Abstract
Background
Histone deacetylase (HDAC) inhibitor-based therapeutic drug tolerance is a major obstacle to glioblastoma (GBM) treatment. Meanwhile, non-coding RNAs have been reported to be involved in the regulation of HDAC inhibitor (SAHA) tolerance in some human tumors. However, the relationship between circular RNAs (circRNAs) and SAHA tolerance is still unknown. Herein, we explored the role and mechanism of circ_0000741 on SAHA tolerance in GBM.
Methods
Circ_0000741, microRNA-379-5p (miR-379-5p), and tripartite motif-containing 14 (TRIM14) level were detected by real-time quantitative polymerase chain reaction (RT-qPCR). (4-5-dimethylthiazol-2-yl)-2,5-diphenyl tetrazolium bromide (MTT), 5-ethynyl-2’-deoxyuridine (EdU), Colony formation, flow cytometry, and transwell assays were used to detect SAHA tolerance, proliferation, apoptosis, and invasion in SAHA-tolerant GBM cells. Western blot analysis of protein levels of E-cadherin, N-cadherin, and TRIM14. After Starbase2.0 analysis, the binding between miR-379-5p and circ_0000741 or TRIM14 was proved using a dual-luciferase reporter. The role of circ_0000741 on drug tolerance was assessed using a xenograft tumor model in vivo.
Results
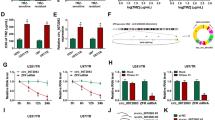
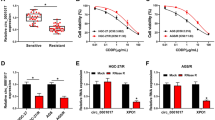
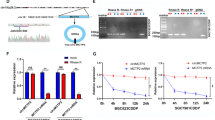
Circ_0000741 and TRIM14 were upregulated, and miR-379-5p was reduced in SAHA-tolerant GBM cells. Furthermore, circ_0000741 absence reduced SAHA tolerance, suppressed proliferation, invasion, and induced apoptosis in SAHA-tolerant GBM cells. Mechanistically, circ_0000741 might affect TRIM14 content via sponging miR-379-5p. Besides, circ_0000741 silencing enhanced the drug sensitivity of GBM in vivo.
Conclusion
Circ_0000741 might accelerate SAHA tolerance by regulating the miR-379-5p/TRIM14 axis, which provided a promising therapeutic target for GBM treatment.








Similar content being viewed by others
Data availability
The datasets used and analyzed during the current study are available from the corresponding author on reasonable request.
References
Abbas MN, Kausar S, Cui H (2020) Therapeutic potential of natural products in glioblastoma treatment: targeting key glioblastoma signaling pathways and epigenetic alterations. Clin Transl Oncol 22(7):963–977
Adams CM, Eischen CM (2016) Histone deacetylase inhibition reveals a tumor-suppressive function of MYC-regulated miRNA in breast and lung carcinoma. Cell Death Differ 23(8):1312–1321
Barrett SP, Salzman J (2016) Circular RNAs: analysis, expression and potential functions. Development 143(11):1838–1847
Bartel DP (2018) Metazoan MicroRNAs Cell 173(1):20–51
Belousova EA, Filipenko ML, Kushlinskii NE (2018) Circular RNA: New Regulatory Molecules. Bull Exp Biol Med 164(6):803–815
Bhat AA, Younes SN, Raza SS, Zarif L, Nisar S, Ahmed I et al (2020) Role of non-coding RNA networks in leukemia progression, metastasis and drug resistance. Mol Cancer 19(1):57
Bian X, Liang Z, Feng A, Salgado E, Shim H (2018) HDAC inhibitor suppresses proliferation and invasion of breast cancer cells through regulation of miR-200c targeting CRKL. Biochem Pharmacol 147:30–37
Campos B, Olsen LR, Urup T, Poulsen HS (2016) A comprehensive profile of recurrent glioblastoma. Oncogene 35(45):5819–5825
Chen B, Chen C, Zhang Y, Xu J (2021) Recent incidence trend of elderly patients with glioblastoma in the United States, 2000–2017. BMC Cancer 21(1):54
Deng Y, Zhu H, Xiao L, Liu C, Meng X (2020) Circ_0005198 enhances temozolomide resistance of glioma cells through miR-198/TRIM14 axis. Aging 13(2):2198–2211
Feng S, Cai X, Li Y, Jian X, Zhang L, Li B (2019) Tripartite motif-containing 14 (TRIM14) promotes epithelial-mesenchymal transition via ZEB2 in glioblastoma cells. J Exp Clin Cancer Res 38(1):57
Ghasabi M, Mansoori B, Mohammadi A, Duijf PH, Shomali N, Shirafkan N et al (2019) MicroRNAs in cancer drug resistance: basic evidence and clinical applications. J Cell Physiol 234(3):2152–2168
Glenfield C, McLysaght A (2018) Pseudogenes provide evolutionary evidence for the competitive endogenous RNA hypothesis. Mol Biol Evol 35(12):2886–2899
Goodall GJ, Wickramasinghe VO (2021) RNA in cancer. Nat Rev Cancer 21(1):22–36
Grillone K, Riillo C, Scionti F, Rocca R, Tradigo G, Guzzi PH et al (2020) Non-coding RNAs in cancer: platforms and strategies for investigating the genomic “dark matter”. J Exp Clin Cancer Res 39(1):117
Guo X, Piao H (2021) Research Progress of circRNAs in Glioblastoma. Front Cell Dev Biol 9:791892
Gusyatiner O, Hegi ME (2018) Glioma epigenetics: from subclassification to novel treatment options. Semin Cancer Biol 51:50–58
Hallal S, Mallawaaratchy DM, Wei H, Ebrahimkhani S, Stringer BW, Day BW et al (2019) Extracellular vesicles released by Glioblastoma cells stimulate normal astrocytes to acquire a tumor-supportive phenotype Via p53 and MYC signaling pathways. Mol Neurobiol 56(6):4566–4581
Hansen TB, Jensen TI, Clausen BH, Bramsen JB, Finsen B, Damgaard CK et al (2013) Natural RNA circles function as efficient microRNA sponges. Nature 495(7441):384–388
He C, Wang X, Du M, Dong Y (2021) LncRNA MSC-AS1 Promotes Colorectal Cancer Progression by Regulating miR-325/TRIM14 Axis. J Oncol 2021:9954214
Hu G, Pen W, Wang M (2019) TRIM14 promotes breast Cancer cell proliferation by inhibiting apoptosis. Oncol Res 27(4):439–447
Jiang Y, Zhao J, Xu J, Zhang H, Zhou J, Li H et al (2022) Glioblastoma-associated microglia-derived exosomal circKIF18A promotes angiogenesis by targeting FOXC2. Oncogene 41(26):3461–3473
Jones PA, Issa JP, Baylin S (2016) Targeting the cancer epigenome for therapy. Nat Rev Genet 17(10):630–641
Khathayer F, Taylor MA, Ray SK (2020) Synergism of 4HPR and SAHA increases anti-tumor actions in glioblastoma cells. Apoptosis 25(3–4):217–232
Kristensen LS, Andersen MS, Stagsted LVW, Ebbesen KK, Hansen TB, Kjems J (2019) The biogenesis, biology and characterization of circular RNAs. Nat Rev Genet 20(11):675–691
Kristensen LS, Jakobsen T, Hager H, Kjems J (2022) The emerging roles of circRNAs in cancer and oncology. Nat Rev Clin Oncol 19(3):188–206
Kunadis E, Lakiotaki E, Korkolopoulou P, Piperi C (2021) Targeting post-translational histone modifying enzymes in glioblastoma. Pharmacol Ther 220:107721
Li Y, Seto E (2016) HDACs and HDAC inhibitors in Cancer Development and Therapy. Cold Spring Harb Perspect Med 6(10):a026831
Li X, Yang L, Chen LL (2018) The Biogenesis, Functions, and Challenges of Circular RNAs. Mol Cell 71(3):428–442
Li S, Teng S, Xu J, Su G, Zhang Y, Zhao J et al (2019) Microarray is an efficient tool for circRNA profiling. Brief Bioinform 20(4):1420–1433
Li X, Wang N, Leng H, Yuan H, Xu L (2022) Hsa_circ_0043949 reinforces temozolomide resistance via upregulating oncogene ITGA1 axis in glioblastoma. Metab Brain Dis 37(8):2979–2993
Lopez-Jimenez E, Rojas AM, Andres-Leon E (2018) RNA sequencing and prediction tools for circular RNAs analysis. Adv Exp Med Biol 1087:17–33
Lv X, Wang M, Qiang J, Guo S (2019) Circular RNA circ-PITX1 promotes the progression of glioblastoma by acting as a competing endogenous RNA to regulate miR-379-5p/MAP3K2 axis. Eur J Pharmacol 863:172643
Ostrom QT, Gittleman H, Farah P, Ondracek A, Chen Y, Wolinsky Y et al (2013) CBTRUS statistical report: primary brain and central nervous system tumors diagnosed in the United States in 2006–2010. Neuro Oncol 15(Suppl 2):ii1–56
Panda AC (2018) Circular RNAs act as miRNA sponges. Adv Exp Med Biol 1087:67–79
Pastorino O, Gentile MT, Mancini A, Del Gaudio N, Di Costanzo A, Bajetto A et al (2019) Histone deacetylase inhibitors impair vasculogenic mimicry from Glioblastoma cells. Cancers (Basel) 11(6):747
Rajaratnam V, Islam MM, Yang M, Slaby R, Ramirez HM, Mirza SP (2020) Glioblastoma: pathogenesis and current status of Chemotherapy and other Novel treatments. Cancers (Basel) 12(4):937
Ramaiah MJ, Tangutur AD, Manyam RR (2021) Epigenetic modulation and understanding of HDAC inhibitors in cancer therapy. Life Sci 277:119504
Rasmussen RD, Gajjar MK, Jensen KE, Hamerlik P (2016) Enhanced efficacy of combined HDAC and PARP targeting in glioblastoma. Mol Oncol 10(5):751–763
Singh D, Assaraf YG, Gacche RN (2022) Long non-coding RNA mediated drug resistance in breast cancer. Drug Resist Updat 63:100851
Tan Z, Song L, Wu W, Zhou Y, Zhu J, Wu G et al (2018) TRIM14 promotes chemoresistance in gliomas by activating Wnt/beta-catenin signaling via stabilizing Dvl2. Oncogene 37(40):5403–5415
Toden S, Zumwalt TJ, Goel A (2021) Non-coding RNAs and potential therapeutic targeting in cancer. Biochim Biophys Acta Rev Cancer 1875(1):188491
Touat M, Idbaih A, Sanson M, Ligon KL (2017) Glioblastoma targeted therapy: updated approaches from recent biological insights. Ann Oncol 28(7):1457–1472
Uddin MS, Mamun AA, Alghamdi BS, Tewari D, Jeandet P, Sarwar MS et al (2022) Epigenetics of glioblastoma multiforme: from molecular mechanisms to therapeutic approaches. Semin Cancer Biol 83:100–120
Wang C, Liu WR, Tan S, Zhou JK, Xu X, Ming Y et al (2022) Characterization of distinct circular RNA signatures in solid tumors. Mol Cancer 21(1):63
Wei JW, Huang K, Yang C, Kang CS (2017) Non-coding RNAs as regulators in epigenetics (review). Oncol Rep 37(1):3–9
West AC, Johnstone RW (2014) New and emerging HDAC inhibitors for cancer treatment. J Clin Invest 124(1):30–39
Wirsching HG, Galanis E, Weller M (2016) Glioblastoma. Handb Clin Neurol 134:381–397
Xue F, Cheng Y, Xu L, Tian C, Jiao H, Wang R et al (2020) LncRNA NEAT1/miR-129/Bcl-2 signaling axis contributes to HDAC inhibitor tolerance in nasopharyngeal cancer. Aging 12(14):14174–14188
Yamamoto K, Seike M, Takeuchi S, Soeno C, Miyanaga A, Noro R et al (2014) MiR-379/411 cluster regulates IL-18 and contributes to drug resistance in malignant pleural mesothelioma. Oncol Rep 32(6):2365–2372
Zhang HD, Jiang LH, Sun DW, Hou JC, Ji ZL (2018a) CircRNA: a novel type of biomarker for cancer. Breast Cancer 25(1):1–7
Zhang Z, Yang T, Xiao J (2018b) Circular RNAs: promising biomarkers for Human Diseases. EBioMedicine 34:267–274
Zhang C, Zhou Y, Gao Y, Zhu Z, Zeng X, Liang W et al (2022) Radiated glioblastoma cell-derived exosomal circ_0012381 induce M2 polarization of microglia to promote the growth of glioblastoma by CCL2/CCR2 axis. J Transl Med 20(1):388
Acknowledgements
None.
Funding
This study was supported by Basic Research project of Knowledge Innovation Plan of Wuhan Science and Technology Bureau (Grant Agreement number: 2022020801010549) and The Seventh Batch of Wuhan Young and Middle-aged Medical Backbone Talents Project.
Author information
Authors and Affiliations
Contributions
J.Y. designed the research study. L.M. and Y.W. performed the research. Q.T. collected and analyzed the data. Y.Z. contributed essential reagents or tools. X.D. operated the software and wrote the manuscript. All authors have read and approved the final manuscript.
Corresponding author
Ethics declarations
Ethics approval and consent to participate
This project was approved by the Ethics Committee of Wuhan Third Hospital.
Approval to perform this animal experiment was acquired from the Wuhan Third Hospital.
Consent for publication
Not applicable.
Conflict of interest
The authors declare that they have no conflicts of interest.
Additional information
Publisher’s Note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Electronic supplementary material
Below is the link to the electronic supplementary material.
Rights and permissions
Springer Nature or its licensor (e.g. a society or other partner) holds exclusive rights to this article under a publishing agreement with the author(s) or other rightsholder(s); author self-archiving of the accepted manuscript version of this article is solely governed by the terms of such publishing agreement and applicable law.
About this article
Cite this article
Meng, L., Wang, Y., Tu, Q. et al. Circular RNA circ_0000741/miR-379-5p/TRIM14 signaling axis promotes HDAC inhibitor (SAHA) tolerance in glioblastoma. Metab Brain Dis 38, 1351–1364 (2023). https://doi.org/10.1007/s11011-023-01184-9
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s11011-023-01184-9