Abstract
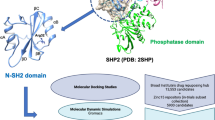
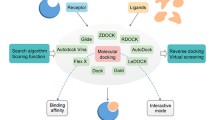
Matrix metalloproteinase-2 (MMP-2) is capable of degrading Collage TypeIV in the vascular basement membrane and extracellular matrix. Studies have shown that MMP-2 is tightly associated with the biological behavior of malignant tumors. Therefore, the identification of inhibitors targeting MMP-2 could be effective in treating the disease by maintaining extracellular matrix homeostasis. In the pharmaceutical and biomedical fields, many computational tools are widely used, which improve the efficiency of the whole process to some extent. Apart from the conventional cheminformatics approaches (e.g., pharmacophore model and molecular docking), virtual screening strategies based on machine learning also have promising applications. In this study, we collected 2871 compound activity data against MMP-2 from the ChEMBL database and divided the training and test sets in a 3:1 ratio. Four machine learning algorithms were then selected to construct the classification models, and the best-performing model, i.e., the stacking-based fusion model with the highest AUC value in both training and test datasets, was used for the virtual screening of ZINC database. Next, we screened 17 potential MMP-2 inhibitors from the results predicted by the machine learning model via ADME/T analysis. The interactions between these compounds and the target protein were explored through molecular docking calculations, and the results showed that ZINC712249, ZINC4270723, and ZINC15858504 had lower binding free energies than the co-crystal ligand. To further examine the binding stability of the complexes, we performed molecular dynamics simulations and finally identified these three hits as the most promising natural products for MMP-2 inhibitors.
Graphical Abstract










Similar content being viewed by others
Data Availability
The data that support the findings of this study are available from the corresponding author upon reasonable request.
References
Verma RP, Hansch C (2007) Matrix metalloproteinases (MMPs): chemical–biological functions and (Q)SARs. Bioorgan Med Chem 15:2223–2268. https://doi.org/10.1016/j.bmc.2007.01.011
Cui N, Hu M, Khalil RA (2017) Biochemical and biological attributes of matrix metalloproteinases. Prog Mol Biol Transl 147:1–73. https://doi.org/10.1016/bs.pmbts.2017.02.005
Nissinen L, Kähäri V (2014) Matrix metalloproteinases in inflammation. BBA-Gen Subjects 1840:2571–2580. https://doi.org/10.1016/j.bbagen.2014.03.007
Mezentsev A, Nikolaev A, Bruskin S (2014) Matrix metalloproteinases and their role in psoriasis. Genes-Basel 540:1–10. https://doi.org/10.1016/j.gene.2014.01.068
Knapinska AM, Estrada C, Fields GB (2017) The roles of matrix metalloproteinases in pancreatic cancer. Prog Mol Biol Transl 148:339–354. https://doi.org/10.1016/bs.pmbts.2017.03.004
Alg VS, Ke X, Grieve J et al (2018) Association of functional MMP-2 gene variant with intracranial aneurysms: case-control genetic association study and meta-analysis. Brit J Neurosurg 32:255–259. https://doi.org/10.1080/02688697.2018.1427213
Ulrich D, Hrynyschyn K, Pallua N (2003) Matrix metalloproteinases and tissue inhibitors of metalloproteinases in sera and tissue of patients with Dupuytren’ s disease. Plast Reconstr Surg 112:1279–1286. https://doi.org/10.1097/01.PRS.0000081462.40448.49
Amin SA, Adhikari N, Jha T (2017) Is dual inhibition of metalloenzymes HDAC-8 and MMP-2 a potential pharmacological target to combat hematological malignancies? Pharmacol Res 122:8–19. https://doi.org/10.1016/j.phrs.2017.05.002
Huang C, Teng Y, Lu F et al (2017) β-mangostin suppresses human hepatocellular carcinoma cell invasion through inhibition of MMP-2 and MMP-9 expression and activating the ERK and JNK pathways. Environ Toxicol 32:2360–2370. https://doi.org/10.1002/tox.22449
Newman DJ (2020) Modern traditional Chinese medicine: identifying, defining and usage of TCM components. Adv Pharmacol 87:113–158. https://doi.org/10.1016/bs.apha.2019.07.001
Liu S, Xu X, Xu J et al (2017) Multi-drug resistant uropathogenic Escherichia coli and its treatment by Chinese medicine. Chin J Integr Med 23:763–769. https://doi.org/10.1007/s11655-016-2738-0
Leuci R, Brunetti L, Poliseno V et al (2021) Natural compounds for the prevention and treatment of cardiovascular and neurodegenerative diseases. Foods 10:29. https://doi.org/10.3390/foods10010029
Bian Y, Feng Z, Yang P et al (2017) integrated in silico fragment-based drug design: case study with allosteric modulators on metabotropic glutamate receptor 5. AAPS J 19:1235–1248. https://doi.org/10.1208/s12248-017-0093-5
Herrera-Acevedo C, Perdomo-Madrigal C, Herrera-Acevedo K et al (2021) Machine learning models to select potential inhibitors of acetylcholinesterase activity from SistematX: a natural products database. Mol Divers 25:1553–1568. https://doi.org/10.1007/s11030-021-10245-z
Islam MA, Rallabandi VPS, Mohammed S et al (2021) Screening of β1- and β2-adrenergic receptor modulators through advanced pharmacoinformatics and machine learning approaches. Int J Mol Sci 22:11191. https://doi.org/10.3390/ijms222011191
Fernández-de Gortari E, García-Jacas CR, Martinez-Mayorga K et al (2017) Database fingerprint (DFP): an approach to represent molecular databases. J Cheminform 9:9. https://doi.org/10.1186/s13321-017-0195-1
Lin H, Han L, Yap C et al (2007) Prediction of factor Xa inhibitors by machine learning methods. J Mol Graph 26:505–518. https://doi.org/10.1016/j.jmgm.2007.03.003
Pedregosa F, Varoquaux G, Gramfort A et al (2011) Scikit-learn: machine learning in python. J Mach Learn Res 12:2825–2830. https://doi.org/10.5555/1953048.2078195
Trott O, Olson AJ (2009) AutoDock Vina: improving the speed and accuracy of docking with a new scoring function, efficient optimization, and multithreading. J Comput Chem. https://doi.org/10.1002/jcc.21334
Morris GM, Huey R, Lindstrom W et al (2009) AutoDock4 and AutoDockTools4: automated docking with selective receptor flexibility. J Comput Chem 30:2785–2791. https://doi.org/10.1002/jcc.21256
Salentin S, Schreiber S, Haupt VJ et al (2015) PLIP: fully automated protein–ligand interaction profiler. Nucl Acids Res 43:W443–W447. https://doi.org/10.1093/nar/gkv315
Yuan S, Chan HCS, Hu Z (2017) Using PyMOL as a platform for computational drug design. Wiley interdisciplinary reviews. Wires Comput Mol Sci. https://doi.org/10.1002/wcms.1298
Pronk S, Páll S, Schulz R et al (2013) GROMACS 4.5: a high-throughput and highly parallel open source molecular simulation toolkit. Bioinformatics 29:845–854. https://doi.org/10.1093/bioinformatics/btt055
Gramatica P, Giani E, Papa E (2007) Statistical external validation and consensus modeling: a QSPR case study for Koc prediction. J Mol Graph Model 25:755–766. https://doi.org/10.1016/j.jmgm.2006.06.005
Mak KK, Pichika MR (2019) Artificial intelligence in drug development: present status and future prospects. Drug Discov Today 24:773–780. https://doi.org/10.1016/j.drudis.2018.11.014
Funding
This research was funded by Sub-project of the National Ministry of Health Major New Drug Creation Science and Technology Major Project (No. 2014ZX09509001001).
Author information
Authors and Affiliations
Corresponding authors
Ethics declarations
Conflict of interest
The authors declare no conflict of interest.
Additional information
Publisher's Note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary Information
Below is the link to the electronic supplementary material.
Rights and permissions
About this article
Cite this article
Yang, R., Zhao, G., Cheng, B. et al. Identification of potential matrix metalloproteinase-2 inhibitors from natural products through advanced machine learning-based cheminformatics approaches. Mol Divers 27, 1053–1066 (2023). https://doi.org/10.1007/s11030-022-10467-9
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s11030-022-10467-9