Abstract
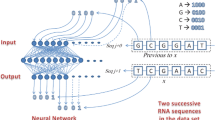
Recently, we defined the randomness within a protein as an important force engineering mutations. Thereafter we build a cause–mutation relationship, where one side is the quantified randomness and the other side is the occurrence or non-occurrence of mutation. This way, we switch the prediction of mutation into the problem of classification, which can be solved using either logistic regression or neural network. Very recently, we attempted to apply the logistic regression to predicting the mutation positions in proteins from influenza A virus. In this study, we attempt to explore the possibility of applying the neural network to predicting the mutation positions in H1 neuraminidase from influenza A virus. Then we applied the amino-acid mutating probability to predicting the would-be-mutated amino acids at predicted positions. The results confirm the possibility of prediction of mutation using this approach and pave the way for future development.
Similar content being viewed by others
References
Everitt BS (1999). Chance rules: an informal guide to probability, risk and statistics. Springer, New York
Wu G and Yan S (2002). Randomness in the primary structure of protein: methods and implications. Mol Biol Today 3: 55–69
Wu G and Yan S (2006). Fate of influenza A virus proteins. Protein Pept Lett 13: 377–384
Wu G and Yan S (2006). Mutation trend of hemagglutinin of influenza A virus: a review from computational mutation viewpoint. Acta Pharmacol Sin 27: 513–526
Draper NR and Smith H (1981). Applied regression analysis, 2nd edn. Wiley, New York
Hosmer DW Jr and Lemeshow S (2000). Applied logistic regression, 2nd edn. Wiley, New York
Jordan MI, Bishop CM (1997) Neural networks. In: Tucker AB (ed) The computer science and engineering handbook. CRC Press, pp 536–556
Wu G and Yan S (2005). Determination of mutation trend in proteins by means of translation probability between RNA codes and mutated amino acids. Biochem Biophys Res Commun 337: 692–700
Wu G and Yan S (2006). Determination of mutation trend in hemagglutinins by means of translation probability between RNA codons and mutated amino acids. Protein Pept Lett 13: 601–609
Wu G and Yan S (2007). Translation probability between RNA codons and translated amino acids and its applications to protein mutations. In: Ostrovskiy, MH (eds) Leading-edge messenger RNA research communications, chap 3, pp 47–65. Nova Science Publishers, New York
Wu G and Yan S (2006). Prediction of possible mutations in H5N1 hemagglutinins of influenza A virus by means of logistic regression. Comp Clin Pathol 15: 255–261
Wu G and Yan S (2006). Prediction of mutations in H5N1 hemagglutinins from influenza A virus. Protein Pept Lett 13: 971–976
Wu G and Yan S (2007). Improvement of model for prediction of hemagglutinin mutations in H5N1 influenza viruses with distinguishing of arginine, leucine and serine. Protein Pept Lett 14: 191–196
Hochgürtel M, Kroth H and Piecha D et al. (2002). Target-induced formation of neuraminidase inhibitors from in vitro virtual combinatorial libraries. PNAS 99: 3382–3387
Garman E and Laver G (2004). Controlling influenza by inhibiting the virus’s neuraminidase. Curr Drug Targ 5: 119–136
Oxford J, Balasingam S and Lambkin R (2004). A new millennium conundrum: how to use a powerful class of influenza anti-neuraminidase drugs (NAIs) in the community. J Antimicrob Chemother 53: 133–136
Influenza virus resources (2006) http://www.ncbi.nlm.nih.gov/genomes/FLU/Database/multiple.cgi
Hagan MT, Demuth HB and Beale MH (1996). Neural network design. PWS Publishing Company, Boston, MA
Demuth H, Beale M (2001) Neural network toolbox for use with MatLab. User’s guide, version 4
PROSITE (2002) A dictionary of protein sites and patterns user manual. http://www.expasy.ch/prosite/
Feller W (1968). An introduction to probability theory and its applications, 3rd edn, vol I. Wiley, New York, 34–40
Wu G and Yan S (2000). Prediction of distributions of amino acids and amino acid pairs in human haemoglobin α-chain and its seven variants causing α-thalassemia from their occurrences according to the random mechanism. Comp Haematol Int 10: 80–84
MathWorks Inc (2001) MatLab—the language of technical computing. (version 6.1.0.450, release 12.1) 1984–200
Healy MJR (1979). Outliers in clinical chemistry quality-control schemes. Clin Chem 25: 675–677
SYSTAT Software Inc (2004) Systat for Windows, version 11.00.01
Wasserman PD (1993). Advanced methods in neural computing. Van Nostrand Reinhold, New York, 35–55
Kohonen T (1987). Self-organization and associative memory, 2nd edn. Springer-Verlag, Berlin
Wu G and Yan S (2005). Timing of mutation in hemagglutinins from influenza A virus by means of unpredictable portion of amino-acid pair and fast Fourier transform. Biochem Biophys Res Commun 333: 70–78
Wu G and Yan S (2006). Timing of mutation in hemagglutinins from influenza A virus by means of amino-acid distribution rank and fast Fourier transform. Protein Pept Lett 13: 143–148
Author information
Authors and Affiliations
Corresponding author
Rights and permissions
About this article
Cite this article
Wu, G., Yan, S. Prediction of mutations in H1 neuraminidases from North America influenza A virus engineered by internal randomness. Mol Divers 11, 131–140 (2007). https://doi.org/10.1007/s11030-008-9067-y
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s11030-008-9067-y