Abstract
Owing to their superior agronomic performance, the hybrids of vegetable crops are currently applied extensively. However, effective hybrid production requires a laborious manual emasculation to ensure the purity of hybrid seeds in tomato because of the lack of an effective male sterility system. Here, we created two types of tomato nuclear male-sterile lines with different screening markers in a clustered regularly interspaced short palindromic repeats (CRISPR)/CRISPR-associated protein 9 (Cas9) system. Co-knockouts of male sterile 1035 (Ms1035) and glutathione S-transferase (GSTAA) created a male-sterile line marked by a green hypocotyl. The Ms1035 biallelic mutation was introduced into the woolly tomato background, resulting in the linkage of male sterility and a non-woolly phenotype. Two types of male-sterile lines were easily selected at the seedling stage by hypocotyl color or trichome density and further showed high seed purity during hybrid seed production. Our work established the procedure for a rapid transfer of the male-sterile phenotype to the parents of hybrids without extra-modification by the CRISPR/Cas9 system that can be practically applied to hybrid seed production in tomato. This method will be the basis and example for sterile parent creation of multiple crops for hybrid production with the CRISPR/Cas9 system.




Similar content being viewed by others
References
Atanassova B (1999) Functional male sterility (ps-2) in tomato (Lycopesicon esculentum Mill.) and its application in breeding and hybrid seed production. Euphytica 107:13–21. https://doi.org/10.1023/A:1003527714805
Atanassova B (2000) Functional male sterility in tomato (Lycopersicon esculentum Mill.) and its application in hybrid seed production. Acta Physiol Plant 22:221–225. https://doi.org/10.1007/s11738-000-0015-4
Barman HN et al (2019) Generation of a new thermo-sensitive genic male sterile rice line by targeted mutagenesis of TMS5 gene through CRISPR/Cas9 system. BMC Plant Biol 19:109. https://doi.org/10.1186/s12870-019-1715-0
Brooks C, Nekrasov V, Lippman ZB, Van Eck J (2014) Efficient gene editing in tomato in the first generation using the clustered regularly interspaced short palindromic repeats/CRISPR-associated 9 system. Plant Physiol 166:1292–1297. https://doi.org/10.1104/pp.114.247577
Čermák T, Baltes NJ, Čegan R, Zhang Y, Voytas DF (2015) High-frequency, precise modification of the tomato genome. Genome Biol 16:232. https://doi.org/10.1186/s13059-015-0796-9
Chandrasekaran J et al (2016) Development of broad virus resistance in non-transgenic cucumber using CRISPR/Cas9 technology. Mol Plant Pathol 17:1140–1153. https://doi.org/10.1111/mpp.12375
Chang Z et al (2016) Construction of a male sterility system for hybrid rice breeding and seed production using a nuclear male sterility gene. Proc Natl Acad Sci U S A 113:14145–14150. https://doi.org/10.1073/pnas.1613792113
Chen L, Liu Y (2014) Male sterility and fertility restoration in crops. In: Merchant SS (ed) Annual Review of Plant Biology, vol 65, pp 579–606. https://doi.org/10.1146/annurev-arplant-050213-040119
Cigan AM et al (2017) Targeted mutagenesis of a conserved anther-expressed P450 gene confers male sterility in monocots. Plant Biotechnol J 15:379–389. https://doi.org/10.1111/pbi.12633
Cong L et al (2013) Multiplex genome engineering using CRISPR/Cas systems. Science 339:819–823. https://doi.org/10.1126/science.1231143
Crane MB (1915) Heredity of types of inflorescence and fruits in tomato. J Genet 5:1–11. https://doi.org/10.1007/BF02982149
Dellaporta SL, Wood J, Hicks JB (1983) A plant DNA minipreparation: Version II. Plant Mol Biol Report 1:19–21. https://doi.org/10.1007/BF02712670
Du M et al (2020) A biotechnology-based male-sterility system for hybrid seed production in tomato. Plant J 102:1090–1100. https://doi.org/10.1111/tpj.14678
Goldsbrough A, Belzile F, Yoder JI (1994) Complementation of the tomato anthocyanin without (aw) mutant using the dihydroflavonol 4-reductase gene. Plant Physiol 105:491–496. https://doi.org/10.1104/pp.105.2.491
Jeong H et al (2014) Tomato male sterile 10(35) is essential for pollen development and meiosis in anthers. J Exp Bot 65:6693–6709. https://doi.org/10.1093/jxb/eru389
Ji Y et al (2007) Co-dominant SCAR markers for detection of the Ty3 and Ty3a loci from Solanum chilense at 25 cM of chromosome 6 of tomato. Tomato Genet Cooper 57:25–29
Jung YJ et al (2020) Knockout of SlMS10 gene (Solyc02g079810) encoding bHLH transcription factor using CRISPR/Cas9 system confers male sterility phenotype in tomato. Plants (Basel) 9. https://doi.org/10.3390/plants9091189
Kim Y, Zhang D (2018) Molecular control of male fertility for crop hybrid breeding. Trends Plant Sci 23:53–65. https://doi.org/10.1016/j.tplants.2017.10.001
Liu J, Van Eck J, Cong B, Tanksley SD (2002) A new class of regulatory genes underlying the cause of pear-shaped tomato fruit. Proc Natl Acad Sci U S A 99:13302–13306. https://doi.org/10.1073/pnas.162485999
Liu X et al (2019) A putative bHLH transcription factor is a candidate gene for male sterile 32, a locus affecting pollen and tapetum development in tomato. Hortic Res-England 6. https://doi.org/10.1038/s41438-019-0170-2
Maeder ML et al (2008) Rapid “open-source” engineering of customized zinc-finger nucleases for highly efficient gene modification. Mol Cell 31:294–301. https://doi.org/10.1016/j.molcel.2008.06.016
Maloney GS, DiNapoli KT, Muday GK (2014) The anthocyanin reduced tomato mutant demonstrates the role of flavonols in tomato lateral root and root hair development. Plant Physiol 166:254–614. https://doi.org/10.1104/pp.114.240507
Mussolino C, Morbitzer R, Lütge F, Dannemann N, Lahaye T, Cathomen T (2011) A novel TALE nuclease scaffold enables high genome editing activity in combination with low toxicity. Nucleic Acids Res 39:9283–9293. https://doi.org/10.1093/nar/gkr597
Mutschler M, Tanksley S, Rick CM (1987) Linkage maps of the tomato (Lycopersicon esculentum) Vol 37.,
Okada A et al (2019) CRISPR/Cas9-mediated knockout of Ms1 enables the rapid generation of male-sterile hexaploid wheat lines for use in hybrid seed production. Plant Biotechnol J 17:1905–1913. https://doi.org/10.1111/pbi.13106
Park SH, Morris JL, Park JE, Hirschi KD, Smith RH (2003) Efficient and genotype-independent Agrobacterium-mediated tomato transformation. J Plant Physiol 160:1253–1257. https://doi.org/10.1078/0176-1617-01103
Pucci A, Picarella ME, Mazzucato A (2017) Phenotypic, genetic and molecular characterization of 7B-1, a conditional male-sterile mutant in tomato. Theor Appl Genet 130:2361–2374. https://doi.org/10.1007/s00122-017-2964-7
Semagn K, Babu R, Hearne S, Olsen M (2013) Single nucleotide polymorphism genotyping using Kompetitive Allele Specific PCR (KASP): Overview of the technology and its application in crop improvement. Mol Breed 33:1–14. https://doi.org/10.1007/s11032-013-9917-x
Shilling, Richard P (1959) Investigation of the hereditary character, woolly, in the tomato
Song Y, Wang J, Zhang G, Zhao X, Zhang P, Niu N, Ma S (2015) Microspore abortion and abnormal tapetal degeneration in a male-sterile wheat line induced by the chemical hybridizing agent SQ-1. Crop Sci 55:1117–1128. https://doi.org/10.2135/cropsci2014.08.0538
Uluisik S et al (2016) Genetic improvement of tomato by targeted control of fruit softening. Nat Biotechnol 34:950–952. https://doi.org/10.1038/nbt.3602
Valverde P, Fornoni J, Núñez-Farfán J (2001) Defensive role of leaf trichome in resistance to herbivorous in Datura stramonium. J Evol Biol 14:424–432. https://doi.org/10.1046/j.1420-9101.2001.00295.x
Wang S, Zhang S, Wang W, Xiong X, Meng F, Cui X (2015) Efficient targeted mutagenesis in potato by the CRISPR/Cas9 system. Plant Cell Rep 34:1473–1476. https://doi.org/10.1007/s00299-015-1816-7
Woo JW et al (2015) DNA-free genome editing in plants with preassembled CRISPR-Cas9 ribonucleoproteins. Nat Biotechnol 33:1156–1162. https://doi.org/10.1038/nbt.3389
Xing H et al. (2014) A CRISPR/Cas9 toolkit for multiplex genome editing in plants. BMC Plant Biol. 14:327 doi:https://doi.org/10.1186/s12870-014-0327-y
Yang C, Li H, Zhang J, Wang T, Ye Z (2011) Fine-mapping of the woolly gene controlling multicellular trichome formation and embryonic development in tomato. Theor Appl Genet 123:625–633. https://doi.org/10.1007/s00122-011-1612-x
Yang C, Gao Y, Gao S, Yu G, Xiong C, Chang J, Li H, Ye Z (2015) Transcriptome profile analysis of cell proliferation molecular processes during multicellular trichome formation induced by tomato Wo (v) gene in tobacco. BMC Genomics 16:868. https://doi.org/10.1186/s12864-015-2099-7
Ye M et al (2018) Generation of self-compatible diploid potato by knockout of S-RNase. Nat Plants 4:651–654. https://doi.org/10.1038/s41477-018-0218-6
Zhang L, Huang Z, Wang X, Gao J, Guo Y, Du Y, Hu H (2016) Fine mapping and molecular marker development of anthocyanin absent, a seedling morphological marker for the selection of male sterile 10 in tomato. Mol Breed. 36 doi:https://doi.org/10.1007/s11032-016-0531-6
Acknowledgements
We thank Prof Qijun Chen (China Agricultural University, Beijing) for kindly providing the plant genome editing vector pKSE401 and pCBC1-DT1T2 plasmids.
Funding
This work was financially supported by NSFC (No. 31972430) and Research Collaborative Innovation of Yangling Demonstration Zone (2018CXY-09) to Xiaofeng Wang, “Light of the West” Talent Training and Introduction Program of Chinese Academy of Sciences (XAB2018AW16) to Yanfeng Zhang.
Author information
Authors and Affiliations
Contributions
J.L., S.W., H.W., Y.Z., and X.W. designed the studies. J.L., S.W., H.W., B.L., Y.C., and X.L. performed the experiments. J.L., S.W., H.W., Y.Z., and X.W. wrote the manuscript. All the authors gave final approval for submission of the manuscript.
Corresponding authors
Ethics declarations
Ethics approval and consent to participate
Not applicable.
Consent for publication
Not applicable.
Competing interests
The authors declare no competing interests.
Additional information
Publisher’s note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary information
Supplementary Fig. 1
Morphological analysis of Ms1035ms1035/WoDWo and CR-ms1035/Wo. a Two-month-old plants in the field, woollyand non-woolly phenotype plants from two dependent male-sterile lines. b Morphological comparison of Ms1035ms1035/WoDWo and CR-ms1035/Wo flowers; scale bar: 1 cm. c Morphology of Ms1035ms1035/WoDWo and CR-ms1035/Wo anther cones, red arrow indicates exerted stigmata; scale bar: 1 cm. (PNG 2262 kb)
Supplementary Fig. 2
CR-Ms035/Wo and CR-Ms035/gstaa can set normal fruits by artificial pollination. The red arrow indicates the fruit set by artificial pollination. (TIF 6068 kb) (PNG 2090 kb)
Supplementary Fig. 3
Analysis of hypocotyl color and genotype in Ms1035ms1035/F3Hf3h self-crossed progeny. a The second exon of F3H was targeted by the CRISPR/Cas9 system using single-guide RNAs (red arrows indicate target). Black arrows indicate forward (F) and reverse (R) primers used for PCR genotyping and sequencing. The target sequences in F3H are underlined, minus symbols represent deletions. The numbers on the right show the type of mutation and how many nucleotides are involved, with “−” indicating deletion of the given number of nucleotides. b Morphological comparison of wild-type (WT) and CR-ms1035/f3h hypocotyls; scale bar: 1 cm. c Editing analysis of F3H using restriction enzyme site Xba I loss. The two bands and three bands indicate WT and heterozygous, respectively. (PNG 4642 kb)
Supplementary Fig. 4
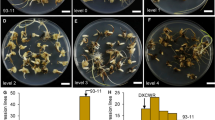
a Morphology of fruits and seeds in 6Q71, 5N21, CR-ms1035/Wo and CR-ms1035/gstaa during tomato seed production. b Germination rate of hybrid F1 seeds from 6Q71, 5N21, CR-ms1035/Wo and CR-ms1035/gstaa. Per plug, 128 seeds were sowing, and the germination rate was counted seven days after sowing. Three plugs were sown for each type of seed as three independent biological repeats. Statistically significant differences were test by Student’s t-test (P < 0.05). (PNG 1007 kb)
Supplementary Table 1.
Detection of mutations in potential off-target sites in T0 edited plants. (PDF 184 kb)
Supplementary Table 2.
Linkage between ms1035 and f3h in the T2 and T3 generations. (PDF 246 kb)
Supplementary Table 3.
Primers used for this study. (PDF 255 kb)
ESM 8
(DOCX 151 kb)
Rights and permissions
About this article
Cite this article
Liu, J., Wang, S., Wang, H. et al. Rapid generation of tomato male-sterile lines with a marker use for hybrid seed production by CRISPR/Cas9 system. Mol Breeding 41, 25 (2021). https://doi.org/10.1007/s11032-021-01215-2
Received:
Accepted:
Published:
DOI: https://doi.org/10.1007/s11032-021-01215-2