Abstract
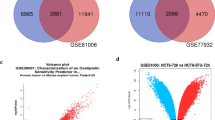
Colorectal cancer (CRC) is the third most diagnosed and highly fatal malignancy, presenting serious health concerns worldwide. The search for an effective cure for CRC is challenging and poses a serious concern. Kaempferol is a potent anti-cancerous bioactive compound often suggested for treating various cancers, including CRC. However, its underlying molecular mechanism against CRC remains unclear. The present study delves into kaempferol's molecular pathways and underlying molecular mechanisms against CRC targets. The target protein-coding genes for kaempferol were retrieved, and the CRC-associated genes were curated. Twelve common targets with a disease specificity index of > 0.6 were validated for their protein expression at different stages of CRC. Over-expressed USP1, SETD7, POLH, TDP1 and RACGAP1 were selected for further studies. The binding affinities of kaempferol to the corresponding proteins were evaluated using molecular docking and Molecular Dynamics (MD) simulations. SETD7 exhibited the highest binding affinity with the lowest binding energy (− 8.06 kcal/mol). Additionally, the MD simulation, and MM-PBSA conferred SETD7-kaempferol complex had the least root-mean-square deviation with lower interaction energy and higher conformational stability. The protein–protein interaction of SETD7 constructed revealed direct interactors, namely, DNMT1, FOXO1, FOXO3, FOXO4, H3-3B, H3-4, H3C12, H3C13, SETD7, SIRT1 and TP53, have a potential role in cancer progression through FOXO signalling. In summary, our study revealed kaempferol's multi-target and synergistic effect on multiple CRC targets and its underlying mechanisms. Finally, the study recommends in-vitro and in-vivo trials for validation of anti-cancerous drugs for CRC.








Similar content being viewed by others
Data availability
All data generated during this study are included in this article.
References
Spaander MCW, Zauber AG, Syngal S et al (2023) Young-onset colorectal cancer. Nat Rev Dis Prim 9:21. https://doi.org/10.1038/s41572-023-00432-7
Krasteva N, Georgieva M (2022) Promising therapeutic strategies for colorectal cancer treatment based on nanomaterials. Pharmaceutics 14:1213. https://doi.org/10.3390/pharmaceutics14061213
Negarandeh R, Salehifar E, Saghafi F et al (2020) Evaluation of adverse effects of chemotherapy regimens of 5-fluoropyrimidines derivatives and their association with DPYD polymorphisms in colorectal cancer patients. BMC Cancer 20:560. https://doi.org/10.1186/s12885-020-06904-3
Bousbaa H (2021) Novel anticancer strategies. Pharmaceutics 13:275. https://doi.org/10.3390/pharmaceutics13020275
Esmeeta A, Adhikary S, Dharshnaa V et al (2022) Plant-derived bioactive compounds in colon cancer treatment: an updated review. Biomed Pharmacother 153:113384. https://doi.org/10.1016/j.biopha.2022.113384
Huang X, Yang Z, Xie Q et al (2019) Natural products for treating colorectal cancer: a mechanistic review. Biomed Pharmacother 117:109142. https://doi.org/10.1016/j.biopha.2019.109142
Roy A, Datta S, Bhatia KS et al (2022) Role of plant derived bioactive compounds against cancer. South African J Bot 149:1017–1028. https://doi.org/10.1016/j.sajb.2021.10.015
Benarba B, Pandiella A (2018) Colorectal cancer and medicinal plants: principle findings from recent studies. Biomed Pharmacother 107:408–423. https://doi.org/10.1016/j.biopha.2018.08.006
Macharia JM, Mwangi RW, Rozmann N et al (2022) Medicinal plants with anti-colorectal cancer bioactive compounds: potential game-changers in colorectal cancer management. Biomed Pharmacother 153:113383. https://doi.org/10.1016/j.biopha.2022.113383
Li X, Khan I, Huang G et al (2022) Kaempferol acts on bile acid signaling and gut microbiota to attenuate the tumor burden in ApcMin/+ mice. Eur J Pharmacol 918:174773. https://doi.org/10.1016/j.ejphar.2022.174773
Wang L, Tu Y-C, Lian T-W et al (2006) Distinctive antioxidant and antiinflammatory effects of flavonols. J Agric Food Chem 54:9798–9804. https://doi.org/10.1021/jf0620719
Ginwala R, Bhavsar R, Chigbu DI et al (2019) Potential role of flavonoids in treating chronic inflammatory diseases with a special focus on the anti-inflammatory activity of apigenin. Antioxidants 8:35. https://doi.org/10.3390/antiox8020035
Maleki SJ, Crespo JF, Cabanillas B (2019) Anti-inflammatory effects of flavonoids. Food Chem 299:125124. https://doi.org/10.1016/j.foodchem.2019.125124
Luo H, Rankin GO, Liu L et al (2009) Kaempferol inhibits angiogenesis and VEGF expression through both HIF dependent and independent pathways in human ovarian cancer cells. Nutr Cancer 61:554–563. https://doi.org/10.1080/01635580802666281
Seifried HE, Anderson DE, Fisher EI, Milner JA (2007) A review of the interaction among dietary antioxidants and reactive oxygen species. J Nutr Biochem 18:567–579. https://doi.org/10.1016/j.jnutbio.2006.10.007
Luo H, Jiang B-H, King SM, Chen YC (2008) Inhibition of cell growth and VEGF expression in ovarian cancer cells by flavonoids. Nutr Cancer 60:800–809. https://doi.org/10.1080/01635580802100851
Nguyen TTT, Tran E, Ong CK et al (2003) Kaempferol-induced growth inhibition and apoptosis in A549 lung cancer cells is mediated by activation of MEK-MAPK. J Cell Physiol 197:110–121. https://doi.org/10.1002/jcp.10340
Chen AY, Chen YC (2013) A review of the dietary flavonoid, kaempferol on human health and cancer chemoprevention. Food Chem 138:2099–2107. https://doi.org/10.1016/j.foodchem.2012.11.139
Wu H, Du J, Li C et al (2022) Kaempferol can reverse the 5-Fu resistance of colorectal cancer cells by inhibiting PKM2-mediated glycolysis. Int J Mol Sci 23:3544. https://doi.org/10.3390/ijms23073544
Riahi-Chebbi I, Souid S, Othman H et al (2019) The Phenolic compound Kaempferol overcomes 5-fluorouracil resistance in human resistant LS174 colon cancer cells. Sci Rep 9:195. https://doi.org/10.1038/s41598-018-36808-z
Nirmala P, Ramanathan M (2011) Effect of kaempferol on lipid peroxidation and antioxidant status in 1,2-dimethyl hydrazine induced colorectal carcinoma in rats. Eur J Pharmacol 654:75–79. https://doi.org/10.1016/j.ejphar.2010.11.034
Li L, Wang X, Guo X et al (2022) Network pharmacology and computer-aided drug design to explored potential targets of Lianhua Qingwen and Qingfei Paidu decoction for COVID-19. Front Pharmacol. https://doi.org/10.3389/fphar.2022.1013428
Noor F, Tahir ul Qamar M, Ashfaq UA et al (2022) Network pharmacology approach for medicinal plants: review and assessment. Pharmaceuticals 15:572. https://doi.org/10.3390/ph15050572
Li J-X, Li R-Z, Sun A et al (2021) Metabolomics and integrated network pharmacology analysis reveal Tricin as the active anti-cancer component of Weijing decoction by suppression of PRKCA and sphingolipid signaling. Pharmacol Res 171:105574. https://doi.org/10.1016/j.phrs.2021.105574
Mathpal S, Sharma P, Joshi T et al (2022) Identification of zinc-binding inhibitors of matrix metalloproteinase-9 to prevent cancer through deep learning and molecular dynamics simulation approach. Front Mol Biosci. https://doi.org/10.3389/fmolb.2022.857430
Gaulton A, Bellis LJ, Bento AP et al (2012) ChEMBL: a large-scale bioactivity database for drug discovery. Nucleic Acids Res 40:D1100–D1107. https://doi.org/10.1093/nar/gkr777
Bento AP, Gaulton A, Hersey A et al (2014) The ChEMBL bioactivity database: an update. Nucleic Acids Res 42:D1083–D1090. https://doi.org/10.1093/nar/gkt1031
Piñero J, Bravo À, Queralt-Rosinach N et al (2017) DisGeNET: a comprehensive platform integrating information on human disease-associated genes and variants. Nucleic Acids Res 45:D833–D839. https://doi.org/10.1093/nar/gkw943
Chandrashekar DS, Karthikeyan SK, Korla PK et al (2022) UALCAN: an update to the integrated cancer data analysis platform. Neoplasia 25:18–27. https://doi.org/10.1016/j.neo.2022.01.001
Apweiler R (2001) The InterPro database, an integrated documentation resource for protein families, domains and functional sites. Nucleic Acids Res 29:37–40. https://doi.org/10.1093/nar/29.1.37
O’Boyle NM, Banck M, James CA et al (2011) Open babel: an open chemical toolbox. J Cheminform 3:33. https://doi.org/10.1186/1758-2946-3-33
Hanwell MD, Curtis DE, Lonie DC et al (2012) Avogadro: an advanced semantic chemical editor, visualization, and analysis platform. J Cheminform 4:17. https://doi.org/10.1186/1758-2946-4-17
Knox C, Wilson M, Klinger CM et al (2024) DrugBank 6.0: the DrugBank knowledgebase for 2024. Nucleic Acids Res 52:D1265–D1275. https://doi.org/10.1093/nar/gkad976
Morris GM, Huey R, Lindstrom W et al (2009) AutoDock4 and AutoDockTools4: automated docking with selective receptor flexibility. J Comput Chem 30:2785–2791. https://doi.org/10.1002/jcc.21256
Debroy R, Ramaiah S (2023) Translational protein RpsE as an alternative target for novel nucleoside analogues to treat MDR Enterobacter cloacae ATCC 13047: network analysis and molecular dynamics study. World J Microbiol Biotechnol 39:187. https://doi.org/10.1007/s11274-023-03634-z
Varghese R, Basu S, Neeravi A et al (2022) Emergence of meropenem resistance among cefotaxime non-susceptible streptococcus pneumoniae: evidence and challenges. Front Microbiol. https://doi.org/10.3389/fmicb.2021.810414
Peela SCM, Basu S, Sharma J et al (2023) Structure elucidation and interaction dynamics of MefA-MsrD efflux proteins in streptococcus pneumoniae : impact on macrolide susceptibility. ACS Omega 8:39454–39467. https://doi.org/10.1021/acsomega.3c05210
Yuan S, Chan HCS, Hu Z (2017) Using PyMOL as a platform for computational drug design. WIREs Comput Mol Sci. https://doi.org/10.1002/wcms.1298
Pawar SS, Rohane SH (2021) Review on discovery studio: an important tool for molecular docking. Asian J Res Chem 14:1–3. https://doi.org/10.5958/0974-4150.2021.00014.6
Naha A, Banerjee S, Debroy R et al (2022) Network metrics, structural dynamics and density functional theory calculations identified a novel ursodeoxycholic acid derivative against therapeutic target Parkin for Parkinson’s disease. Comput Struct Biotechnol J 20:4271–4287. https://doi.org/10.1016/j.csbj.2022.08.017
Miryala SK, Basu S, Naha A et al (2021) Identification of bioactive natural compounds as efficient inhibitors against Mycobacterium tuberculosis protein-targets: a molecular docking and molecular dynamics simulation study. J Mol Liq 341:117340. https://doi.org/10.1016/j.molliq.2021.117340
Joshi T, Sharma P, Joshi T et al (2022) Repurposing of FDA approved drugs against Salmonella enteric serovar Typhi by targeting dihydrofolate reductase: an in silico study. J Biomol Struct Dyn 40:3731–3744. https://doi.org/10.1080/07391102.2020.1850356
Singh AK, Kushwaha PP, Prajapati KS et al (2021) Identification of FDA approved drugs and nucleoside analogues as potential SARS-CoV-2 A1 pp domain inhibitor: an in silico study. Comput Biol Med 130:104185. https://doi.org/10.1016/j.compbiomed.2020.104185
Kabsch W, Sander C (1983) Dictionary of protein secondary structure: pattern recognition of hydrogen-bonded and geometrical features. Biopolymers 22:2577–2637. https://doi.org/10.1002/bip.360221211
Kollman PA, Massova I, Reyes C et al (2000) Calculating structures and free energies of complex molecules: combining molecular mechanics and continuum models. Acc Chem Res 33:889–897. https://doi.org/10.1021/ar000033j
Kumari R, Kumar R, Lynn A (2014) g_mmpbsa —a GROMACS tool for high-throughput MM-PBSA calculations. J Chem Inf Model 54:1951–1962. https://doi.org/10.1021/ci500020m
Mathpal S, Joshi T, Sharma P et al (2024) In silico screening of chalcone derivatives as promising EGFR-TK inhibitors for the clinical treatment of cancer. 3Biotech 14:18. https://doi.org/10.1007/s13205-023-03858-8
Szklarczyk D, Gable AL, Nastou KC et al (2021) The STRING database in 2021: customizable protein-protein networks, and functional characterization of user-uploaded gene/measurement sets. Nucleic Acids Res 49:D605–D612. https://doi.org/10.1093/nar/gkaa1074
Chin C-H, Chen S-H, Wu H-H et al (2014) cytoHubba: identifying hub objects and sub-networks from complex interactome. BMC Syst Biol 8:S11. https://doi.org/10.1186/1752-0509-8-S4-S11
Ashok G, Ramaiah S (2023) FN1 and cancer-associated fibroblasts markers influence immune microenvironment in clear cell renal cell carcinoma. J Gene Med. https://doi.org/10.1002/jgm.3556
Priyamvada P, Ramaiah S (2023) Potential signature therapeutic biomarkers TOP2A, MAD2L1, and CDK1 in colorectal cancer: a systems biomedicine-based approach. Biochem Genet. https://doi.org/10.1007/s10528-023-10544-0
Luo W, Pant G, Bhavnasi YK et al (2017) Pathview Web: user friendly pathway visualization and data integration. Nucleic Acids Res 45:W501–W508. https://doi.org/10.1093/nar/gkx372
Torre LA, Siegel RL, Ward EM, Jemal A (2016) Global cancer incidence and mortality rates and trends—an update. Cancer Epidemiol Biomarkers Prev 25:16–27. https://doi.org/10.1158/1055-9965.EPI-15-0578
Oh SM, Kim YP, Chung KH (2006) Biphasic effects of kaempferol on the estrogenicity in human breast cancer cells. Arch Pharm Res 29:354–362. https://doi.org/10.1007/BF02968584
Amjad E, Sokouti B, Asnaashari S (2022) A systematic review of anti-cancer roles and mechanisms of kaempferol as a natural compound. Cancer Cell Int 22:260. https://doi.org/10.1186/s12935-022-02673-0
Kim S-H, Choi K-C (2013) Anti-cancer effect and underlying mechanism(s) of kaempferol, a phytoestrogen, on the regulation of apoptosis in diverse cancer cell models. Toxicol Res 29:229–234. https://doi.org/10.5487/TR.2013.29.4.229
Li Q, Wei L, Lin S et al (2019) Synergistic effect of kaempferol and 5-fluorouracil on the growth of colorectal cancer cells by regulating the PI3K/Akt signaling pathway. Mol Med Rep. https://doi.org/10.3892/mmr.2019.10296
Zhou Q, Fang G, Pang Y, Wang X (2023) Combination of kaempferol and docetaxel induces autophagy in prostate cancer cells in vitro and in vivo. Int J Mol Sci 24:14519. https://doi.org/10.3390/ijms241914519
Xu X, Li S, Cui X et al (2019) Inhibition of ubiquitin specific protease 1 sensitizes colorectal cancer cells to DNA-damaging chemotherapeutics. Front Oncol. https://doi.org/10.3389/fonc.2019.01406
Kee Y, D’Andrea AD (2010) Expanded roles of the Fanconi anemia pathway in preserving genomic stability. Genes Dev 24:1680–1694. https://doi.org/10.1101/gad.1955310
Meisenberg C, Gilbert DC, Chalmers A et al (2015) Clinical and cellular roles for TDP1 and TOP1 in modulating colorectal cancer response to irinotecan. Mol Cancer Ther 14:575–585. https://doi.org/10.1158/1535-7163.MCT-14-0762
Leung E, Patel J, Hollywood JA et al (2021) Validating TDP1 as an inhibition target for the development of chemosensitizers for camptothecin-based chemotherapy drugs. Oncol Ther 9:541–556. https://doi.org/10.1007/s40487-021-00158-0
Laporte GA, Leguisamo NM, Gloria HC et al (2020) The role of double-strand break repair, translesion synthesis, and interstrand crosslinks in colorectal cancer progression—clinicopathological data and survival. J Surg Oncol 121:906–916. https://doi.org/10.1002/jso.25737
Imaoka H, Toiyama Y, Saigusa S et al (2015) RacGAP1 expression, increasing tumor malignant potential, as a predictive biomarker for lymph node metastasis and poor prognosis in colorectal cancer. Carcinogenesis 36:346–354. https://doi.org/10.1093/carcin/bgu327
Duan B, Bai J, Qiu J et al (2018) Histone-lysine N-methyltransferase SETD7 is a potential serum biomarker for colorectal cancer patients. EBioMedicine 37:134–143. https://doi.org/10.1016/j.ebiom.2018.10.036
Duan B, Bai J, Qiu J et al (2023) Corrigendum to “Histone-lysine N-methyltransferase SETD7 is a potential serum biomarker for colorectal cancer patients” [EBioMedicine 37 (2018) 134–143]. EBioMedicine 91:104580. https://doi.org/10.1016/j.ebiom.2023.104580
Monteiro FL, Williams C, Helguero LA (2022) A systematic review to define the multi-faceted role of lysine methyltransferase SETD7 in cancer. Cancers (Basel) 14:1414. https://doi.org/10.3390/cancers14061414
Oudhoff MJ, Braam MJS, Freeman SA et al (2016) SETD7 controls intestinal regeneration and tumorigenesis by regulating Wnt/β-Catenin and Hippo/YAP signaling. Dev Cell 37:47–57. https://doi.org/10.1016/j.devcel.2016.03.002
Farooqi AA, de la Roche M, Djamgoz MBA, Siddik ZH (2019) Overview of the oncogenic signaling pathways in colorectal cancer: mechanistic insights. Semin Cancer Biol 58:65–79. https://doi.org/10.1016/j.semcancer.2019.01.001
Kashyap D, Sharma A, Tuli HS et al (2017) Kaempferol—a dietary anticancer molecule with multiple mechanisms of action: recent trends and advancements. J Funct Foods 30:203–219. https://doi.org/10.1016/j.jff.2017.01.022
Laissue P (2019) The forkhead-box family of transcription factors: key molecular players in colorectal cancer pathogenesis. Mol Cancer 18:5. https://doi.org/10.1186/s12943-019-0938-x
Tenbaum SP, Ordóñez-Morán P, Puig I et al (2012) β-catenin confers resistance to PI3K and AKT inhibitors and subverts FOXO3a to promote metastasis in colon cancer. Nat Med 18:892–901. https://doi.org/10.1038/nm.2772
Luo H, Hao E, Tan D et al (2019) Apoptosis effect of Aegiceras corniculatum on human colorectal cancer via activation of FoxO signaling pathway. Food Chem Toxicol 134:110861. https://doi.org/10.1016/j.fct.2019.110861
Shi F, Li T, Liu Z et al (2018) FOXO1: another avenue for treating digestive malignancy? Semin Cancer Biol 50:124–131. https://doi.org/10.1016/j.semcancer.2017.09.009
Kashafi E, Moradzadeh M, Mohamadkhani A, Erfanian S (2017) Kaempferol increases apoptosis in human cervical cancer HeLa cells via PI3K/AKT and telomerase pathways. Biomed Pharmacother 89:573–577. https://doi.org/10.1016/j.biopha.2017.02.061
Danielsen SA, Eide PW, Nesbakken A et al (2015) Portrait of the PI3K/AKT pathway in colorectal cancer. Biochim Biophys Acta Rev Cancer 1855:104–121. https://doi.org/10.1016/j.bbcan.2014.09.008
Choi J-B, Kim J-H, Lee H et al (2018) Reactive oxygen species and p53 mediated activation of p38 and caspases is critically involved in kaempferol induced apoptosis in colorectal cancer cells. J Agric Food Chem 66:9960–9967. https://doi.org/10.1021/acs.jafc.8b02656
Funding
The authors gratefully acknowledge the Indian Council of Medical Research (ICMR), New Delhi, Government of India agency, for the research grant (IRIS ID: 2021–11889).
Author information
Authors and Affiliations
Contributions
Conceptualization and methodology-P. P and G.A.; Validation and formal analysis- P.P, G.A and T.J; Investigation- A.A and S.R; writing-original draft preparation- P.P and G.A; Writing- review and editing- A.A and S.R; visualization- P.P., G.A. and S.A.; supervision- A.A and S.R.
Corresponding author
Ethics declarations
Conflict of interest
The authors declare that they have no conflicts of interest.
Ethical approval
Not applicable.
Consent to publish
Not applicable.
Consent to participate
Not applicable.
Additional information
Publisher's Note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary Information
Below is the link to the electronic supplementary material.
Rights and permissions
Springer Nature or its licensor (e.g. a society or other partner) holds exclusive rights to this article under a publishing agreement with the author(s) or other rightsholder(s); author self-archiving of the accepted manuscript version of this article is solely governed by the terms of such publishing agreement and applicable law.
About this article
Cite this article
Priyamvada, P., Ashok, G., Joshi, T. et al. Unravelling the molecular mechanistic pathway underlying the anticancer effects of kaempferol in colorectal cancer: a reverse pharmacology network approach. Mol Divers (2024). https://doi.org/10.1007/s11030-024-10890-0
Received:
Accepted:
Published:
DOI: https://doi.org/10.1007/s11030-024-10890-0