Abstract
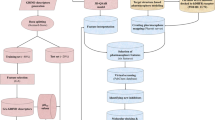
Cytochrome P450-1B1 is a majorly overexpressed drug-metabolizing enzyme in tumors and is responsible for inactivation and subsequent resistance to a variety of anti-cancer drugs, i.e., docetaxel, tamoxifen, and cisplatin. In the present study, a 3D quantitative structure–activity relationship (3D-QSAR) model has been constructed for the identification, design, and optimization of novel CYP1B1 inhibitors. The model has been built using a set of 148 selective CYP1B1 inhibitors. The developed model was evaluated based on certain statistical parameters including q2 and r2 which showed the acceptable predictive and descriptive capability of the generated model. The developed 3D-QSAR model assisted in understanding the key molecular fields which were firmly related to the selective CYP1B1 inhibition. A theoretic approach for the generation of new lead compounds with optimized CYP1B1 receptor affinity has been performed utilizing bioisosteric replacement analysis. These generated molecules were subjected to a developed 3D-QSAR model to predict the inhibitory activity potentials. Furthermore, these compounds were scrutinized through the activity atlas model, molecular docking, electrostatic complementarity, molecular dynamics, and waterswap analysis. The final hits might act as selective CYP1B1 inhibitors which could address the issue of resistance. This 3D-QSAR includes several chemically diverse selective CYP1B1 receptor ligands and well accounts for the individual ligand's inhibition affinities. These features of the developed 3D-QSAR model will ensure future prospective applications of the model to speed up the identification of new potent and selective CYP1B1 receptor ligands.
Graphical abstract















Similar content being viewed by others
References
Lang D, Radtke M, Bairlein M (2019) Highly variable expression of CYP1A1 in human liver and impact on pharmacokinetics of riociguat and granisetron in humans. Chem Res Toxicol 32(6):1115. https://doi.org/10.1021/acs.chemrestox.8b00413
Raju B, Verma H, Narendra G, Sapra B, Silakari O (2021) Multiple machine learning, molecular docking, and ADMET screening approach for identification of selective inhibitors of CYP1B1. J Biomol Struct Dyn. https://doi.org/10.1080/07391102.2021.1905552
Raju B, Narendra G, Verma H, Kumar M, Sapra B, Kaur G, Jain SK, Silakari O (2022) Machine learning enabled structure-based drug repurposing approach to identify potential CYP1B1 inhibitors. ACS Omega 7:31999–32013. https://doi.org/10.1021/acsomega.2c02983
Dong J, Zhang Q, Cui Q, Huang G, Pan X, Li S (2016) Flavonoids and naphthoflavonoids: wider roles in the modulation of cytochrome P450 family 1 enzymes. ChemMedChem 11(19):2102. https://doi.org/10.1002/cmdc.201600316
Raju B, Choudhary S, Narendra G, Verma H, Silakari O (2021) Molecular modeling approaches to address drug-metabolizing enzymes (DMEs) mediated chemoresistance: a review. Drug Metab Rev 53:45–75. https://doi.org/10.1080/03602532.2021.1874406
Shimada T, Hayes CL, Yamazaki H, Amin S, Hecht SS, Guengerich FP, Sutter TR (1996) Activation of chemically diverse procarcinogens by human cytochrome P-450 1B1. Cancer Res 56(13):2979
Guengerich FP, Chun Y-J, Kim D, Gillam EM, Shimada T (2003) Cytochrome P450 1B1: a target for inhibition in anticarcinogenesis strategies. Mutat Res 523:173. https://doi.org/10.1016/S0027-5107(02)00333-0
Don M-J, Lewis DF, Wang S-Y, Tsai M-W, Ueng Y-F (2003) Effect of structural modification on the inhibitory selectivity of rutaecarpine derivatives on human CYP1A1, CYP1A2, and CYP1B1. Bioorg Med Chem Lett 13(15):2535. https://doi.org/10.1016/S0960-894X(03)00469-4
Rl D, Cortés-Benítez F, Roy J, Poirier D (2017) Structure-based design and synthesis of new estrane-pyridine derivatives as cytochrome P450 (CYP) 1B1 inhibitors. ACS Med Chem Lett 8(11):1159. https://doi.org/10.1021/acsmedchemlett.7b00265
Dong J, Wang Z, Cui J, Meng Q, Li S (2020) Synthesis and structure-activity relationship studies of α-naphthoflavone derivatives as CYP1B1 inhibitors. Eur J Med Chem 187:111938. https://doi.org/10.1016/j.ejmech.2019.111938
Murray GI, Taylor MC, McFadyen MC, McKay JA, Greenlee WF, Burke MD, Melvin WT (1997) Tumor-specific expression of cytochrome P450 CYP1B1. Cancer Res 57(14):3026
Shimada T, Yamazaki H, Foroozesh M, Hopkins NE, Alworth WL, Guengerich FP (1998) Selectivity of polycyclic inhibitors for human cytochrome P450s 1A1, 1A2, and 1B1. Chem Res Toxicol 11(9):1048. https://doi.org/10.1021/tx980090+
Wang A, Savas U, Stout CD, Johnson EF (2011) Structural characterization of the complex between α-naphthoflavone and human cytochrome P450 1B1. J Biol Chem 286(7):5736. https://doi.org/10.1074/jbc.M110.204420
Takemura H, Nagayoshi H, Matsuda T, Sakakibara H, Morita M, Matsui A, Ohura T, Shimoi K (2010) Inhibitory effects of chrysoeriol on DNA adduct formation with benzo [a] pyrene in MCF-7 breast cancer cells. Toxicology 274(1–3):42. https://doi.org/10.1016/j.tox.2010.05.009
Kim S, Ko H, Park JE, Jung S, Lee SK, Chun Y-J (2002) Design, synthesis, and discovery of novel trans-stilbene analogues as potent and selective human cytochrome P450 1B1 inhibitors. J Med Chem 45(1):160. https://doi.org/10.1021/jm010298j
Endo K, Uno S, Seki T, Ariga T, Kusumi Y, Mitsumata M, Yamada S, Makishima M (2008) Inhibition of aryl hydrocarbon receptor transactivation and DNA adduct formation by CYP1 isoform-selective metabolic deactivation of benzo [a] pyrene. Toxicol Appl Pharmacol 230(2):135. https://doi.org/10.1016/j.taap.2008.02.009
Hayes CL, Spink DC, Spink BC, Cao JQ, Walker NJ, Sutter TR (1996) 17 beta-estradiol hydroxylation catalyzed by human cytochrome P450 1B1. Proc Natl Acad Sci 93(18):9776. https://doi.org/10.1073/pnas.93.18.9776
Narendra G, Choudhary S, Raju B, Verma H, Silakari O (2022) Role of genetic polymorphisms in drug-metabolizing enzyme-mediated toxicity and pharmacokinetic resistance to anti-cancer agents: a review on the pharmacogenomics aspect. Clin Pharmacokinet. https://doi.org/10.1007/s40262-022-01174-7
Verma H, Bahia MS, Choudhary S, Singh PK, Silakari O (2019) Drug metabolizing enzymes-associated chemo resistance and strategies to overcome it. Drug Metab Rev 51:196–223. https://doi.org/10.1080/03602532.2019.1632886
McFadyen MC, McLeod HL, Jackson FC, Melvin WT, Doehmer J, Murray GI (2001) Cytochrome P450 CYP1B1 protein expression: a novel mechanism of anticancer drug resistance. Biochem Pharmacol 62(2):207. https://doi.org/10.1016/S0006-2952(01)00643-8
Rochat B, Morsman JM, Murray GI, Figg WD, McLeod HL (2001) Human CYP1B1 and anticancer agent metabolism: mechanism for tumor-specific drug inactivation? J Pharmacol Exp Ther 296(2):537
McFadyen MC, Murray GI (2005) Cytochrome P450 1B1: a novel anticancer therapeutic target. J Med Chem 91:10901. https://doi.org/10.1517/14796694.1.2.259
Reid JM, Kuffel MJ, Miller JK, Rios R, Ames MM (1999) Metabolic activation of dacarbazine by human cytochromes P450: the role of CYP1A1, CYP1A2, and CYP2E1. Clin Cancer Res 5(8):2192
Androutsopoulos V, Wilsher N, Arroo RR, Potter GA (2009) Bioactivation of the phytoestrogen diosmetin by CYP1 cytochromes P450. Cancer Lett 274(1):54. https://doi.org/10.1016/j.canlet.2008.08.032
Takemura H, Itoh T, Yamamoto K, Sakakibara H, Shimoi K (2010) Selective inhibition of methoxyflavonoids on human CYP1B1 activity. Bioorg Med Chem 18(17):6310. https://doi.org/10.1016/j.bmc.2010.07.020
Sridhar J, Liu J, Foroozesh M, Klein Stevens CL (2012) Inhibition of cytochrome p450 enzymes by quinones and anthraquinones. Chem Res Toxicol 25(2):357. https://doi.org/10.1021/tx2004163
Mammen JS, Kleiner HE, DiGiovanni J, Sutter TR, Strickland PT (2005) Coumarins are competitive inhibitors of cytochrome P450 1B1, with equal potency for allelic variants. Pharmacogenet Genomics 15(3):183
Dutour R, Poirier D (2017) Inhibitors of cytochrome P450 (CYP) 1B1. Eur J Med Chem 135:296. https://doi.org/10.1016/j.ejmech.2017.04.042
Williams IS, Joshi P, Gatchie L, Sharma M, Satti NK, Vishwakarma RA, Chaudhuri B, Bharate SB (2017) Synthesis and biological evaluation of pyrrole-based chalcones as CYP1 enzyme inhibitors, for possible prevention of cancer and overcoming cisplatin resistance. Bioorg Med Chem Lett 27(16):3683. https://doi.org/10.1016/j.bmcl.2017.07.010
Horley NJ, Beresford KJ, Chawla T, McCann GJ, Ruparelia KC, Gatchie L, Sonawane VR, Williams IS, Tan HL, Joshi P (2017) Discovery and characterization of novel CYP1B1 inhibitors based on heterocyclic chalcones: overcoming cisplatin resistance in CYP1B1-overexpressing lines. Eur J Med Chem 129:159. https://doi.org/10.1016/j.ejmech.2017.02.016
Dong J, Huang G, Zhang Q, Wang Z, Cui J, Wu Y, Meng Q, Li S (2019) Development of benzochalcone derivatives as selective CYP1B1 inhibitors and anticancer agents. MedChemComm 10(9):1606. https://doi.org/10.1039/C9MD00258H
Tan HL (2006) Selective inhibitors of the cytochrome P450 enzyme CYP1B1: De Montfort University. https://ethos.bl.uk/OrderDetails.do?uin=uk.bl.ethos.502100
Rl D, Roy J, Cortés-Benítez F, Maltais R, Poirier D (2018) Targeting Cytochrome P450 (CYP) 1B1 Enzyme with four series of a-ring substituted estrane derivatives: design, synthesis, inhibitory activity, and selectivity. J Med Chem 61(20):9229. https://doi.org/10.1021/acs.jmedchem.8b00907
Meng Q, Wang Z, Cui J, Cui Q, Dong J, Zhang Q, Li S (2018) Design, synthesis, and biological evaluation of cytochrome P450 1B1 targeted molecular imaging probes for colorectal tumor detection. J Med Chem 61(23):10901. https://doi.org/10.1021/acs.jmedchem.8b01633
Chun Y-J, Lim C, Ohk SO, Lee JM, Lee JH, Choi S, Kim S (2011) trans-Stilbenoids: potent and selective inhibitors for human cytochrome P450 1B1. MedChemComm 2(5):402. https://doi.org/10.1039/C0MD00242A
Cui J, Meng Q, Zhang X, Cui Q, Zhou W, Li S (2015) Design and synthesis of new α-naphthoflavones as cytochrome P450 (CYP) 1B1 inhibitors to overcome docetaxel-resistance associated with CYP1B1 overexpression. J Med Chem 58(8):3534. https://doi.org/10.1021/acs.jmedchem.5b00265
Kubo M, Yamamoto K, Itoh T (2019) Design and synthesis of selective CYP1B1 inhibitor via dearomatization of α-naphthoflavone. Bioorg Med Chem 27(2):285. https://doi.org/10.1016/j.bmc.2018.11.045
Mikstacka R, Przybylska D, Rimando AM, Baer-Dubowska W (2007) Inhibition of human recombinant cytochromes P450 CYP1A1 and CYP1B1 by trans-resveratrol methyl ethers. Mol Nutr Food Res 51(5):517. https://doi.org/10.1002/mnfr.200600135
Mikstacka R, Baer-Dubowska W, Wieczorek M, Sobiak S (2008) Thiomethylstilbenes as inhibitors of CYP1A1, CYP1A2 and CYP1B1 activities. Mol Nutr Food Res 52(S1):S77. https://doi.org/10.1002/mnfr.200700202
Siddique MUM, McCann GJ, Sonawane V, Horley N, Williams IS, Joshi P, Bharate SB, Jayaprakash V, Sinha BN, Chaudhuri B (2016) Biphenyl urea derivatives as selective CYP1B1 inhibitors. Org Biomol Chem 14(38):8931. https://doi.org/10.1039/C6OB01506A
Iimoto D (2018) Preventing carcinogenesis with compounds that inhibit cytochrome P450 1A1 and 1B1. Biochem Physiol 7(230):2. https://doi.org/10.4172/2168-9652.1000230
Cheeseright T, Mackey M, Rose S, Vinter A (2006) Molecular field extrema as descriptors of biological activity: definition and validation. J Chem Info Model 46(2):665. https://doi.org/10.1021/ci050357s
De Jong S (1993) SIMPLS: an alternative approach to partial least squares regression. Chemometr Intell Lab Syst 18(3):251. https://doi.org/10.1016/0169-7439(93)85002-X
Alam S, Khan F (2014) QSAR and docking studies on xanthone derivatives for anticancer activity targeting DNA topoisomerase IIα. Drug Des Devel Ther 8:183. https://doi.org/10.2147/DDDT.S51577
Floresta G, Rescifina A, Marrazzo A, Dichiara M, Pistarà V, Pittalà V, Prezzavento O, Amata E (2017) Hyphenated 3D-QSAR statistical model-scaffold hopping analysis for the identification of potentially potent and selective sigma-2 receptor ligands. Eur J Med Chem 139:884. https://doi.org/10.1016/j.ejmech.2017.08.053
Wagener M, Lommerse JP (2006) The quest for bioisosteric replacements. J Chem Info Model 46(2):677. https://doi.org/10.1021/ci0503964
Langdon SR, Ertl P, Brown N (2010) Bioisosteric replacement and scaffold hopping in lead generation and optimization. Mol Inform 29(5):366. https://doi.org/10.1002/minf.201000019
Walsh AA, Szklarz GD, Scott EE (2013) Human cytochrome P450 1A1 structure and utility in understanding drug and xenobiotic metabolism. J Biol Chem 288(18):12932. https://doi.org/10.1074/jbc.M113.452953
Scrocco E, Tomasi J (1973) The electrostatic molecular potential as a tool for the interpretation of molecular properties. New concept II. Springer, New York, p 95
Verma H, Narendra G, Raju B, Kumar M, Jain SK, Tung GK, Singh PK, Silakari O (2022) 3D-QSAR and scaffold hopping based designing of benzo [d] ox-azol-2 (3H)-one and 2-oxazolo [4, 5-b] pyridin-2 (3H)-one derivatives as selective aldehyde dehydrogenase 1A1 inhibitors: synthesis and biological evaluation. Arch Pharm 355(9):1–25. https://doi.org/10.1002/ardp.202200108
Bauer MR, Mackey MD (2019) Electrostatic complementarity as a fast and effective tool to optimize binding and selectivity of protein–ligand complexes. J Med Chem 62(6):3036. https://doi.org/10.1021/acs.jmedchem.8b01925
Gupta S, Jadaun A, Kumar H, Raj U, Varadwaj PK, Rao A (2015) Exploration of new drug-like inhibitors for serine/threonine protein phosphatase 5 of Plasmodium falciparum: a docking and simulation study. J Biomol Struct Dyn 33(11):2421. https://doi.org/10.1080/07391102.2015.1051114
Acknowledgements
BR would like to acknowledge the Indian Council of Medical Research (ICMR), New Delhi for providing a Senior Research Fellowship (SRF).
Funding
This work was financially supported by the Indian Council of Medical Research (ICMR), New Delhi [ISRM/12(10)/2019].
Author information
Authors and Affiliations
Corresponding author
Ethics declarations
Conflict of interest
No potential conflict of interest was reported by the author(s).
Additional information
Publisher's Note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary Information
Below is the link to the electronic supplementary material.
Rights and permissions
Springer Nature or its licensor (e.g. a society or other partner) holds exclusive rights to this article under a publishing agreement with the author(s) or other rightsholder(s); author self-archiving of the accepted manuscript version of this article is solely governed by the terms of such publishing agreement and applicable law.
About this article
Cite this article
Raju, B., Sapra, B. & Silakari, O. 3D-QSAR assisted identification of selective CYP1B1 inhibitors: an effective bioisosteric replacement/molecular docking/electrostatic complementarity analysis. Mol Divers 27, 2673–2693 (2023). https://doi.org/10.1007/s11030-022-10574-7
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s11030-022-10574-7