinct intracellular Scaffolds. Annu Rev Biochem 73:749–789
Ivaska J et al (2007) Novel functions of vimentin in cell adhesion, migration, and signaling. Exp Cell Res 313(10):2050–2062
Wang N, Stamenovic D (2002) Mechanics of vimentin intermediate filaments. J Muscle Res Cell Motil 23(5–6):535–540
Duprey P, Paulin D (1995) What can be learned from intermediate filament gene regulation in the mouse embryo. Int J Dev Biol 39(3):443–457
Kokkinos MI et al (2007) Vimentin and epithelial-mesenchymal transition in human breast cancer–observations in vitro and in vivo. Cells Tissues Organs 185(1–3):191–203
Rittling SR, Baserga R (1987) Functional analysis and growth factor regulation of the human vimentin promoter. Mol Cell Biol 7(11):3908–3915
Benazzouz A, Duprey P (1999) The vimentin promoter as a tool to analyze the early events of retinoic acid-induced differentiation of cultured embryonal carcinoma cells. Differentiation 65(3):171–180
Evans RM (1998) Vimentin: the conundrum of the intermediate filament gene family. Bioessays 20(1):79–86
Zhang X et al (2009) Inhibition of vimentin or beta1 integrin reverts morphology of prostate tumor cells grown in laminin-rich extracellular matrix gels and reduces tumor growth in vivo. Mol Cancer Ther 8(3):499–508
Hendrix MJ et al (1997) Experimental co-expression of vimentin and keratin intermediate filaments in human breast cancer cells results in phenotypic interconversion and increased invasive behavior. Am J Pathol 150(2):483–495
Gilles C (1999) Vimentin contributes to human mammary epithelial cell migration. J Cell Sci 112(Pt 24):4615–4625
Perreau J et al (1988) Nucleotide sequence of the human vimentin gene and regulation of its transcription in tissues and cultured cells. Gene 62(1):7–16
Pieper FR et al (1992) Regulation of vimentin expression in cultured epithelial cells. Eur J Biochem 210(2):509–519
Izmailova ES et al (1999) A GC-box is required for expression of the human vimentin gene. Gene 235(1–2):69–75
Sax CM et al (1988) Multiple elements are required for expression of an intermediate filament gene. Nucleic Acids Res 16(16):8057–8076
Kryszke MH, Vicart P (1998) Regulation of the expression of the human vimentin gene: application to cellular immortalization. Pathol Biol (Paris) 46(1):39–45
Moura-Neto V et al (1996) A 28-bp negative element with multiple factor-binding activity controls expression of the vimentin-encoding gene. Gene 168(2):261–266
Wu Y et al (2007) The zinc finger repressor, ZBP-89, recruits histone deacetylase 1 to repress vimentin gene expression. Genes Cells 12(8):905–918
Izmailova ES, Zehner ZE (1999) An antisilencer element is involved in the transcriptional regulation of the human vimentin gene. Gene 230(1):111–120
Wu Y et al (2004) Stat3 enhances vimentin gene expression by binding to the antisilencer element and interacting with the repressor protein, ZBP-89. Oncogene 23(1):168–178
Condorelli DF et al (1999) GFAPbeta mRNA expression in the normal rat brain and after neuronal injury. Neurochem Res 24(5):709–714
Nielsen AL et al (2002) A new splice variant of glial fibrillary acidic protein, GFAP epsilon, interacts with the presenilin proteins. J Biol Chem 277(33):29983–29991
Blechingberg J et al (2007) Regulatory mechanisms for 3′-end alternative splicing and polyadenylation of the glial fibrillary acidic protein, GFAP, transcript. Nucleic Acids Res 35(22):7636–7650
Condorelli DF et al (1999) Structural features of the rat GFAP gene and identification of a novel alternative transcript. J Neurosci Res 56(3):219–228
Hol EM et al (2003) Neuronal expression of GFAP in patients with Alzheimer pathology and identification of novel GFAP splice forms. Mol Psychiatry 8(9):786–796
Aparicio O et al. (2005) Chromatin immunoprecipitation for determining the association of proteins with specific genomic sequences in vivo. Curr Protoc Mol Biol. Chapter 21: p. Unit 21 3
Kragh PM et al. (2009) Hemizygous minipigs produced by random gene insertion and handmade cloning express the Alzheimer’s disease-causing dominant mutation APPsw. Transgenic Res 18:545–558
Yamamichi-Nishina M et al (2003) SW13 cells can transition between two distinct subtypes by switching expression of BRG1 and Brm genes at the post-transcriptional level. J Biol Chem 278(9):7422–7430
Hedberg KK, Chen LB (1986) Absence of intermediate filaments in a human adrenal cortex carcinoma-derived cell line. Exp Cell Res 163(2):509–517
Butler R, Robertson J, Gallo JM (2000) Mutually exclusive expression of beta(III)-tubulin and vimentin in adrenal cortex carcinoma SW13 cells. FEBS Lett 470(2):198–202
Sarria AJ (1994) The presence or absence of a vimentin-type intermediate filament network affects the shape of the nucleus in human SW-13 cells. J Cell Sci 107(Pt 6):1593–1607
Sommers CL et al (1994) Regulation of vimentin gene transcription in human breast cancer cell lines. Cell Growth Differ 5(8):839–846
Stover DM et al (1994) A negative regulatory factor is missing in a human metastatic breast cancer cell line. Cancer Res 54(12):3092–3095
Carninci P et al (2006) Genome-wide analysis of mammalian promoter architecture and evolution. Nat Genet 38(6):626–635
Heintzman ND et al (2007) Distinct and predictive chromatin signatures of transcriptional promoters and enhancers in the human genome. Nat Genet 39(3):311–318
Mikkelsen TS et al (2007) Genome-wide maps of chromatin state in pluripotent and lineage-committed cells. Nature 448(7153):553–560
Guenther MG et al (2007) A chromatin landmark and transcription initiation at most promoters in human cells. Cell 130(1):77–88
Kimura K et al (2006) Diversification of transcriptional modulation: large-scale identification and characterization of putative alternative promoters of human genes. Genome Res 16(1):55–65
Kim TH et al (2005) A high-resolution map of active promoters in the human genome. Nature 436(7052):876–880
Martianov I et al (2007) Repression of the human dihydrofolate reductase gene by a non-coding interfering transcript. Nature 445(7128):666–670
Acknowledgments
This project was supported by funding from The Danish Cancer Society, The Danish Medical Research Foundation, Fonden til Lægevidenskabens Fremme, Christinan X′s Foundation, The Lundbeck Foundation, and The NovoNordisk Foundation.
Author information
Authors and Affiliations
Corresponding author
Rights and permissions
About this article
Cite this article
Zhou, Z., Kahns, S. & Nielsen, A.L. Identification of a novel vimentin promoter and mRNA isoform. Mol Biol Rep 37, 2407–2413 (2010). https://doi.org/10.1007/s11033-009-9751-8
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s11033-009-9751-8