Abstract
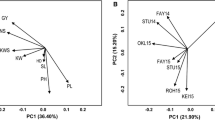
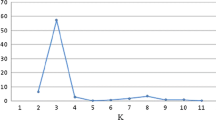
To make advances in rice breeding it is important to understand the relatedness and ancestry of introduced rice accessions, and identify SSR markers associated with agronomically important phenotypic traits, for example yield. Ninety-two rice germplasm accessions recently introduced from seven geographic regions of Africa, Asia, and Latin America, and eleven US cultivars, included as checks, were evaluated for yield and kernel characteristics, and genotyped with 123 SSR markers. The SSR markers were highly polymorphic across all accessions. Population structure analysis identified eight main clusters for the accessions which corresponded to the major geographic regions, indicating agreement between genetic and predefined populations. Linkage disequilibrium (LD) patterns and distributions are of fundamental importance for genome-wide mapping association. LD between linked markers decreased with distance and with a substantial drop in LD decay values between 20 and 30 cM, suggesting it should be possible to achieve resolution down to the 25 cM level. For the 103 cultivars, the complex traits yield, kernel width, kernel length, kernel width/length ratio, and 1000-kernel weight, were estimated by analysis of variety trial data. The mixed linear model method was used to disclose marker-trait associations. Many of the associated markers were located in regions where QTL had previously been identified. In conclusion, association mapping in rice is a viable alternative to QTL mapping based on crosses between different lines.




Similar content being viewed by others
Abbreviations
- SSR:
-
Simple sequence repeat
- QTL:
-
Quantitative trait loci
- cM:
-
CentiMorgan
References
Akkaya MS, Bhagwat AA, Cregan PB (1992) Length polymorphisms of simple sequence repeat DNA in soybean. Genetics 132:1131–1139
Abdallah JM, Goffinet B, Ayrolles CC, Pérez-Enciso M (2003) Linkage disequilibrium fine mapping of quantitative trait loci: a simulation study. Genet Sel Evol 35:513–532
Ardlie KG, Kruglyak L, Seielstad M (2002) Patterns of linkage disequilibrium in the human genome. Nat Rev Genet 3:299–309
Breseghello F, Sorrells ME (2006) Association mapping of kernel size and milling quality in wheat (Triticum aestivum L.) cultivars. Genetics 172:1165–1177
Cardon LR, Bell JI (2001) Association study designs for complex diseases. Nature Rev Genet 2:91–98
Chen X, Cho YG, McCouch SR (2002) Sequence divergence of rice microsatellites in Oryza and other plant species. Mol Genet Genomics 268:331–343
Ching A, Caldwell KS, Jung M, Dolan M, Smith OS, Tingey S, Morgante M, Rafalski A (2002) SNP frequency, haplotype structure and linkage disequilibrium in elite maize inbred lines. BMC Genet 3:19. pp 1–14
Cho YG, Ishii T, Temnykh S, Chen X, Lipovich L, McCouch SR, Park WD, Ayres N, Cartinhour S (2000) Diversity of microsatellites derived from genomic libraries and GenBank sequences in rice (Oryza sativa L.). Theor Appl Genet 100:713–722
Condon F, Smith K (2005) Linkage disequilibrium and marker-trait associations within six-rowed barley breeding germplasm. Plant & Animal Genomes XIII Conference. January 15–19, 2005
Dilday RH, Yan W, Moldenhauer KAK, Gibbons JW, Lee FN, Bryant RJ (2001) Chinese and other foreign germplasm evaluation. In: Norman RJ, Meullenet J-F (eds) B.R. Wells Rice Research Studies 2000. University of Arkansas, Agricultural Experiment Station. Research Series 485:1–12
Eaves IA, Barber RA, Merriman TR (1998) Comparison of linkage disequilibrium in populations from UK and Finland. Am J Hum Gen A221
Eizenga GC, Agrama HA, Lee FN, Yan W, Jia Y (2006) Identifying novel resistance genes in newly introduced Blast resistant rice germplasm. Crop Sci 46:1870–1878
Excoffier L, Laval G, Schneider S (2005) Arlequin ver. 3.0: An integrated software package for population genetics data analysis. Evol Bioinformatics Online 1:47–50
Estoup A, Tailliez C, Cornuet JM, Solignac M (1995) Size homoplasy and mutational processes of interrupted microsatellites in two bee species, Apis mellifera and Bombus terrestris (Apidae). Mol Biol Evol 12:1074–1084
Farnir F, Coppieters W, Arranz W, Berzi P, Cambisano N, Grisart B, Karim L, Marcq F, Moreau L, Mni M, Nezer C, Simon P, Vanmanshoven P, Wagenaar D, Georges M (2000) Extensive genome-wide linkage disequilibrium in cattle. Genome Res 10:220–227
Falush D, Stephens M, Pritchard JK (2003) Inference of population structure using multilocus genotype data: linked loci and correlated allele frequencies. Genetics 164:1567–1587
Fjellstrom RG, Conaway-Bormans CA, McClung AM, Marchetti MA, Shank AR, Park WD (2004) Development of DNA markers suitable for marker assisted selection of three Pi genes conferring resistance to multiple Pyricularia grisea pathotypes. Crop Sci 44:1790–1798
Flint-Garcia SA, Thuillet AC, Yu J, Pressoir G, Romero SM, Mitchell S, Doebley J, Kresovich S, Goodman MM, Buckler ES (2005) Maize association population: a high-resolution platform for quantitative trait locus dissection. Plant J 44:1054–1064
Foster M, Sharp RR (2002) Race, ethnicity, and genomics: social classifications as proxies of biological heterogeneity. Genome Res 12:844–850
Gao LZ, Ge S, Hong DY (2000) Low levels of genetic diversity within populations and high genetic differentiation among populations of a wild rice, Oryza granulate Nees et Arn. Ex Watt. From China. Int J Plant Sci 161:691–697
Gao LZ, Cheng W, Ge S, Hong DY, Jiang WZ, Wang W (2001) Genetic erosion in northern marginal population of common wild rice Oryza rufipogon Griff. and its conservation, revealed by allozyme analysis. Hereditas 133:47–53
Gao LZ, Schaal BA, Zhang CH, Jia JZ (2002) Assessment of population genetic structure in common wild rice Oryza rufipogon Griff. Using microsatellite and allozyme markers. Theor Appl Genet 106:173–180
Gao LZ, Zhang CH (2005) Comparisons of microsatellite variability and population genetic of two endangered wild rice species, Oryza rufipogon and O. officinalis, and their conservation implications. Biodivers Conserv 14:1663–1679
Garris A, McCouch SR, Kresovich S (2003) Population structure and its effect on haplotype diversity and linkage disequilibrium surrounding the xa5 locus of rice (Oryza sativa L.). Genetics 165:759–769
Garris A, Tai T, Coburn J, Kresovich S, McCouch S (2005) Genetic structure and diversity in Oryza sativa L. Genetics 169:1631–1638
Ge XJ, Xing YZ, Xu CG, He YQ (2005) QTL analysis of cooked rice grain elongation, volume expansion, and water absorption using a recombinant inbred population. Plant Breed 124:121–126
Glaszmann JC (1987) Isozymes and classification of Asian rice varieties. Theor Appl Genet 74:21–30
Guo LB, Xing YZ, Mei HW, Xu CG, Shi CH, Wu P, Luo LJ (2005) Dissection of component QTL expression in yield formation in rice. Plant Breed 124:127–132
Gupta PK, Rustgi S, Kulwal PL (2005) Linkage disequilibrium and association studies in higher plants: present status and future prospects. Plant Mol Biol 57:461–485
Hardy OJ, Vekemans X (2002) SPAGeDi: a versatile computer program to analyze spatial genetic structure at the individual or population levels. Mol Ecol Notes 2:618–620
Hill WG, Robertson A (1968) Linkage disequilibrium in finite populations. Theor Appl Genet 38:226–231
Hittalmani S, Shashidhar HE, Bagali PG, Huang N, Sidhu JS, Singh VP, Khush GS (2002) Molecular mapping of quantitative trait loci for plant growth, yield and yield related traits across three diverse locations in a doubled haploid rice population. Euphytica 125:207–214
Ishii T, McCouch SR (2000) Microsatellites and microsynteny in the chloroplast genomes of Oryza and eight other Gramineae species. Theor Appl Genet 100:1257–1266
Ishii T, Xu Y, McCouch SR (2001) Nuclear- and chloroplast-microsatellite variation in A-genome species of rice. Genome 44:658–666
Jannink JL, Walsh B (2002) Association mapping in plant populations. In: Kang MS (ed) Quantitative genetics, genomics and plant breeding. CAB International
Jing YH, Sun CQ, Tan LB, Fu YC, Zhang PJ, Xu ZJ, Chen WF, Wang XK (2005) Mapping QTLs controlling vascular bundle and panicle-related traits from Yuanjiang common wild rice (Oryza rufipogon Griff.). Acta-Genetica-Sinica 32:178–182
Jorde LR (1995) Linkage disequilibrium as a gene-mapping tool. Am J Hum Genet 56:11–14
Jorde LR (2000) Linkage disequilibrium and the search for complex disease genes. Genome Res 10:1435–1444
Kraakman ATW, Niks RE, van den Berg PM, Stam P, van Eeuwijk FA (2004) Linkage disequilibrium mapping of yield and yield stability in modern spring barley cultivars. Genetics 168:435–446
Kraakman ATW, Martínez F, Mussiraliev B, van Eeuwijk FA, Niks RE (2006) Linkage disequilibrium mapping of morphological, resistance, and other agronomically relevant traits in modern spring barley cultivars. Mol Breed 17:41–58
Kraft T, Hansen M, Nilsson N-O (2000) Linkage disequilibrium and fingerprinting in sugar beet. Theor Appl Genet 101:323–326
Kruger SA, Able JA, Chalmers KJ, Langridge P (2004) Linkage disequilibrium analysis of hexaploid wheat. In: Plant & Animal Genomes XII Conference, 10–14 January, Town & Country Convention Center, San Diego, CA, p 321
Kruglyak L (1999) Prospects for whole-genome linkage disequilibrium mapping of common disease genes. Nat Genet 22:139–144
Lander ES, Schork NJ (1994) Genetic dissection of quantitative traits. Science 256:2037–2048
Li J, Xiao J, Grandillo S, Jiang L, Wan Y, Deng Q, Yuan L, McCouch SR (2004) QTL detection for rice grain quality traits using an interspecific backcross population derived from cultivated Asian (O. sativa L.) and African (O. glaberrima S.) rice. Genome 47:697–704
Li ZK, Yu SB, Lafitte HR, Huang N, Courtois B, Hittalmani S, Vijayakumar CHM, Liu GF, Wang GC, Shashidhar HE, Zhuang JY, Zheng KL, Singh VP, Sidhu JS, Srivantaneeyakul S, Khush GS (2003) QTL × environment interactions in rice. I. Heading date and plant height. Theor Appl Genet 108:141–153
Liu K, Muse S (2004) PowerMarker: new genetic data analysis software. Version 3.0. Free program distributed by the author over the internet from http://www.powermarker.net
Lu H, Redus MA, Coburn JR, Rutger JN, McCouch SR, Tai TH (2005) Population structure and breeding patterns of 145 US rice cultivars based on SSR marker analysis. Crop Sci 45:66–76
Maccaferri M, Sanguineti MC, Noli E, Tuberosa R (2005) Population structure and long-range linkage disequilibrium in a durum wheat elite collection. Mol Breed 15:271–289
Mackill DJ (1995) Classifying japonica rice cultivars with RAPD markers. Crop Sci 35:889–894
Mather DE, Hyes PM, Chalmers KJ, Eglinton J, Matus I, Richardson K, VonZitzewitz J, Marquez-Cedillo L, Hearnden P, Pal N (2004) Use of SSR marker data to study linkage disequilibrium and population structure in Hordeum vulgare: Prospects for association mapping in barley. In: International barley genetics symposium, Brno, Czech Republic, 20–26 June 2004. pp 302–307
McCouch SR, Chen X, Panaud O, Temnykh S, Xu Y, Cho YG, Huang N, Ishii T, Blair MW (1997) Microsatellite marker development, mapping and applications in rice genetics and breeding. Plant Mol Biol 35:89–99
McCouch SRM, Teytelman L, Xu Y, Lobos KB, Clare K, Walton M, Fu B, Maghirang R, Li Z, Xing Y, Zhang Q, Kono I, Yano M, Fjellstrom R, DeClerck G, Schneider D, Carinhour S, Ware D, Stein L (2002) Development and mapping of 2240 new SSR markers for rice (Oryza sativa L.). DNA Res 9:199–207
McRae AF, McEwan JC, Dodds KG, Wilson T, Crawford AM, Slate J (2002) Linkage disequilibrium in domestic sheep. Genetics 160:1113–1122
N’Goran JAK, Laurent V, Risterucci AM, Lanaud C (2000) The genetic structure of cocoa populations (Theobroma cacao L.) revealed by RFLP analysis. Euphytica 115:83–90
Ni J, Colowit PM, Mackill DJ (2002) Evaluation of genetic diversity in rice subspecies using microsatellite markers. Crop Sci 42:601–607
Nordborg M, Borevitz JO, Bergelson J, Berry CC, Chory J, Hagenblad J, Kreitman M, Maloof JN, Noyes T, Oefner PJ, Stahl EA, Weigel D (2002) The extent of linkage disequilibrium in Arabidopsis thaliana. Nat Genet 30:190–193
Nordborg M, Hu TT, Ishino Y, Jhaveri J, Toomanjian C, Zheng H, Bakker E, Calabrese P, Gladstone J, Goyal R, Jakobsson M, Kim S, Morozov Y, Padhukasahasram B, Plagnol V, Rosenberg NA, Shah C, Wall JD, Wang J, Zhao K, Kalbfeisch T, Schulz V, Kreitman M, Bergelson J (2005) The pattern of polymorphism in Arabidopsis thaliana. PLoS Biol 3:1289–1299
Olsen KM, Caicedo AL, Polato N, McClung A, McCouch S, Purugganan D (2006) Selection under domestication: evidence for a sweep in the rice Waxy genomic region. Genetics 173:975–983
Parsons BJ, Newbury HJ, Jackcon MT, Ford-Lloyd BV (1999) The genetic structure and conservation of aus, aman and bro rices from Bangladesh. Genet Res Crop Evol 46:587–598
Pritchard JK, Rosenberg NA (1999) Use of unlinked genetic markers to detect population stratification in association studies. Am J Hum Genet 65:220–228
Pritchard JK, Donnelly P (2001) Case-control studies of association in structured or admixed populations. Theor Popul Biol 60:227–237
Pritchard JK, Stephens M, Donnelly P (2000) Inference of population structure using multilocus genotype data. Genetics 155:945–959
Remington DL, Thornsberry JM, Matsuola Y, Wilson LM, Whitt SR, Doebley J, Kresovich S, Goodman MM, Buckler IV ES (2001) Structure of linkage disequilibrium and phenotypic associations in the maize genome. Proc Natl Acad Sci USA 98:11479–11484
Risch N, Merikangas K (1996) The future of genetic studies of complex human diseases. Science 273:1516–1517
Risch NJ (2000) Searching for genetic determination for the new millennium. Nature 405:847–856
Rosenberg NA, Pritchard JK, Weber JL, Cann HM, Kidd KK, Zhivotovsky LA, Feldman MW (2002) Genetic structure of human populations. Science 298:2381–2385
Skøt L, Humphreys MO, Armstead I, Heywood S, Skøt KP, Sanderson R, Thomas ID, Chorlton KH, Hamilton NRS (2005) An association mapping approach to identify flowering time genes in natural populations of Lolium perenne (L.). Mol Breed 15:233–245
Smith MW, O’Brien SJ (2005) Mapping by admixture linkage disequilibrium: advances, limitations and guidelines. Nat Rev (online) doi:10.1038/nrgl1657
Stich B, Melchinger AE, Frisch M, Maurer HP, Heckenberger M, Reif JC (2005) Linkage disequilibrium in European elite maize germplasm investigated with SSRs. Theor Appl Genet 111:723–730
Thornsberry JM, Goodman MM, Doebley J, Kresovich S, Nielsen D, Buckler IV ES (2001) Dwarf8 polymorphisms associate with variation in flowering time. Nat Genet 28:286–289
Viard F, Franck P, Dubois MP, Estoup A, Jarne P (1998) Variation of microsatellite size homoplasy across electromorphs, loci, and populations in three invertebrate species. J Mol Evol 47:42–51
Watkins WS, Rogers AR, Ostler CT, Wooding S, Bamshad MJ, Brassington A-ME, Carroll ML, Nguyen SV, Walker JA, Prasad BVR, Reddy PG, Das PK, Batzer MA, Jorde1 LB (2003) Genetic variation among world populations: inferences from 100 Alu insertion polymorphisms. Genome Res 13:1607–1618
Weber J, May PE (1989) Abundant class of human DNA polymorphisms which can be typed using the polymerase chain reaction. Am J Hum Genet 44:388–396
Weir BS, Cockerham CC (1984) Estimating F-statistics for the analysis of population structure. Evolution 38:1358–1370
Weir BS (1996) Genetic data analysis II: Methods for discrete population genetic data. Sinauer Assoc., Inc., Sunderland, MA, USA
Wilson LM, Whitt SR, Ibáñez AM, Rocheford TR, Goodman MM, Buckler IV ES (2004) Dissection of maize kernel composition and starch production by candidate gene association. Plant Cell 16:2719–2733
Yan WG, Rutger JN, Bryant RJ, Lee FN, Gibbons JW (2003) Characteristics of newly-introduced accessions in the USDA-ARS rice quarantine program In: Norman RJ, Meullenet J-F (eds) B.R. Wells Rice Research Studies 2002. Univ. of Arkansas Agric Exp Stn Res Ser 504:112–124
Yang GP, Saghai Maroof MA, Xu CG, Zhang Q, Biyashev RM (1994) Comparative analysis of microsatellite DNA polymorphism in landraces and cultivars of rice. Mol Gen Genet 245:187–194
Yu J, Buckler ES (2006) Genetic association mapping and genome organization of maize. Curr Opin Biotech 17:155–160
Yu J, Pressoir G, Briggs WH, Bi IV, Yamasaki M, Doebley J, McMullen MD, Gaut BS, Nielsen DM, Holland JB, Kresovich S, Buckler ES (2006) A unified mixed-model method for association mapping that accounts for multiple levels of relatedness. Nat Genet 38:203–208
Zhang N, Xu Y, Akash M, McCouch S, Oard JH (2005) Identification of candidate markers associated with agronomic traits in rice using discriminant analysis. Theor Appl Genet 110:721–729
Acknowledgement
The authors acknowledge the support of H. Raeann Refeld and Dr Hesham A. Agrama from the Arkansas Rice Research and Promotion Board. Technical contributions to this research were made by H. Raeann Refeld and Quynh P. Ho. Contributions of Melissa H. Jia, Gordon H. Miller and the late Mark A. Redus of the DB NRRC Genomics Core Facility also are acknowledged.
Author information
Authors and Affiliations
Corresponding author
Rights and permissions
About this article
Cite this article
Agrama, H.A., Eizenga, G.C. & Yan, W. Association mapping of yield and its components in rice cultivars. Mol Breeding 19, 341–356 (2007). https://doi.org/10.1007/s11032-006-9066-6
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s11032-006-9066-6