Abstract
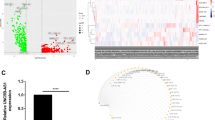
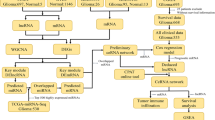
Evidence suggests that non-coding RNAs have a role in glioblastoma multiforme (GBM), although the regulatory mechanisms controlled by competing endogenous RNAs (ceRNAs) in GBM are still poorly understood and infrequently described. This research extensively analyzed circRNA, lncRNA, miRNA, and mRNA expression changes in GBM patients. RNA-sequencing analyses were conducted to investigate differentially expressed genes (DEGs), lncRNAs (DELs), miRNAs (DEMs), and circRNAs (DECs) in the GBM. In this study, researchers found that GBM patients and healthy controls differed in the presence of 1224 DECs, 1406 DELs, 229 DEMs, and 2740 DEGs. PPI network analysis demonstrated that CEACAM5, CXCL17, FAM83A, TMPRSS4, and GGPRC5A were hub genes and enriched in modules. Then a ceRNA network was constructed with 8 circRNA, 7 lncRNAs, 16 miRNAs, and 17 mRNAs. Overall, the ceRNA interaction axes that were found may prove to be pivotal therapeutic targets for treating GBM.







Similar content being viewed by others
Data Availability
The datasets used and/or analyzed during the current study are available from the corresponding author upon reasonable request.
Abbreviations
- GBM:
-
Glioblastoma multiforme
- TCGA:
-
The cancer genome atlas
- ceRNAs:
-
Competing endogenous RNAs
- circRNA:
-
circular RNA
- lncRNAs:
-
Long non-coding RNAs
- miRNA/miR:
-
microRNA
- DEGs:
-
Differentially expressed genes
- PPI:
-
Protein-protein interaction
- CEACAM5:
-
Carcinoembryonic antigen-related cell adhesion molecule 5
- CXCL17:
-
C-X-C motif chemokine ligand 17
- FAM83A:
-
Family with sequence similarity 83 member A
- TMPRSS4:
-
Transmembrane serine protease 4
- GPRC5A:
-
G protein-coupled receptor class C group 5 member A
References
Abdul-Wahid A, Cydzik M, Fischer NW, Prodeus A, Shively JE, Martel A et al (2018) Serum-derived carcinoembryonic antigen (CEA) activates fibroblasts to induce a local re-modeling of the extracellular matrix that favors the engraftment of CEA-expressing tumor cells. Int J Cancer 143:1963–1977
Ambrosi C, Scribano D, Sarshar M, Zagaglia C, Singer BB, Palamara AT (2020) Acinetobacter baumannii targets Human Carcinoembryonic Antigen-Related cell adhesion molecules (CEACAMs) for Invasion of Pneumocytes. mSystems. ;5
Azari F, Kennedy GT, Chang A, Bernstein E, Nadeem B, Pèlegrin A et al (2022) Glycoprotein receptor CEACAM5-Targeted intraoperative Molecular Imaging Tracer in Non-Small Cell Lung Cancer. The Annals of thoracic surgery
Baek DS, Kim YJ, Vergara S, Conard A, Adams C, Calero G et al (2022) A highly-specific fully-human antibody and CAR-T cells targeting CD66e/CEACAM5 are cytotoxic for CD66e-expressing cancer cells in vitro and in vivo. Cancer Lett 525:97–107
Basso J, Paggi MG, Fortuna A, Vitorino C (2022) Deciphering specific miRNAs in brain tumors: a 5-miRNA signature in glioblastoma. 297:507–521
Cheng J, Meng J, Zhu L, Peng Y (2020) Exosomal non-coding RNAs in glioma: biological functions and potential clinical applications. Mol Cancer 19:66
Choreño-Parra JA, Thirunavukkarasu S, Zúñiga J, Khader SA (2020) The protective and pathogenic roles of CXCL17 in human health and disease: potential in respiratory medicine. Cytokine Growth Factor Rev 53:53–62
Gisina A, Novikova S, Kim Y, Sidorov D, Bykasov S, Volchenko N et al (2021) CEACAM5 overexpression is a reliable characteristic of CD133-positive colorectal cancer stem cells. Cancer Biomark A 32:85–98
Guo X, Piao H (2021) Research Progress of circRNAs in Glioblastoma. Front cell Dev biology 9:791892
Holmer R, Wätzig GH, Tiwari S, Rose-John S, Kalthoff H (2015) Interleukin-6 trans-signaling increases the expression of carcinoembryonic antigen-related cell adhesion molecules 5 and 6 in colorectal cancer cells. BMC Cancer 15:975
Ji H, Song H, Wang Z, Jiao P, Xu J, Li X et al (2021) FAM83A promotes proliferation and metastasis via Wnt/β-catenin signaling in head neck squamous cell carcinoma. 19:423
Jia B, Liu W, Gu J, Wang J, Lv W, Zhang W et al (2019) MiR-7-5p suppresses stemness and enhances temozolomide sensitivity of drug-resistant glioblastoma cells by targeting Yin Yang 1. Exp Cell Res 375:73–81
Kelleher M, Singh R, O’Driscoll CM, Melgar S (2019) Carcinoembryonic antigen (CEACAM) family members and inflammatory bowel disease. Cytokine Growth Factor Rev 47:21–31
Le Rhun E, Preusser M, Roth P, Reardon DA, van den Bent M, Wen P et al (2019) Molecular targeted therapy of glioblastoma. Cancer Treat Rev 80:101896
Li L, Ji Y, Chen Y, Zhen Z (2021) MiR-325-3p mediate the CXCL17/CXCR8 axis to regulate angiogenesis in hepatocellular carcinoma. Cytokine 141:155436
Liu R, Dai W (2021) CircCDC45 promotes the malignant progression of glioblastoma by modulating the miR-485-5p/CSF-1 axis. 21:1090
Ma R, Taphoorn MJB, Plaha P (2021) Advances in the management of glioblastoma. J Neurol Neurosurg Psychiatry 92:1103–1111
Ohba S, Hirose Y (2016) Current and future drug treatments for glioblastomas. Curr Med Chem 23:4309–4316
Ohgaki H, Kleihues P (2013) The definition of primary and secondary glioblastoma. Clin cancer research: official J Am Association Cancer Res 19:764–772
Omuro A, DeAngelis LM (2013) Glioblastoma and other malignant gliomas: a clinical review. JAMA 310:1842–1850
Peng Z, Liu C, Wu M (2018) New insights into long non-coding RNAs and their roles in glioma. Mol Cancer 17:61
Powell E, Shao J, Picon HM, Bristow C, Ge Z, Peoples M et al (2018) A functional genomic screen in vivo identifies CEACAM5 as a clinically relevant driver of breast cancer metastasis. NPJ breast cancer 4:9
Robin AM, Lee I, Kalkanis SN (2017) Reoperation for recurrent Glioblastoma Multiforme. Neurosurg Clin North Am 28:407–428
Shea A, Harish V, Afzal Z, Chijioke J, Kedir H, Dusmatova S et al (2016) MicroRNAs in glioblastoma multiforme pathogenesis and therapeutics. Cancer Med 5:1917–1946
Simpson L, Galanis E (2006) Recurrent glioblastoma multiforme: advances in treatment and promising drug candidates. Expert Rev Anticancer Ther 6:1593–1607
Wang Y, Xu R, Zhang D, Lu T, Yu W, Wo Y et al (2019) Circ-ZKSCAN1 regulates FAM83A expression and inactivates MAPK signaling by targeting mir-330-5p to promote non-small cell lung cancer progression. Translational lung cancer research 8:862–875
Zhang F, Jin Liu X, Qu X, Sheng Hu Z, Yang YM, Ma L et al (2012) Osteopontin’s colocalization with the adhesion molecule CEACAM5 in cytoplasm of carcinoma of tongue and its correlation with the invasion of that diease. Cancer Cell Int 12:33
Zhang M, Zhao K, Xu X, Yang Y, Yan S, Wei P et al (2018) A peptide encoded by circular form of LINC-PINT suppresses oncogenic transcriptional elongation in glioblastoma. 9:4475
Zhang X, Niu W, Mu M, Hu S, Niu C (2020) Long non-coding RNA LPP-AS2 promotes glioma tumorigenesis via miR-7-5p/EGFR/PI3K/AKT/c-MYC feedback loop. Journal of experimental & clinical cancer research: CR. ;39:196
Zhao C, Guo R, Guan F (2020) MicroRNA-128-3p enhances the chemosensitivity of Temozolomide in Glioblastoma by Targeting c-Met and EMT. 10:9471
Zhou J, Fan X, Chen N, Zhou F, Dong J, Nie Y et al (2015) Identification of CEACAM5 as a biomarker for prewarning and prognosis in gastric Cancer. J Histochem cytochemistry: official J Histochem Soc 63:922–930
Acknowledgements
Not Applicable.
We thank all participants involved in this study and all authors. We would like to thank Dr. Pu Yong of Tianjin Ruijiyin Biotechnology Co., Ltd. for his assistance in our bioinformatics analysis.
Funding
This work was supported by the National Natural Science Foundation of China, No. 81871086 (to LHQ), the Tianjin Science and Technology Project, No. 21JCQNJC01400 (to WRJ).
Author information
Authors and Affiliations
Contributions
RW, QL, HL and FH conceived and designed the study. RW and QL collected and processed data. XC and NL analyzed data. RW and QL drafted the manuscript. HL and FH revised the manuscript. All authors read and approved the final manuscript.
Acknowledgements.
Corresponding authors
Ethics declarations
Ethics approval and consent to participate
This study was performed conforming to the tenets of Declaration of Helsinki and written informed consent was obtained from all patients prior to the investigation. An ethics approval was obtained from the Characteristic Medical Center of Chinese People’s Armed Police Force (No. 2022 − 0002.1) prior to the collection of tissue samples. The funders had no role in the study design, data collection and analysis, data interpretation, or preparation of the manuscript.
Consent for publication
Not Applicable.
Competing interests
The authors declare no competing interests.
Additional information
Renjie Wang, Qi Li and Xiaolei Chu contributed equally to this work.
Publisher’s Note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Rights and permissions
Springer Nature or its licensor (e.g. a society or other partner) holds exclusive rights to this article under a publishing agreement with the author(s) or other rightsholder(s); author self-archiving of the accepted manuscript version of this article is solely governed by the terms of such publishing agreement and applicable law.
About this article
Cite this article
Wang, R., Li, Q., Chu, X. et al. Sequencing and Bioinformatics analysis of lncRNA/circRNA-miRNA-mRNA in Glioblastoma multiforme. Metab Brain Dis 38, 2289–2300 (2023). https://doi.org/10.1007/s11011-023-01256-w
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s11011-023-01256-w