Abstract
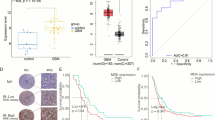
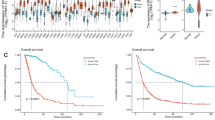
Gliomas are the most prevalent type of primary brain tumor, with poor prognosis reported in patients with high-grade glioma. Kinesin family member 4 A (KIF4A) stimulates the proliferation, migration, and invasion of tumor cells. However, its function in gliomas has not been clearly established. Therefore, this study aimed to investigate the effects of KIF4A on the epithelial–mesenchymal transition and invasion of glioma cells. We searched The Cancer Genome Atlas and Chinese Glioma Genome Atlas databases to identify KIF4A-related signaling pathways and downstream genes. We further validated them using western blotting, transwell migration and invasion, wound-healing scratch, and dual-luciferase reporter assays in U251 and U87 human glioblastoma cells. Our analysis of the Cancer Genome Atlas and Chinese Glioma Genome Atlas data showed elevated KIF4A expression in patients with gliomas and was associated with clinical grade. Here, KIF4A overexpression promoted the migration, invasion, and proliferation of glioma cells, whereas KIF4A knockdown showed contrasting results. Gene Ontology (GO) and Gene Set Enrichment Analysis (GSEA) analyses demonstrated that KIF4A positively controls TGF-β/SMAD signaling in glioma cells. Additionally, genetic correlation analysis revealed that KIF4A transcriptionally controls benzimidazoles-1 expression in glioma cells. KIF4A promotes the epithelial–mesenchymal transition by regulating the TGF-β/SMAD signaling pathway via benzimidazoles-1 in glioma cells.







Similar content being viewed by others
Data availability
The datasets generated during and/or analyzed during the current study are available from the corresponding author upon reasonable request, the TCGA database (https://portal.gdc.cancer.gov/), and the CGGA database (https://www.cgga.org.cn).
References
Lara-Velazquez M, Al-Kharboosh R, Jeanneret S, Vazquez-Ramos C, Mahato D, Tavanaiepour D, Rahmathulla G, Quinones-Hinojosa A (2017) Advances in brain tumor surgery for glioblastoma in adults. Brain Sci 7:166. https://doi.org/10.3390/brainsci7120166
Wen PY, Kesari S (2008) Malignant gliomas in adults. N Engl J Med 359:492–507. https://doi.org/10.1056/NEJMra0708126
Eisemann T, Costa B, Strelau J, Mittelbronn M, Angel P, Peterziel H (2018) An advanced glioma cell invasion assay based on organotypic brain slice cultures. BMC Cancer 18:103. https://doi.org/10.1186/s12885-018-4007-4
Iwadate Y (2016) Epithelial-mesenchymal transition in glioblastoma progression. Oncol Lett 11:1615–1620. https://doi.org/10.3892/ol.2016.4113
Paw I, Carpenter RC, Watabe K, Debinski W, Lo HW (2015) Mechanisms regulating glioma invasion. Cancer Lett 362:1–7. https://doi.org/10.1016/j.canlet.2015.03.015
Lamouille S, Xu J, Derynck R (2014) Molecular mechanisms of epithelial-mesenchymal transition. Nat Rev Mol Cell Biol 15:178–196. https://doi.org/10.1038/nrm3758
Vale RD, Reese TS, Sheetz MP (1985) Identification of a novel force-generating protein, kinesin, involved in microtubule-based motility. Cell 42:39–50. https://doi.org/10.1016/s0092-8674(85)80099-4
Rath O, Kozielski F (2012) Kinesins and cancer. Nat Rev Cancer 12:527–539. https://doi.org/10.1038/nrc3310
Cao Q, Song Z, Ruan H, Wang C, Yang X, Bao L, Wang K, Cheng G, Xu T, Xiao W, Xiong Z, Liu D, Yang M, Zhou D, Yang H, Chen K, Zhang X (2020) Targeting the KIF4A/AR axis to reverse endocrine therapy resistance in castration-resistant prostate cancer. Clin Cancer Res 26:1516–1528. https://doi.org/10.1158/1078-0432.CCR-19-0396
Taniwaki M, Takano A, Ishikawa N, Yasui W, Inai K, Nishimura H, Tsuchiya E, Kohno N, Nakamura Y, Daigo Y (2007) Activation of KIF4A as a prognostic biomarker and therapeutic target for lung cancer. Clin Cancer Res 13:6624–6631. https://doi.org/10.1158/1078-0432.CCR-07-1328
Hu G, Yan Z, Zhang C, Cheng M, Yan Y, Wang Y, Deng L, Lu Q, Luo S (2019) FOXM1 promotes hepatocellular carcinoma progression by regulating KIF4A expression. J Exp Clin Cancer Res 38:188. https://doi.org/10.1186/s13046-019-1202-3
Cho SY, Kim S, Kim G, Singh P, Kim DW (2019) Integrative analysis of KIF4A, 9, 18A, and 23 and their clinical significance in low-grade glioma and glioblastoma. Sci Rep 9:4599. https://doi.org/10.1038/s41598-018-37622-3
Nyati S, Schinske-Sebolt K, Pitchiaya S, Chekhovskiy K, Chator A, Chaudhry N, Dosch J, Van Dort ME, Varambally S, Kumar-Sinha C, Nyati MK, Ray D, Walter NG, Yu H, Ross BD, Rehemtulla A (2015) The kinase activity of the Ser/Thr kinase BUB1 promotes TGF-β signaling. Sci Signal 8:ra1. https://doi.org/10.1126/scisignal.2005379
Chin L, Andersen JN, Futreal PA (2011) Cancer genomics: from discovery science to personalized medicine. Nat Med 17:297–303. https://doi.org/10.1038/nm.2323
The Gene Ontology Consortium (2019) The Gene Ontology Resource: 20 years and still GOing strong. Nucleic Acids Res 47:D330–D338. https://doi.org/10.1093/nar/gky1055
Subramanian, A., Tamayo, P., Mootha, V. K., Mukherjee, S., Ebert, B. L., Gillette, M. A., Paulovich, A., Pomeroy, S. L., Golub, T. R., Lander, E. S., & Mesirov, J. P. (2005). Gene set enrichment analysis: a knowledge-based approach for interpreting genome-wide expression profiles. Proc Nati Acad Sci United States America 102(43):15545–15550. https://doi.org/10.1073/pnas.0506580102
Wu T, Hu E, Xu S, Chen M, Guo P, Dai Z, Feng T, Zhou L, Tang W, Zhan L, Fu X, Liu S, Bo X, Yu G (2021) clusterProfiler 4.0: a universal enrichment tool for interpreting omics data. Innov (Camb) 2:100141. https://doi.org/10.1016/j.xinn.2021.100141
Eckel-Passow JE, Lachance DH, Molinaro AM, Walsh KM, Decker PA, Sicotte H, Pekmezci M, Rice T, Kosel ML, Smirnov IV, Sarkar G, Caron AA, Kollmeyer TM, Praska CE, Chada AR, Halder C, Hansen HM, McCoy LS, Bracci PM, Marshall R, Zheng S, Reis GF, Pico AR, O’Neill BP, Buckner JC, Giannini C, Huse JT, Perry A, Tihan T, Berger MS, Chang SM, Prados MD, Wiemels J, Wiencke JK, Wrensch MR, Jenkins RB (2015) Glioma groups based on 1p/19q, IDH, and tert promoter mutations in tumors. N Engl J Med 372:2499–2508. https://doi.org/10.1056/NEJMoa1407279
Han S, Liu Y, Cai SJ, Qian M, Ding J, Larion M, Gilbert MR, Yang C (2020) IDH mutation in glioma: molecular mechanisms and potential therapeutic targets. Br J Cancer 122(11):1580–1589. https://doi.org/10.1038/s41416-020-0814-x
Hersh DS, Harder BG, Roos A, Peng S, Heath JE, Legesse T, Kim AJ, Woodworth GF, Tran NL, Winkles JA (2018) The TNF receptor family member Fn14 is highly expressed in recurrent glioblastoma and in GBM patient-derived xenografts with acquired temozolomide resistance. Neurooncology 20(10):1321–1330. https://doi.org/10.1093/neuonc/noy063
Lapointe S, Perry A, Butowski NA (2018) Primary brain tumours in adults. Lancet (London England) 392(10145):432–446. https://doi.org/10.1016/S0140-6736(18)30990-5
Delgado-Martín B, Medina MÁ (2020) Advances in the knowledge of the Molecular Biology of Glioblastoma and its impact in patient diagnosis, stratification, and treatment. Advanced science (Weinheim. Baden-Wurttemberg Germany) 7(9):1902971. https://doi.org/10.1002/advs.201902971
Radisky DC, LaBarge MA (2008) Epithelial-mesenchymal transition and the stem cell phenotype. Cell Stem Cell 2(6):511–512. https://doi.org/10.1016/j.stem.2008.05.007
Pastushenko I, Blanpain C (2019) EMT Transition States during Tumor Progression and Metastasis. Trends Cell Biol 29(3):212–226. https://doi.org/10.1016/j.tcb.2018.12.001
Mittal V (2018) Epithelial mesenchymal transition in Tumor Metastasis. Annu Rev Pathol 13:395–412. https://doi.org/10.1146/annurev-pathol-020117-043854
Derynck R, Zhang YE (2003) Smad-dependent and smad-independent pathways in TGF-beta family signalling. Nature 425(6958):577–584. https://doi.org/10.1038/nature02006
David CJ, Massagué J (2018) Contextual determinants of TGFβ action in development, immunity and cancer. Nat Rev Mol Cell Biol 19(7):419–435. https://doi.org/10.1038/s41580-018-0007-0
Moon H, Han KH, Ro SW (2017) Pro-tumorigenic roles of TGF-β signaling during the early stages of liver tumorigenesis through upregulation of snail. BMB Rep 50(12):599–600. https://doi.org/10.5483/bmbrep.2017.50.12.201
Korkut A, Zaidi S, Kanchi RS, Rao S, Gough NR, Schultz A, Li X, Lorenzi PL, Berger AC, Robertson G, Kwong LN, Datto M, Roszik J, Ling S, Ravikumar V, Manyam G, Rao A, Shelley S, Liu Y, Ju Z, Akbani R (2018) A Pan-cancer Analysis reveals high-frequency genetic alterations in mediators of signaling by the TGF-β superfamily. Cell Syst 7(4):422–437e7. https://doi.org/10.1016/j.cels.2018.08.010
Shathasivam P, Kollara A, Ringuette MJ, Virtanen C, Wrana JL, Brown TJ (2015) Human ortholog of Drosophila Melted impedes SMAD2 release from TGF-β receptor I to inhibit TGF-β signaling. Proc Natl Acad Sci USA 112(23):E3000–E3009. https://doi.org/10.1073/pnas.1504671112
Crane CA, Han SJ, Barry JJ, Ahn BJ, Lanier LL, Parsa AT (2010) TGF-beta downregulates the activating receptor NKG2D on NK cells and CD8 + T cells in glioma patients. Neurooncology 12(1):7–13. https://doi.org/10.1093/neuonc/nop009
Luo D, Xu X, Li J, Chen C, Chen W, Wang F, Xie Y, Li F (2018) The PDK1/cJun pathway activated by TGFβ induces EMT and promotes proliferation and invasion in human glioblastoma. Int J Oncol 53(5):2067–2080. https://doi.org/10.3892/ijo.2018.4525
Rouam S, Moreau T, Broët P (2010) Identifying common prognostic factors in genomic cancer studies: a novel index for censored outcomes. BMC Bioinformatics 11:150. https://doi.org/10.1186/1471-2105-11-150
Cuijpers SAG, Willemstein E, Ruppert JG, van Elsland DM, Earnshaw WC, Vertegaal ACO (2020) Chromokinesin KIF4A teams up with stathmin 1 to regulate abscission in a SUMO-dependent manner. J Cell Sci 133:jcs248591. https://doi.org/10.1242/jcs.248591
Hou PF, Jiang T, Chen F, Shi PC, Li HQ, Bai J, Song J (2018) KIF4A facilitates cell proliferation via induction of p21-mediated cell cycle progression and promotes metastasis in colorectal cancer. Cell Death Dis 9:477. https://doi.org/10.1038/s41419-018-0550-9
Zhang H, Meng S, Chu K, Chu S, Fan YC, Bai J, Yu ZQ (2022) KIF4A drives gliomas growth by transcriptional repression of Rac1/Cdc42 to induce cytoskeletal remodeling in glioma cells. J Cancer 13:3640–3651. https://doi.org/10.7150/jca.77238
Kuburich NA, den Hollander P, Pietz JT, Mani SA (2022) Vimentin and cytokeratin: good alone, bad together. Sem Cancer Biol 86(Pt 3):816–826. https://doi.org/10.1016/j.semcancer.2021.12.006
Acknowledgements
We would like to thank Professor Hongtao Zhang for providing access to the Suzhou Key Laboratory for Molecular Cancer Genetics.
Funding
This work was supported by Soochow University Mentorship funding (Grant numbers 22122134 and 23122220).
Author information
Authors and Affiliations
Contributions
XL provided experimental funding. YX completed major experiments and wrote the main manuscript text. XL and JZ provided the laboratory methods and supervised the laboratory process. GW and LH assisted with some of the experiments. LZ and SM analyzed experimental data. All authors read and approved the final manuscript.
Corresponding authors
Ethics declarations
Competing interests
The authors declare no competing interests.
Ethical approval
Not applicable, as neither human nor animal experimentation was performed in this study.
Additional information
Publisher’s Note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary Information
Below is the link to the electronic supplementary material.
Rights and permissions
Springer Nature or its licensor (e.g. a society or other partner) holds exclusive rights to this article under a publishing agreement with the author(s) or other rightsholder(s); author self-archiving of the accepted manuscript version of this article is solely governed by the terms of such publishing agreement and applicable law.
About this article
Cite this article
Xu, Y., Xue, G., Zhou, L. et al. KIF4A promotes epithelial–mesenchymal transition by activating the TGF-β/SMAD signaling pathway in glioma cells. Mol Cell Biochem (2024). https://doi.org/10.1007/s11010-024-04943-z
Received:
Accepted:
Published:
DOI: https://doi.org/10.1007/s11010-024-04943-z