Abstract
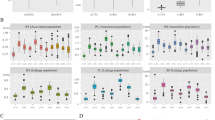
Ramie fiber extracted from stem bast is one of the most important natural fibers. The fiber yield of ramie is a valuable trait and is decided by several components, including stem number per plant (SN), the fiber yield per stem (FYPS), stem length (SL), stem diameter (SD), and bark thickness (BT). All of these fiber yield-related traits are inherited in a quantitative manner. The genetic basis for these traits is still uncharacterized, which has hindered the improvement of yield traits through selective ramie breeding. In this study, an F2 population derived from two ramie varieties, Zhongzhu 1 and Qingyezhuma, with striking differences in fiber yield-related traits, was used for cutting propagation and to develop an F2 agamous line (FAL) population. A genetic linkage map with 132 DNA loci spanning 2,265.1 cM was first constructed. The analysis of quantitative trait locus (QTL) for fiber yield-related traits was performed in ramie for the first time. Finally, a total of 6, 9, 5, 7, and 6 QTLs for FYPS, SL, SN, SD, and BT, respectively, were identified in the FAL population in two environments. Among these 33 QTLs, 9 QTLs were detected in both environments and 24 QTLs exhibited overdominance. The overdominance of these QTLs possibly contributed to the heterosis of these yield-related traits in ramie. Moreover, there were 7 QTL clusters identified. The identification of the QTLs for fiber yield-related traits will be helpful for improving the fiber yield in ramie breeding programs.


Similar content being viewed by others
References
Chen M, Presting G, Barbazuk W, Goicoechea J, Blackmon B, Fang G, Kim H, Frisch D, Yu Y, Sun S et al (2002) An Integrated Physical and Genetic Map of the Rice Genome. Plant Cell 14:537–545
Chen J, Luan M, Song S, Zou Z, Wang X, Xu Y, Sun Z (2011) Isolation and characterization of EST-SSRs in the Ramie. Afr J Microbiol Res 5:3504–3508
Fan C, Xing Y, Mao H, Lu T, Han B, Xu C, Li X, Zhang Q (2006) GS3, a major QTL for grain length and weight and minor QTL for grain width and thickness in rice, encodes a putative transmembrane protein. Theor Appl Genet 112:1164–1171
Frary A, Nesbitt T, FraryA Grandillo S, Knaap E, Cong B, Liu J, Meller J, Elber R, Alpert K et al (2000) fw2.2: A quantitative trait locus key to the evolution of tomato fruit size. Science 289:85–88
Grattapaglia D, Sederoff R (1994) Genetic Linkage maps of Eucalyptus grandis and Eucalyptus urophylla using a pseudo-testcross: mapping strategy and RAPD markers. Genetics 137:1121–1137
Groos C, Robert N, Bervas E, Charmet G (2003) Genetic analysis of grain protein-content, grain yield and thousand-kernel weight in bread wheat. Theor Appl Genet 106:1032–1040
Guo Q, Liu F (1989) A study on variation and segregation of self-bred progeny of ramie. J Hunan Agric Coll 15(S1):54–59
Hua J, XingY WuW, Xu C, Sun X, Yu S, Zhang Q (2003) Single-locus heterotic effects and dominance by dominance interactions can adequately explain the genetic basis of heterosis in an elite rice hybrid. Proc Natl Acad Sci USA 100:2574–2579
Jones DF (1917) Dominance of linked factors as a means of accounting for heterosis. Genetics 2:466–479
Kusterer B, Muminovic J, Utz H, Piepho H, Barth S, Heckenberger M, Meyer R, Altmann T, Melchinger A (2007) Analysis of a triple testcross design with recombinant inbred lines reveals a significant role of epistasis in heterosis for biomass-related traits in Arabidopsis. Genetics 175:2009–2017
Li Z, Luo L, Mei H, Wang D, Shu Q, Tabien R, Zhong D, Ying C, Stansel J, Khush G et al (2001) Overdominant epistatic loci are the primary genetic basis of inbreeding depression and heterosis in rice. I. Biomass and grain yield. Genetics 158:1737–1753
Li L, Lu K, Chen Z, Mu T, Hu Z, Li X (2008) Dominance, overdominance and epistasis condition the heterosis in two heterotic rice hybrids. Genetics 180:1725–1742
Lincoln SE, Daly MJ, Lander ES (1993) Mapping genes controlling quantitative traits with MAPMAKER/QTL1.1: a tutorial and reference manual, 2nd edn. Whitehead Institute Technical Report, Cambridge
Liu T, Mao D, Zhang S, Xu C, Xing Y (2009) Fine mapping SPP1, a QTL controlling the number of spikelets per panicle, to a BAC clone in rice (Oryza sativa). Theor Appl Genet 118:1509–1517
Liu T, Zhang Y, Xue W, Xu C, Li X, Xing Y (2010) Comparison of quantitative trait loci for 1,000-grain weight and spikelets per panicle across three connected rice populations. Euphytica 175:383–394
Liu T, Li L, Zhang Y, Xu C, Li X, Xing Y (2011) Comparison of quantitative trait loci for rice yield, panicle length and spikelet density across three connected populations. J Genet 90:377–382
Liu T, Liu H, Zhang H, Xing Y (2013a) Validation and characterization of Ghd7.1, a major QTL with pleiotropic effects on spikelets per panicle, plant height, and heading date in rice (Oryza sativa L.). J Integr Plant Biol 55:917–927
Liu T, Zhu S, Fu L, Tang Q, Yu Y, Chen P, Luan M, Wang C, Tang S (2013b) Development and characterization of 1827 expressed sequence tag-derived simple sequence repeat markers in ramie (Boehmeria nivea L. Gaud). PLoS ONE 8:e60346
Liu T, Zhu S, Tang Q, Chen P, Yu Y, Tang S (2013c) De novo assembly and characterization of transcriptome using Illumina paired-end sequencing and identification of CesA gene in ramie (Boehmeria nivea L. Gaud). BMC Genom 14:125
Liu T, Zhu S, Tang Q, Yu Y, Tang S (2013d) Identification of drought stress-responsive transcription factors in ramie (Boehmeria nivea L. Gaud). BMC Plant Biol 13:130
Luo L, Li Z, Mei H, Shu Q, Tabien R, Zhong D, Ying C, Stansel J, Khush G, Paterson A (2001) Overdominant epistatic loci are the primary genetic basis of inbreeding depression and heterosis in rice. II. Grain yield components. Genetics 158:1755–1771
Messmer R, Fracheboud Y, Bänziger M, Vargas M, Stamp P, Ribaut J (2009) Drought stress and tropical maize: QTL-by-environment interactions and stability of QTLs across environments for yield components and secondary traits. Theor Appl Genet 119:913–930
Ni J, Pujar A, Youens-Clark K, Yap I, Jaiswal P, Tecle I, Tung C, Ren L, Spooner W, Wei X, et al (2009) Gramene QTL database: development, content and applications. Database 2009:bap005
Paterson A, Lander E, Hewitt J, Peterson S, Lincoln S, Tanksley S (1988) Resolution of quantitative traits into Mendelian factors by using a complete linkage map of restriction fragment length polymorphisms. Nature 335:721–726
Paterson A, Saranga Y, Menz M, Jiang C, Wright R (2003) QTL analysis of genotype × environment interactions affecting cotton fiber quality. Theor Appl Genet 106:384–396
Salvi S, Tuberosa R, Chiapparino E, Maccaferri M, Veillet S, Beuningen L, Isaac P, Edwards K, Phillips R (2002) Toward positional cloning of Vgt1, a QTL controlling the transition from the vegetative to the reproductive phase in maize. Plant Mol Biol 48:601–613
Schnell FW, Cockerham CC (1992) Multiplicative vs. arbitrary gene action in heterosis. Genetics 131:461–469
Semel Y, Nissenbaum J, Menda N, Zinder M, Krieger U, Issman N, Pleban T, Lippman Z, Gur A, Zamir D (2006) Overdominant quantitative trait loci for yield and fitness in tomato. Proc Natl Acad Sci USA 103:12981–12986
Shull GH (1908) The composition of a field of maize. Am Breeders Assoc Rep 4:296–301
Stuber CW, Lincoln SE, Wolff DW, Helentjaris T, Lander ES (1992) Identification of genetic factors contributing to heterosis in a hybrid from two elite maize inbred lines using molecular markers. Genetics 132:823–839
Tang J, Yan J, Ma X, Teng W, Wu W, Dai J, Dhillon B, Melchinger A, Li J (2010) Dissection of the genetic basis of heterosis in an elite maize hybrid by QTL mapping in an immortalized F2 population. Theor Appl Genet 120:333–340
Wang S, Basten C, Zeng Z (2012) Windows QTL Cartographer 2.5. Department of Statistics, North Carolina State University, Raleigh, NC. (http://statgen.ncsu.edu/qtlcart/WQTLCart.htm)
Welter L, Gokturk-Baydar N, Akkurt M, Maul E, Eibach R, Topfer R, Zyprian E (2007) Genetic mapping and localization of quantitative trait loci affecting fungal disease resistance and leaf morphology in grapevine (Vitis vinifera L). Mol Breed 20:359–374
Wu KS, Tanksley SD (1993) Abundance, polymorphism and genetic mapping of microsatellites in rice. Mol Gen Genet 241:225–235
Xiao J, Li J, Yuan L, Tanksley SD (1995) Dominance is the major genetic basis of heterosis in rice as revealed by QTL analysis using molecular markers. Genetics 140:745–754
Xiong H, Jiang J, Yu C, Guo Y (1998) Relation between yield-related traits and yield in ramie. ACTA Agron Sinica 24:155–160
Xue W, Xing Y, Weng X, Zhao Y, Tang W, Wang L, Zhou H, Yu S, Xu C, Li X et al (2008) Natural variation in Ghd7 is an important regulator of heading date and yield potential in rice. Nat Genet 40:761–767
Yan W, Liu H, Zhou X, Li Q, Zhang J, Lu L, Liu T, Liu H, Zhang C, Zhang Z et al (2013) Natural variation in Ghd7.1 plays an important role in grain yield and adaptation in rice. Cell Res 23:969–971
Yu S, Li J, Xu C, Tan Y, Gao Y, Li X, Zhang Q (1997) Importance of epistasis as the genetic basis of heterosis in an elite rice hybrid. Proc Natl Acad Sci USA 94:9226–9231
Zhou G, Chen Y, Yao W, Zhang C, Xie W, Hua J, Xing Y, Xiao J, Zhang Q (2012) Genetic composition of yield heterosis in an elite rice hybrid. Proc Natl Acad Sci USA 109:15847–15852
Acknowledgments
This work was supported by grants from the National Natural Science Foundation of China (31101189).
Author information
Authors and Affiliations
Corresponding author
Electronic supplementary material
Below is the link to the electronic supplementary material.
Rights and permissions
About this article
Cite this article
Liu, T., Tang, S., Zhu, S. et al. QTL mapping for fiber yield-related traits by constructing the first genetic linkage map in ramie (Boehmeria nivea L. Gaud). Mol Breeding 34, 883–892 (2014). https://doi.org/10.1007/s11032-014-0082-7
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s11032-014-0082-7