Abstract
Kingella negevensis belongs to the Neisseriaceae family. It is implied that it has significant virulence potential due to RTX toxin production, which can cause hemolysis. It usually colonizes the orophayrynx of pediatric population, along with Kingella kingae but has also been isolated from vagina. Todate no report on its drug targets is present, therefore putative therapeutic targets were identified from its genomic sequence data. Traditional Chinese (n > 36,000) and Indian medicinal compounds (n > 2000) were then screened against its pyridoxine 5'-phosphate synthase, a vital therapeutic target. Prioritized TCM compounds included ZINC02525131, ZINC33833737 and ZINC85486932, and Cadiyenol, 9,11,13-Octadecatrienoic acid and 6-Gingerol from Indian medicinal library. Molecular dynamics simulation of top compounds revealed ZINC02525131 as having best stability for 100 ns, compared to Cadiyenol. ADMET profiling was then done, along with physiologically based pharmacokinetic simulation of these compounds in a population of 200 individuals, for 12 h to see fate of the ingested compound. Additionally, the impact of these compounds in a population with cirrhosis and renal impairment was also simulated. We imply in light of all the studied parameters of safety and bioavailability, etc., that 6-Gingerol from Zingiber officinalis rhizome must be proceeded further for in vitro and in vivo testing for inhibition of K. negevensis.
Similar content being viewed by others
Introduction
Kingella negevensis is a gram-negative bacterium and produces RTX toxin associated hemolysis [1]. Its colonization is known in respiratory tract and oropharynx, with carriage and colonization resembling K. kingae [2, 3]. Its occurrence is known to increase from 6 to 24 month children and decrease thereafter [4]. Its role in the septic arthritis of an infant [2, 4, 5], endocarditis, pediatric osteomyelitis and bacteremia has been implicated [1], while the bacterial spread is through person to person contact [4].
Its genome has been sequenced, with genome size around 2 MB [6]. This bacterium has been reported to show heterogeneity in genetic makeup in different strains [4]. Among several key virulence factors, human epithelial binding through elements such as an exopolysaccharide, a polysaccharide capsule, an adhesin autotransporter, and a pili (type IV) are shared between K. negevensis and K. kingae [7]. Its integrative and conjugative elements (ICE) also resemble the Neisseria gonorrhea [8], with high homology between type IV secretion system protein virB4, involved in human endothelial cell subversion. Apart from this, type IV coupling protein T4CP, DNA transesterase enzyme relaxase and integrase are also exceptionally conserved. DNA uptake sequences with ICE sequences, match that of Neisseria gonorrhea and Neisseria meningitides [8].
Genome sequence availability is a boon for bioinformatics based studies, where algorithms and softwares can be used to mine therapeutic targets and explore druggable potential of a bacteria [9]. This approach has previously been implemented for identifying therapeutic targets in drug resistant Salmonella typhi [10], Enterobacteriacea family [11], Bacillus sp. [12], Chlamydia pneumoniae [13] etc. Main principle is application of the ‘essentiality & selectivity criteria’, where a gene product presence should be necessary for the bacterial survival and lacking in host. This guarantees that drug molecules targeted against the pathogen will not disrupt the host system [14]. The capability to discern molecules having a robust modulatory activity against a pathogen, using softwares (via lock and key fit contrivance) is a blessing for drug discovery. A large number of molecules with selectivity for an enzyme target can be screened in a small time. Molecular docking estimates the best binding through a regression or classification based scoring and thus, prioritizes hits in a library [15]. However, the hits should be subjected to several filters as decoys, toxic, less bioavailable and non-active molecules are of little use in real scenario. For this purpose, the absorption, distribution, metabolism, and excretion (ADME) properties, along with physiologically based pharmacokinetic (PBPK) profiling helps confirm drug efficacy and tolerability [16]. In the past, poor PK properties (e.g., small bioavailability) have led to the failure of a large fraction of lead compounds [17]. This is why, good PK properties can be of motivation to further explore molecule as a drug. It can also shed light on dosing [18] and explicit adaptations of the regimen in different ethnicities [19], as well as health conditions.
Natural products have gained a lot of importance in drug design against pathogens and around half of the FDA approved drugs (USFDA, 1981–2019) are sourced or based on natural compounds [20]. Their large chemical space [21] as well as already established medicinal properties as traditional remedy against ailments makes them an important treasure trove for screening against pathogenic bacteria. Traditional Indian (Ayurvedic) and Traditional Chinese Medicine (TCM), have a deep-rooted history in the pharmacopeia of respective areas. Drugs derived from these medicinal systems have been implicated in diseases like cancer [22,23,24,25,26,27,28], COVID-19 [28, 29]. Although traditional medicine comprises single or multi-component preparation, computational screening has lent a quick evaluation strategy for single compound assessment. Therapeutic efficacy of compounds screened from these sources is not disease but rather structure based.
In this study, we inferred therapeutic targets and carried out virtual screening of two natural product libraries of Indian and Chinese origin, against pdxJ gene product of K. nevegensis. ADMET and PBPK properties were also studied for prioritized compounds of our selected natural product libraries. To the best of authors knowledge, this is the first report of therapeutic target map of K. nevegensis and screening of natural product inhibitors against it.
Material and methods
Data acquisition
Genome data of the Kingella negevensis strain Sch538, with accession CCNJ00000000, was obtained from NCBI. Coding DNA sequences and proteome was also procured.
Subtractive genomics
The data were subjected to subtractive genomics for therapeutic target identification using previously described pipeline [30]. Core i7 (7th generation) machine was used for analysis, with 8 GB RAM. Paralogous sequences with more than 60% similarity were removed and genomic data were converted to protein dataset. Data for DEG [31] and CEG [32]database, human proteome, gut microbiota (n = 84 bacteria), were obtained from UniProt (https://www.uniprot.org), NCBI (https://www.ncbi.nlm.nih.gov), DEG (http://www.essentialgene.org) and CEG website (http://cefg.uestc.cn/ceg), respectively. Homologous or non-homologous sets of proteins were filtered in the same order, against these datasets using standalone BLAST 2.2.31. One protein pyridoxine 5'-phosphate (PNP) synthase (Accession no: WP_032137481.1), involved in vitamin B6 synthesis, was chosen from obtained therapeutic targets for downstream analysis.
Structure modeling and virtual screening
PNP synthase was subjected to 3D structural modeling using I-Tasser [33], with LOMETS multi-threading algorithm at the back-end. Since LOMETS picks lots of templates from the PDB library, I-TASSER sifts out significant ones and retains top ten based on Z-score. Top template was a crystal structure of PNP synthase from Escherichia coli, with bound 1-deoxy-d-xylulose phosphate (PDB ID: 1m5w), while ten structures in all were used for threading based structure modeling. Alignment of these structures were obtained using several softwares i.e. SPARKS-X, HHSEARCH, FFAS-3D, Neff-PPAS, pGenTHREADER, wdPPAS, PROSPECT2 and SP3. Generated decoys were clustered and top one picked by SPICKER program. C-score was used for final ranking of structures in top cluster. Pro-motif was used to study secondary structure while co-factor and COACH were used to predict ligand binding site residues. Ramachandran analysis was done for structure validation (https://zlab.umassmed.edu/bu/rama/index.pl).
Docking based screening was carried out using Molecular Operating Environment (MOE) software version 2019.1. TCM consisted of 36,000 compounds while Ayurvedic library consisted of 2002 molecules. Triangle placement method was used for initial round of hit prioritization and forcefield based refinement was carried out for further improvement of hits screening. Three complexes with least energy values were saved from each library and ligand interaction diagrams were also drawn for 2D visualization. Top scoring compounds were subjected to dynamics simulation of 100 ns using Desmond, according to previously described parameters [34].
ADMET and PBPK evaluation
These compounds were subjected to ADMET using machine learning based pkCSM (https://biosig.lab.uq.edu.au/pkcsm/) platform. It is centered on a graph-based technique, where distance between atoms is used for training predictive regression and classification models, that calculate ADMET properties. Absorption variables include water solubility, intestinal absorption, Caco-2 and skin permeability, whereas distribution lists blood brain barrier and central nervous system permeability. Metabolism of cytochrome enzymes is available, alongside clearance as excretion parameter. Toxicity is determined by quantity tolerated in rat, minnow, T. pyriformis, hepatoxicity, Ames toxicity and skin sensitization [35].
PBPK modeling is based on a chain of differential equations and has been executed in GastroPlus software (SimulationsPlus LLC). Compound-specific parameters like weight, pKa values are input and plasma concentration is profiled after chosen route of administration. Statistically, oral route has been the most efficacious [36], so it was chosen. GastroPlus includes distribution factors for Pgp, PepT1, HPT1, OCTN1, LAT2 and OATP1A2 transporters. Vmax is adjusted for each compartment, based on the values from these transporters. Health conditions were taken as (a) normal, (b) cirrhosis, (c) renal impairment. Quantity was taken as 100 mg, with intake in fasting state in the humans (population of 200). pH-dependent dissolution model was selected, with pH = 7.2. Diffusion coefficient values were taken as 0.5–1.5 × 10–5 cm2/sec, transit time value for the stomach = 0.25 h, transit time value for caecum = 4.5 h, transit time value for colon = 13.5 h and drug particle density = 1.2 g/mL. Lengths of compartments, radii, transit times and pH values were adjusted in population, with respect to weights of individuals. Simulation time was 12 h and Advanced Compartmental and Transit (ACAT) Model was implemented.
Results and discussion
Therapeutic candidate mapping
After paralog removal, 2037 hits were obtained from the total 2104 CDSs. DEG similar sequences were 944 and CEG similar sequences were 814. Common sequences to both DEG and CEG proteome were only 798. These were compared to the human proteome and 386 non-similar proteins were obtained, with 157 among these dissimilar to the human gut microbiota proteome. These were compared to the DrugBank and 41 druggable proteins were obtained (Supplementary Table 1).
Among the druggable proteins, PNP synthase was selected for further analysis as it is an essential part of pathways involved in amino acid synthesis in prokaryotes but absent in humans [37]. This protein is a product of pdxJ gene and the active form of vitamin B6 contrived by this enzyme serves as a co-enzyme for metabolism of several amino acid, lipid and glucose pathways. This enzyme is vital to many processes in the bacterium, including deamination, transamination, decarboxylation, and racemization. [26]. This enzyme family exists as a small but necessary fraction of prokaryotic genome (~ 1.5% of genome) [38]. Nearly 4% of reactions propelled by enzymes are linked with this family of enzymes, as catalogued in the Enzyme Commission database, but approved drugs against this class of enzymes is scanty. We aimed to explore this enzyme as a drug target against K. negevensis and screen traditional medicinal compounds against it. For this, first of all structure was modeled as no experimental model was present for this specie in the Protein databank.
3D structure modeling of PNP synthase
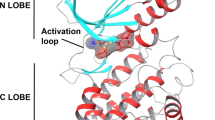
Structure of the PNP synthase (EC 2.6.99.2) was threaded using 10 templates by I-TASSER. Overall ERRAT quality factor was 97%. Top templates with identity of more than 50% with the aligned region as well as whole protein sequence were PNP synthase from Escherichia coli (PDB ID: 1M5W), pyridoxal phosphate biosynthetic protein from Burkholderia pseudomallei (PDB ID: 3GK0), and PNP synthase from Pseudomonas aeruginosa (PDB ID: 5DLC). The obtained model was composed of 1 sheet, 6 beta-alpha–beta motif units (with some residues in loops and some in helices), 1 parallel wide type beta bulge, 8 parallel strands with topology 7X -1X -1X -1X -1X -1X -1X, 13 helices, 17 helix-helix interacs, 8 beta turns, 2 gamma turns (Fig. 1A). Non-glycine and non-proline residues were 218 while glycine and proline residues were 23. Normally, this enzyme depicts TIM-barrel or alpha/beta construction with eight helices and parallel beta strands. However, inner core is hydrophilic and three additional helices are present [37].
A 3D structure of the modeled PNP synthase of K. negevensis, with active site residues indicated by dotted spheres and central gorge visible as a tunnel in the surface representation of the ribbon structure. B Ramachandran plot showing particularly good observations (95.87%) in GREEN Crosses, followed by slightly less favored in brown triangles (2.2%) and questionable ones in red circles (1.8%). Black and gray regions are for preferred conformations, with delta values ≥ 2
Five binding sites were identified by COACH and co-factor. The crystal structure of PNP synthase from template of Escherichia coli consisted of eight binding sites, and predicted active site residues by I-TASSER based on chosen templates consisted of Asn6, His9, Thr12, His42, Arg44, Glu69, Val91, Glu93, Gly102, Phe130, His152, Gly191, Thr193, Gly212, Ile216 residues. Binding site residue prioritization based on all hits via I-TASSER predicted highest bonding capability of Asn6, Glu150, Gly189, His190, Asn210, Ile211, Gly212, His213. Ramachandran plot showed 95% residues in the most favored and 5% in additional allowed region. Only 1.8% (four residues) were in the disallowed region (Fig. 1B).
Docking based screening
Structure modeling was followed by docking with TCM and Ayurvedic medicinal compounds. ZINC02525131, ZINC33833737, ZINC85486932 were mined as top inhibitors from TCM, while Cadiyenol, 9,11,13-Octadecatrienoic acid and 6-Gingerol was prioritized from Ayurvedic library. Prioritized compounds (Table 1) showed bonding with majority of identified active site residues in all of the complexes (Fig. 2). The S-values and MM/PBSA values were in alliance with each other, except for the 9,11,13-Octadecatrienoic acid complex with PNP synthase, as it showed slightly lower value compared to Cadiyenol, which had a lower S-value but slightly higher MM/PBSA value.
2D representation of the docked A PNP synthase and ZINC02525131 complex. B PNP synthase and ZINC33833737 complex C PNP synthase and ZINC85486932 complex D PNP synthase and Cadiyenol complex E PNP synthase and 9,11,13-Octadecatrienoic acid complex F PNP synthase and 6-Gingerol complex. 3D depiction of these representations is shown in the supplementary Fig. 1
Information for every compound was not available in literature but ZINC02525131/β-Hydroxyisovalerylshikonin, the top hit from TCM library, has previously been isolated from Lithospermum radix and known to impart chemotherapeutic properties, conferring apoptotic cell death in human lung cancer DMS114 cells [39] and adenocarcinomic human alveolar basal epithelial A549 cells [39]. This compound has also been isolated from Lithospermum erythrorhizon [40]. It has also been reported that it has fungal properties and reduces mycelium formation in C. albicans [39]. Cadiyenol has been isolated from Centella asiatica (also known as Indian pennywort) [41, 42], and shown apoptotic activity in the murine lymphoma cells. 9,11,13-Octadecatrienoic acid, also called Punicic acid or ɑ-eleostearic acid, is a linoleic acid derivative and occurs in Momordica cochinchinensis and known to inhibit estrogen negative and positive breast cancer proliferation [43]. It also occurs in Pleurocybella porrigens [44], Momordica charantia [45], M. cymbalaria [46] Punica granatum [47] and Aleurites montana [48]. Its antibacterial properties have been reported [49, 50]. 6-Gingerol is present in the rhizome of ginger (Zingiber officinalis) and the physicochemical properties of this tuber have made it a timeless traditional medicinal plant against several ailments [51]. Gingerol is a phenolic constituent and previously, 6-Gingerol has been attributed to reduce allergic rhinitis by suppression of T-cells [52]. Extract with high content of 6-Gingerol has acted as an antioxidant and anti-inflammatory agent in murine models subjected to organophosphate pesticide chlorpyrifos, that causes oxidative damage [53]. It has also shown anti-proliferative effect in prostate cancer cells [54]. Here, we have predicted anti-bacterial property of all the compounds mentioned in Table 1.
Dynamics simulation
Dynamics simulation of two top complexes, PNP synthase with ZINC02525131 and Cadiyenol was conducted. The average RMSD for ZINC02525131 did not exceed 3 Å throughout the whole simulation, depicting the complex as stable (Fig. 3A). Protein ligand interaction (Fig. 3B) showed that Ser214 and Gly191played a significant role in ligand binding through hydrogen bonding, with a robust impact on metabolization, specificity and adsorption of ZINC02525131. Compared to these residues, Asn6, Asp8, His42, Arg44, Thr193, Gly212, His213 made small or transitory contact with ligand through hydrogen bonds. Hydrogen-bonded interactions arbitrated by a water molecule, also known as water bridges were made by Asn6, Asp8, His9, His42, Arg44, His190, Gly191, Thr193, His213. Transitory ionic interactions mediated by the protein backbone were displayed by just two residues, His42 and Glu69, while hydrophobic interaction by just Ile211. Arg44, His213, Ser214 and Gly191 retained interactions for more than 30% of the simulation time.
Interaction plot with Cadiyenol (Fig. 4A) showed some fluctuations in the beginning of the simulation, high disparity from 20 to 30 ns and later 60–80 ns. From 90 to 100 ns, the ligand seemed to move away from the protein but the overall rmsd did not exceed 3 Å on the average. It was tightly bound with PNP synthase from 20 to 30 ns, 60–70 ns and 80–90 ns. However, the pattern is not uniform so this interaction is more unstable than PNP synthase with ZINC02525131. This shows the discrepancy in the binding score value versus dynamics simulation plot, as the binding score of Cadiyenol was higher than ZINC02525131, but binding stability was inferred as less. Glu69, Phe130, and His190 retained interactions for more than 30% of the simulation time. Gly69 and His190 was making water bridge (Fig. 4B) and hydrogen bond interaction while Phe130 depicted a hydrophobic interaction.
ADMET and pharmacokinetics
Good intestinal absorption was seen for the studied compounds, except for ZINC85486932. Cadiyenol, 6-Gingerol and ZINC33833737 had high caco-2 permeability, meaning high absorption of orally consumed drugs. Cadiyenol and 9,11,13-Octadecatrienoic acid were substrates and inhibitors of P-glycoprotein, means that they can bind but inhibit their transport outside of the cell. ZINC85486932 seems to have a high tendency of being purged out of the cell as it did not show inhibition of any of the P-glycoprotein. ZINC02525131 had least tendency to cross skin, but overall almost all other compounds had low skin permeability as well (values less than − 2.5). Steady state volume of distribution (VDss) is calculated to estimate the amount of dose required for uniform distribution of the drug in similar quantity in the plasma. The values were not too high (log VDss > 0.45) or too low (log VDss < − 0.15). Compounds did not show high blood brain barrier (BBB) permeability (none showed logBB > 0.3) but most poor BBB permeability was seen for ZINC85486932 (logBB < − 1). ZINC33833737 showed some possibility of central nervous system penetration while ZINC85486932 depicted no capability to cross central nervous system barrier. Compounds bind to cytochrome (CYP) 450 enzymes for detoxification, excretion or activation. While some compounds were substrates of CYP3A4, none inhibited CYP1A2, CYP2C19, CYP2C9 and CYP2D6. Only 6-Gingerol inhibited CYP3A4. Traditional Indian compounds had better clearance compared to TCM compounds, while no compound showed AMES toxicity. Highest tolerated doses were for ZINC02525131 and 6-Gingerol. ZINC02525131 depicted hepatotoxicity and 9,11,13-Octadecatrienoic acid showed skin sensitization but none of the compounds was an inhibitor of calcium channel hERG I/II. Inhibition of these genes causes long QT syndrome, further leading to ventricular arrhythmia and thus, should be stopped from further processing. In the past, many drugs have been withdrawn after showing this inhibition property.
Gastrointestinal tract absorption and kinetics of the compound were also simulated. ZINC02525131 and 6-Gingerol had maximum bioavailability in healthy state, while Cadiyenol bioavailability improved from ~ 77 to 99% in liver and renal impairment. Absorption of all compounds remained equal to or increased as compared to bioavailability in impaired state. This may be because the transit and excretion are considered as a continuous process in PBPK modeling and rates of these processes deliberated as reciprocal values of individual compartment’s transit time. This means that even after completion of transit time through stomach, intestine and colon, a substantial quantity of drug might still be absorbed in the gut. During enterohepatic recirculation, the absorbed percentage may be more as some of the dose is reabsorbed after secretion in the bile. Impaired state may have an impact on this parameter and thus, values are higher compared to the healthy state. Least time was required for ZINC02525131 to reach highest plasma concentration, while highest for the 9,11,13-Octadecatrienoic acid in healthy state. In impaired health state, time was in slight alliance with healthy state, except for ZINC33833737. It showed a large disparity in time to reach maximum concentration in plasma (increased from approximately 4–5 h to 10 h & 8–9 h in liver and renal impairment, respectively. ZINC33833737 also had the highest AUC till 12 h of simulation in healthy state, while 9,11,13-Octadecatrienoic acid had least AUC in liver impairment. ZINC33833737 concentration quantity was highest among all the compounds in healthy state, while ZINC85486932 showed highest concentration in impaired state.
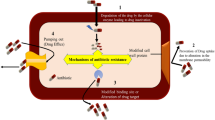
K. negevensis has been isolated from the oral cavity of children [4, 55] and vagina in vaginosis [1]. It is known to produce RTX toxin and thus, designated a pathogen [1]. The mode of action of this class of toxins (part of type I secretion system and acting as a virulence factor) is hemolysis/cytotoxicity through membrane perforation of the host cell [56]. Todate, no report of therapeutic targets exists in literature for K. negevensis. For this reason, a comprehensive subtractive genomics strategy was utilized to infer the candidates that quality as druggable. More than 40 such proteins were inferred (Supplementary Table 1). Among these, one (PNP synthase) was selected for further analysis based on its importance, functional role in cell and novelty. PNP synthase is a homooctameric enzyme, that carries out catalysis of the last step of B6 vitamer biosynthesis, via condensation of deoxyxylulose-5-phosphate and aminoacetone-3-phosphate. This synthesis is essential to many pathways (like amino acid metabolism and antibiotic production) [57, 58]. Additionally, owing to the exclusive occurrence of PNP synthase in some bacteria and not in humans, as well as the vitality for bacterial survival, this enzyme is an encouraging target for screening antibacterial compounds. The structure of this enzyme in K. negevensis is not known yet and necessary for docking, so it was modeled using bioinformatics approach of threading. Full-length PNP synthase model was constructed using iterative procedure, with similar structural fragments cut out from template protein structures and simulated for our sequence. Secondary structure and B-factor values complemented the 3D coordinate information, indicating that the residues with helix or sheet architecture and flexible or rigid. Helices comprised major portion of the protein, followed by coils and strands. The residues were flexible mostly in coil regions, with most flexibility at C and N terminal regions. Active site residues were predicted and used for screening natural product inhibitors (see Table 2).
Virtual screening of natural products is a swift strategy which involves interaction modeling of drug and protein, with favored pose having least energy and showing stable configuration. Therefore, this strategy was adapted and natural products of traditional Indian and Chinese medicine origin, having a large structural as well as physicochemical variety were screened. Docking revealed the binding conformations for the PNP synthase and natural product compounds. Usage of natural products derived from Ayurveda and traditional Chinese medicinal plants dates back to old times and is still utilized in some places of the world [59]. Their revitalization has occurred in cheminformatics based drug mining literature, with new studies looking for natural product based inhibitors against pathogens [30, 60,61,62]. Structure docking was done on hits from TCM and Ayurvedic compounds against PNP synthase to identify potent inhibitors. Previously Ahmad et al. have reported a compound 2-acetyl-3-(2-heptanamidoethyl)-1H-indol-6-yl heptanoate inhibitor of this enzyme through computational screening, with high affinity in Yersinia enterocolitica [63]. We prioritized six compounds from traditional medicinal plants/herbs, based on scoring functions of software’s where ADMET profiling revealed 6-Gingerol as most readily bioavailable and safe. Its medicinal properties have been demonstrated previously as well [52, 53]. Some scientists date the usage of Zingiber officinale to more than 2000 years ago [64] as food condiment. Its use has been implied in both Indian and Chinese medicine [65]. This tuber is “generally recognized as safe” by the Food and Drug Administration, so the constituents of this condiment are nontoxic for consumption [66]. Its antibacterial properties have been demonstrated previously and is an encouraging substitute of synthetic antimicrobials [67,68,69]. We suggest that the 6-Gingerol be tested further in lab on cell lines and in mouse models, for targeted antibacterial action against K. negevensis (see Table 3).
Conclusion
K. negevensis possesses a small sized genome (~ 2 MB) but hosts several virulence factors and causes diseases in children. A case of vaginosis in adult patient has also been reported. The bacterium is currently understudied and similarity as well as co-occurrence with K. kingae in many cases makes it additionally difficult to separately study and link causation with symptoms of the resultant disease. This is why few data is available in literature regarding this bacterium. Only three genome sequences are present in the public NCBI database. In this study, therapeutic targets were mined from the bacterium isolated from pharynx of a child and using biophysical approach, structure modeling and virtual screening of a key drug target PNP synthase was done. We propose on the basis of PBPK and ADMET analysis that 6-Gingerol should be pursued further as an antibacterial compound against K. negevensis.
Data availability
No new data were generated in this work that requires submission in a repository. Details of the data used in this research are included within the manuscript.
References
Opota O, Laurent S, Pillonel T, Leger M, Trachsel S, Prod’hom G, Jaton K, Greub G (2017) Genomics of the new species Kingella negevensis: diagnostic issues and identification of a locus encoding a RTX toxin. Microbes Infect 19:546–552. https://doi.org/10.1016/j.micinf.2017.08.001
Yagupsky P (2022) Pharyngeal colonization by Kingella kingae, transmission, and pathogenesis of invasive infections: a narrative review. Microorganisms. https://doi.org/10.3390/microorganisms10030637
Yagupsky P, El Houmami N, Fournier PE (2018) Respiratory carriage of the novel Kingella negevensis species by young children. New Microbes New Infect 26:59–62. https://doi.org/10.1016/j.nmni.2018.08.011
Yagupsky P (2018) Detection of respiratory colonization by Kingellakingae and the novel Kingella negevensis species in children: uses and methodology. J Clin Microbiol. https://doi.org/10.1128/JCM.00633-18
Yagupsky P (2017) Diagnosing Kingella kingae infections in infants and young children. Exp Rev Anti Infect Ther 15:925–934. https://doi.org/10.1080/14787210.2017.1381557
El Houmami N, Schrenzel J, Yagupsky P, Robert C, Ceroni D, Raoult D, Fournier PE (2017) Draft genome sequence of Kingella negevensis SW7208426, the first European strain of K.negevensis Isolated from a healthy child in Switzerland. Genome Announc. https://doi.org/10.1128/genomeA.00571-17
Porsch EA, Yagupsky P, St Geme JW (2020) Kingella negevensis shares multiple putative virulence factors with Kingella kingae. PLoS One 15:e0241511. https://doi.org/10.1371/journal.pone.0241511
Hughes-Games A, Roberts AP, Davis SA, Hill DJ (2020) Identification of integrative and conjugative elements in pathogenic and commensal Neisseriaceae species via genomic distributions of DNA uptake sequence dialects. Microb Genom. https://doi.org/10.1099/mgen.0.000372
Basharat Z, Akhtar U, Khan K, Alotaibi G, Jalal K, Abbas MN, Hayat A, Ahmad D, Hassan SS (2022) Differential analysis of Orientia tsutsugamushi genomes for therapeutic target identification and possible intervention through natural product inhibitor screening. Comput Biol Med 141:105165. https://doi.org/10.1016/j.compbiomed.2021.105165
Jalal K, Khan K, Hassam M, Abbas MN, Uddin R, Khusro A, Sahibzada MUK, Gajdacs M (2021) Identification of a novel therapeutic target against XDR Salmonella typhi H58 using genomics driven approach followed up by natural products virtual screening. Microorganisms. https://doi.org/10.3390/microorganisms9122512
Hadizadeh M, Tabatabaiepour SN, Tabatabaiepour SZ, Hosseini Nave H, Mohammadi M, Sohrabi SM (2018) Genome-wide identification of potential drug target in Enterobacteriaceae family: a homology-based method. Microb Drug Resist 24:8–17. https://doi.org/10.1089/mdr.2016.0259
AnisAhamed N, Panneerselvam A, Arif IA, Syed Abuthakir MH, Jeyam M, Ambikapathy V, Mostafa AA (2021) Identification of potential drug targets in human pathogen Bacillus cereus and insight for finding inhibitor through subtractive proteome and molecular docking studies. J Infect Public Health 14:160–168. https://doi.org/10.1016/j.jiph.2020.12.005
Kadi RH, Altammar KA, Hassan MM, Shater AF, Saleh FM, Gattan H, Al-Ahmadi BM, AlGabbani Q, Mohammedsaleh ZM (2022) Potential therapeutic candidates against Chlamydia pneumonia discovered and developed in silico using core proteomics and molecular docking and simulation-based approaches. Int J Environ Res Public Health. https://doi.org/10.3390/ijerph19127306
Gupta R, Pradhanb D, Kumar A, Rai CS (2017) TiD: Standalone software for mining putative drug targets from bacterial proteome. Genomics 109:51–57
Ghislat G, Rahman T, Ballester PJ (2021) Recent progress on the prospective application of machine learning to structure-based virtual screening. Curr Opin Chem Biol 65:28–34. https://doi.org/10.1016/j.cbpa.2021.04.009
Ferreira LT, Borba JVB, Moreira-Filho JT, Rimoldi A, Andrade CH, Costa FTM (2021) QSAR-based virtual screening of natural products database for identification of potent antimalarial hits. Biomolecules. https://doi.org/10.3390/biom11030459
Ferreira A, Lapa R, Vale N (2021) PBPK modeling and simulation and therapeutic drug monitoring: possible ways for antibiotic dose adjustment. Processes 9:2087
Li S, Yu Y, Bian X, Yao L, Li M, Lou YR, Yuan J, Lin HS, Liu L, Han B, Xiang X (2021) Prediction of oral hepatotoxic dose of natural products derived from traditional Chinese medicines based on SVM classifier and PBPK modeling. Arch Toxicol 95:1683–1701. https://doi.org/10.1007/s00204-021-03023-1
Adiwidjaja J, Gross AS, Boddy AV, McLachlan AJ (2022) Physiologically-based pharmacokinetic model predictions of inter-ethnic differences in imatinib pharmacokinetics and dosing regimens. Br J Clin Pharmacol 88:1735–1750. https://doi.org/10.1111/bcp.15084
Newman DJ, Cragg GM (2020) Natural products as sources of new drugs over the nearly four decades from 01/1981 to 09/2019. J Nat Prod 83:770–803. https://doi.org/10.1021/acs.jnatprod.9b01285
Santana K, do Nascimento LD, Lima ELA, Damasceno V, Nahum C, Braga RC, Lameira J (2021) Applications of virtual screening in bioprospecting: facts, shifts, and perspectives to explore the chemo-structural diversity of natural products. Front Chem 9:662688. https://doi.org/10.3389/fchem.2021.662688
Fatima N, Baqri SSR, Alsulimani A, Fagoonee S, Slama P, Kesari KK, Roychoudhury S, Haque S (2021) Phytochemicals from Indian ethnomedicines: promising prospects for the management of oxidative stress and cancer. Antioxidants (Basel). https://doi.org/10.3390/antiox10101606
Kumar G, Madka V, Pathuri G, Ganta V, Rao CV (2022) Molecular mechanisms of cancer prevention by gooseberry (Phyllanthus emblica). Nutr Cancer 74:2291–2302. https://doi.org/10.1080/01635581.2021.2008988
Li Z, Feiyue Z, Gaofeng L (2021) Traditional Chinese medicine and lung cancer–from theory to practice. Biomed Pharmacother 137:111381. https://doi.org/10.1016/j.biopha.2021.111381
Sarada K, Puthiyedath R, Philip A, Ravindran GC, Pavithran K (2021) Prevalence of the use of traditional complementary and alternative medicine amongst cancer patients in a tertiary care center in Kerala, India. J Ayurveda Integr Med 12:359–364. https://doi.org/10.1016/j.jaim.2021.04.011
Sun Q, He M, Zhang M, Zeng S, Chen L, Zhao H, Yang H, Liu M, Ren S, Xu H (2021) Traditional Chinese medicine and colorectal cancer: implications for drug discovery. Front Pharmacol 12:685002. https://doi.org/10.3389/fphar.2021.685002
Wang S, Fu JL, Hao HF, Jiao YN, Li PP, Han SY (2021) Metabolic reprogramming by traditional Chinese medicine and its role in effective cancer therapy. Pharmacol Res 170:105728. https://doi.org/10.1016/j.phrs.2021.105728
Yao CL, Zhang JQ, Li JY, Wei WL, Wu SF, Guo DA (2021) Traditional Chinese medicine (TCM) as a source of new anticancer drugs. Nat Prod Rep 38:1618–1633. https://doi.org/10.1039/d0np00057d
Adithya J, Nair B, Aishwarya TS, Nath LR (2021) The plausible role of indian traditional medicine in combating corona virus (SARS-CoV 2): a mini-review. Curr Pharm Biotechnol 22:906–919. https://doi.org/10.2174/1389201021666200807111359
Basharat Z, Khan K, Jalal K, Ahmad D, Hayat A, Alotaibi G, Al Mouslem A, Aba Alkhayl FF, Almatroudi A (2022) An in silico hierarchal approach for drug candidate mining and validation of natural product inhibitors against pyrimidine biosynthesis enzyme in the antibiotic-resistant Shigella flexneri. Infect Genet Evol 98:105233. https://doi.org/10.1016/j.meegid.2022.105233
Zhang R, Ou HY, Zhang CT (2004) DEG: a database of essential genes. Nucleic Acids Res 32:D271-272. https://doi.org/10.1093/nar/gkh024
Liu S, Wang SX, Liu W, Wang C, Zhang FZ, Ye YN, Wu CS, Zheng WX, Rao N, Guo FB (2020) CEG 20: an updated database of clusters of essential genes including eukaryotic organisms. Database (Oxford). https://doi.org/10.1093/database/baaa112
Yang J, Yan R, Roy A, Xu D, Poisson J, Zhang Y (2015) The I-TASSER suite: protein structure and function prediction. Nat Methods 12:7–8. https://doi.org/10.1038/nmeth.3213
Jalal K, Khan K, Hayat A, Ahmad D, Alotaibi G, Uddin R, Mashraqi MM, Alzamami A, Aurongzeb M, Basharat Z (2022) Mining therapeutic targets from the antibiotic-resistant Campylo bactercoli and virtual screening of natural product inhibitors against its riboflavin synthase. Mol Divers. https://doi.org/10.1007/s11030-022-10455-z
Pires DE, Blundell TL, Ascher DB (2015) pkCSM: predicting small-molecule pharmacokinetic and toxicity properties using graph-based signatures. J Med Chem 58:4066–4072. https://doi.org/10.1021/acs.jmedchem.5b00104
Gobeau N, Stringer R, De Buck S, Tuntland T, Faller B (2016) Evaluation of the GastroPlus advanced compartmental and transit (ACAT) model in early discovery. Pharm Res 33:2126–2139. https://doi.org/10.1007/s11095-016-1951-z
Garrido-Franco M (2003) Pyridoxine 5’-phosphate synthase: de novo synthesis of vitamin B6 and beyond. Biochim et Biophys Acta-Protein Proteomics 1647:92–97
Amadasi A, Bertoldi M, Contestabile R, Bettati S, Cellini B, di Salvo ML, Borri-Voltattorni C, Bossa F, Mozzarelli A (2007) Pyridoxal 5’-phosphate enzymes as targets for therapeutic agents. Curr Med Chem 14:1291–1324. https://doi.org/10.2174/092986707780597899
Song S, Sun X, Meng L, Wu Q, Wang K, Deng Y (2021) Antifungal activity of hypocrellin compounds and their synergistic effects with antimicrobial agents against Candida albicans. Microb Biotechnol 14:430–443. https://doi.org/10.1111/1751-7915.13601
Wang J, Iannarelli R, Pucciarelli S, Laudadio E, Galeazzi R, Giangrossi M, Falconi M, Cui L, Navia AM, Buccioni M, Marucci G, Tomassoni D, Serini L, Sut S, Maggi F, Dall’Acqua S, Marchini C, Amici A (2020) Acetylshikonin isolated from Lithospermum erythrorhizon roots inhibits dihydrofolate reductase and hampers autochthonous mammary carcinogenesis in Delta16HER2 transgenic mice. Pharmacol Res 161:1123. https://doi.org/10.1016/j.phrs.2020.105123
Chandrasekara A, Shahidi F (2018) Herbal beverages: Bioactive compounds and their role in disease risk reduction—a review. J Tradit Complement Med 8:451–458. https://doi.org/10.1016/j.jtcme.2017.08.006
Govindan G, Sambandan TG, Govindan M, Sinskey A, Vanessendelft J, Adenan I, Rha CK (2007) A bioactive polyacetylene compound isolated from Centella asiatica. Planta Med 73:597–599. https://doi.org/10.1055/s-2007-981521
Grossmann ME, Mizuno NK, Dammen ML, Schuster T, Ray A, Cleary MP (2009) Eleostearic acid inhibits breast cancer proliferation by means of an oxidation-dependent mechanism. Cancer Prev Res (Phila) 2:879–886. https://doi.org/10.1158/1940-6207.CAPR-09-0088
Amakura Y, Kondo K, Akiyama H, Ito H, Hatano T, Yoshida T, Maitani T (2006) Conjugated ketonic fatty acids from Pleurocybella porrigens. Chem Pharm Bull (Tokyo) 54:1213–1215. https://doi.org/10.1248/cpb.54.1213
Liu L, Hammond EG, Nikolau BJ (1997) In vivo studies of the biosynthesis of [alpha]-eleostearic acid in the seed of Momordica charantia L. Plant Physiol 113:1343–1349. https://doi.org/10.1104/pp.113.4.1343
Hopkins CY, Chisholm MJ, Orgodnik JA (1969) Identity and configuration of conjugated fatty acids in certain seed oils. Lipids 4:89–92
Elfalleh W, Ying M, Nasri N, Sheng-Hua H, Guasmi F, Ferchichi A (2011) Fatty acids from Tunisian and Chinese pomegranate (Punica granatum L.) seeds. Int J Food Sci Nutr 62:200–206. https://doi.org/10.3109/09637486.2010.526932
Radunz A, He P, Schmid GH (2014) Analysis of the seed lipids of Aleurite montana. Zeitschrift für Naturforschung C 53c:305–310
Susilo B, Rohim A, Wahyu ML (2022) Serial extraction technique of rich antibacterial compounds in Sargassum cristaefolium using different solvents and testing their activity. Curr Bioact Compd 18:e100921196341
Naik M, Natarajan V, Rawson A, Rangarajan J, Manickam LE (2021) Extraction kinetics and quality evaluation of oil extracted from bitter gourd (Momardica charantia L.) seeds using emergent technologies. LWT 140:110714
Mao QQ, Xu XY, Cao SY, Gan RY, Corke H, Beta T, Li HB (2019) Bioactive compounds and bioactivities of ginger (Zingiber officinale Roscoe). Foods. https://doi.org/10.3390/foods8060185
Kawamoto Y, Ueno Y, Nakahashi E, Obayashi M, Sugihara K, Qiao S, Iida M, Kumasaka MY, Yajima I, Goto Y, Ohgami N, Kato M, Takeda K (2016) Prevention of allergic rhinitis by ginger and the molecular basis of immunosuppression by 6-gingerol through T cell inactivation. J Nutr Biochem 27:112–122. https://doi.org/10.1016/j.jnutbio.2015.08.025
Abolaji AO, Ojo M, Afolabi TT, Arowoogun MD, Nwawolor D, Farombi EO (2017) Protective properties of 6-gingerol-rich fraction from Zingiber officinale (Ginger) on chlorpyrifos-induced oxidative damage and inflammation in the brain, ovary and uterus of rats. Chem Biol Interact 270:15–23. https://doi.org/10.1016/j.cbi.2017.03.017
Gundala SR, Mukkavilli R, Yang C, Yadav P, Tandon V, Vangala S, Prakash S, Aneja R (2014) Enterohepatic recirculation of bioactive ginger phytochemicals is associated with enhanced tumor growth-inhibitory activity of ginger extract. Carcinogenesis 35:1320–1329. https://doi.org/10.1093/carcin/bgu011
El Houmami N, Bakour S, Bzdrenga J, Rathored J, Seligmann H, Robert C, Armstrong N, Schrenzel J, Raoult D, Yagupsky P, Fournier PE (2017) Isolation and characterization of Kingella negevensis sp. nov., a novel Kingella species detected in a healthy paediatric population. Int J Syst Evol Microbiol 67:2370–2376. https://doi.org/10.1099/ijsem.0.001957
Coote JG (1992) Structural and functional relationships among the RTX toxin determinants of gram-negative bacteria. FEMS Microbiol Rev 8:137–161. https://doi.org/10.1111/j.1574-6968.1992.tb04961.x
Franco MG, Laber B, Huber R, Clausen T (2002) Enzyme–ligand complexes of pyridoxine 5′-phosphate synthase: implications for substrate binding and catalysis. J Mol Biol 321:601–612. https://doi.org/10.1016/S0022-2836(02)00695-2
Franco MG, Laber FB, Huber R, Clausen T (2001) Structural basis for the function of pyridoxine 5′-phosphate synthase. Structure 9:245–253. https://doi.org/10.1016/S0969-2126(01)00584-6
Wu L, Chen W, Wang Z (2021) Traditional Indian medicine in China: the status quo of recognition, development and research. J Ethnopharmacol 279:114317. https://doi.org/10.1016/j.jep.2021.114317
Basharat Z, Jahanzaib M, Yasmin A, Khan IA (2021) Pan-genomics, drug candidate mining and ADMET profiling of natural product inhibitors screened against Yersinia pseudotuberculosis. Genomics 113:238–244. https://doi.org/10.1016/j.ygeno.2020.12.015
Rahaman A, Almalki AA, Rafeeq MM, Akhtar O, Anjum F, Mashraqi MM, Sain ZM, Alzamami A, Ahmad V, Zeng XA, Jamal QMS (2021) Identification of potent natural resource small molecule inhibitor to control Vibrio cholera by targeting its outer membrane protein u: an in silico approach. Molecules. https://doi.org/10.3390/molecules26216517
Salari-Jazi A, Mahnam K, Sadeghi P, Damavandi MS, Faghri J (2021) Discovery of potential inhibitors against New Delhi metallo-beta-lactamase-1 from natural compounds: in silico-based methods. Sci Rep 11:2390. https://doi.org/10.1038/s41598-021-82009-6
Ahmad S, Raza S, Qurat-ul-Ain UR, Rungrotmongkol T, Azam SS (2018) From phylogeny to protein dynamics: a computational hierarchical quest for potent drug identification against an emerging enteropathogen “Yersinia enterocolitica”. J Mol Liq 265:372–389. https://doi.org/10.1016/j.molliq.2018.06.013
Zhang M, Zhao R, Wang D, Wang L, Zhang Q, Wei S, Lu F, Peng W, Wu C (2021) Ginger (Zingiber officinale Rosc.) and its bioactive components are potential resources for health beneficial agents. Phytother Res 35:711–742. https://doi.org/10.1002/ptr.6858
Munda S, Dutta S, Haldar S, Lal M (2018) Chemical analysis and therapeutic uses of ginger (Zingiber officinale Rosc.) essential oil: a review. J Essent Oil Bearing Plants 21:994–1002. https://doi.org/10.1080/0972060X.2018.1524794
Beristain-Bauza SDC, Hernández-Carranza P, Cid-Pérez TS, Ávila-Sosa R, Ruiz-López II, Ochoa-Velasco CE (2019) Antimicrobial activity of ginger (Zingiber officinale) and its application in food products. Food Rev Intl 35:407–426. https://doi.org/10.1080/87559129.2019.1573829
Ahmed N, Karobari MI, Yousaf A, Mohamed RN, Arshad S, Basheer SN, Peeran SW, Noorani TY, Assiry AA, Alharbi AS, Yean CY (2022) The antimicrobial efficacy against selective oral microbes, antioxidant activity and preliminary phytochemical screening of Zingiber officinale. Infect Drug Resist 15:2773–2785. https://doi.org/10.2147/IDR.S364175
Silva FTD, Cunha KFD, Fonseca LM, Antunes MD, Halal S, Fiorentini AM, Zavareze EDR, Dias ARG (2018) Action of ginger essential oil (Zingiber officinale) encapsulated in proteins ultrafine fibers on the antimicrobial control in situ. Int J Biol Macromol 118:107–115. https://doi.org/10.1016/j.ijbiomac.2018.06.079
Silva S, Alves N, Silva P, Vieira T, Maciel P, Castellano LR, Bonan P, Velozo C, Albuquerque D (2019) Antibacterial activity of Rosmarinus officinalis, Zingiber officinale, Citrus aurantium bergamia, and Copaifera officinalis alone and in combination with calcium hydroxide against Enterococcus faecalis. Biomed Res Int 2019:8129439. https://doi.org/10.1155/2019/8129439
Acknowledgements
None.
Funding
None.
Author information
Authors and Affiliations
Contributions
ZB, SMA, AM conceived and designed the experiments. ZB, ZM, AS performed the experiments. ZM, ZB and AS analyzed the data and wrote the paper. SMA, AM curated data and edited the manuscript. ZB, SMA, AM supervised the study.
Corresponding author
Ethics declarations
Conflict of interest
The authors declare that they have no conflict of interest.
Additional information
Publisher's Note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary Information
Below is the link to the electronic supplementary material.
Rights and permissions
Springer Nature or its licensor (e.g. a society or other partner) holds exclusive rights to this article under a publishing agreement with the author(s) or other rightsholder(s); author self-archiving of the accepted manuscript version of this article is solely governed by the terms of such publishing agreement and applicable law.
About this article
Cite this article
Basharat, Z., Murtaza, Z., Siddiqa, A. et al. Therapeutic target mapping from the genome of Kingella negevensis and biophysical inhibition assessment through PNP synthase binding with traditional medicinal compounds. Mol Divers (2023). https://doi.org/10.1007/s11030-023-10604-y
Received:
Accepted:
Published:
DOI: https://doi.org/10.1007/s11030-023-10604-y