Abstract
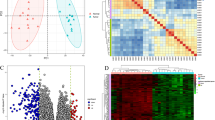
Gastric adenocarcinoma (GAC) is one of the world's most lethal malignant tumors. It has been established that the occurrence and progression of GAC are linked to molecular changes. However, the pathogenesis mechanism of GAC remains unclear. In this study, we sequenced 6 pairs of GAC tumor tissues and adjacent normal tissues and collected GAC gene expression profile data from the TCGA database. Analysis of this data revealed 465 differentially expressed genes (DEGs), of which 246 were upregulated and 219 were downregulated. Kyoto Encyclopedia of Genes and Genomes (KEGG) pathway analysis demonstrated that DEGs were observably enriched in ECM-receptor interaction, protein digestion and absorption, and gastric acid secretion pathways. Six key genes (MATN3, COL1A1, COL5A2, P4HA3, SERPINE1 and VCAN) associated with poor GAC prognosis were screened from the protein‒protein interaction (PPI) network by survival analysis, and P4HA3 and MATN3 have rarely been reported to be associated with GAC. We further analyzed the function of P4HA3 in the GAC cell line SGC-7901 by RT‒qPCR, MTT, flow cytometry, colony formation, wound healing, Transwell and western blot assays. We found that P4HA3 was upregulated in the SGC-7901 cell line versus normal control cells. The outcomes of the loss-of-function assay illustrated that P4HA3 significantly enhanced the ability of GAC cells to proliferate and migrate. This study provides a new basis for the selection of prognostic markers and therapeutic targets for GAC.









Similar content being viewed by others
Data availability
Enquiries about data availability should be directed to the authors.
References
Sung H, Ferlay J, Siegel RL, Laversanne M, Soerjomataram I, Jemal A, Bray F (2021) Global Cancer Statistics 2020: GLOBOCAN estimates of incidence and mortality worldwide for 36 cancers in 185 countries. CA Cancer J Clin 71:209–249. https://doi.org/10.3322/caac.21660
Fuse N, Bando H, Chin K, Ito S, Yoshikawa T, Tsuburaya A, Terashima M, Kawashima Y, Fukunaga T, Gotoh M, Emi Y, Yoshida K, Oki E, Takahashi S, Kuriki H, Sato K, Sasako M (2017) Adjuvant capecitabine plus oxaliplatin after D2 gastrectomy in Japanese patients with gastric cancer: a phase II study. Gastric Cancer 20:332–340. https://doi.org/10.1007/s10120-016-0606-4
Ji XK, Madhurapantula SV, He G, Wang KY, Song CH, Zhang JY, Wang KJ (2021) Genetic variant of cyclooxygenase-2 in gastric cancer: More inflammation and susceptibility. World J Gastroenterol 27:4653–4666. https://doi.org/10.3748/wjg.v27.i28.4653
Wang H, Huang D, Zhou Z (2021) Effect of NTSR1 on the proliferation and invasion of gastric adenocarcinoma cells, the underlying mechanism, and clinical role. Ann Clin Lab Sci 51:163–173
Cikutovic-Molina R, Herrada AA, Gonzalez W, Brown N, Zuniga L (2019) TASK-3 gene knockdown dampens invasion and migration and promotes apoptosis in KATO III and MKN-45 human gastric adenocarcinoma cell lines. Int J Mol Sci. https://doi.org/10.3390/ijms20236077
Wu X, Chen Y, Li G, Xia L, Gu R, Wen X, Ming X, Chen H (2014) Her3 is associated with poor survival of gastric adenocarcinoma: Her3 promotes proliferation, survival and migration of human gastric cancer mediated by PI3K/AKT signaling pathway. Med Oncol 31:903. https://doi.org/10.1007/s12032-014-0903-x
Xie X, Zhou Z, Song Y, Zhang X, Dang C, Zhang H (2021) Mist1 inhibits epithelial-mesenchymal transition in gastric adenocarcinoma via downregulating the Wnt/beta-catenin pathway. J Cancer 12:4574–4584. https://doi.org/10.7150/jca.59138
Bayoumi AS, Sayed A, Broskova Z, Teoh JP, Wilson J, Su H, Tang YL, Kim IM (2016) Crosstalk between long noncoding RNAs and MicroRNAs in health and disease. Int J Mol Sci 17:356. https://doi.org/10.3390/ijms17030356
Zhou K, Cao D, Wang Y, Wang L, Meng X (2021) Hsa-miR-30a-3p attenuates gastric adenocarcinoma proliferation and metastasis via APBB2. Aging (Albany NY) 13:16763–16772. https://doi.org/10.18632/aging.203197
Dhakras P, Uboha N, Horner V, Reinig E, Matkowskyj KA (2020) Gastrointestinal cancers: current biomarkers in esophageal and gastric adenocarcinoma. Transl Gastroenterol Hepatol 5:55. https://doi.org/10.21037/tgh.2020.01.08
Mardis ER (2008) The impact of next-generation sequencing technology on genetics. Trends Genet 24:133–141. https://doi.org/10.1016/j.tig.2007.12.007
Bolger AM, Lohse M, Usadel B (2014) Trimmomatic: a flexible trimmer for Illumina sequence data. Bioinformatics 30:2114–2120. https://doi.org/10.1093/bioinformatics/btu170
Kim D, Pertea G, Trapnell C, Pimentel H, Kelley R, Salzberg SL (2013) TopHat2: accurate alignment of transcriptomes in the presence of insertions, deletions and gene fusions. Genome Biol 14:R36. https://doi.org/10.1186/gb-2013-14-4-r36
Li M, Bai M, Wu Y, Shao W, Zheng L, Sun L, Wang S, Yu C, Huang Y (2021) AGTAR: A novel approach for transcriptome assembly and abundance estimation using an adapted genetic algorithm from RNA-seq data. Comput Biol Med 135:104646. https://doi.org/10.1016/j.compbiomed.2021.104646
Pertea M, Pertea GM, Antonescu CM, Chang TC, Mendell JT, Salzberg SL (2015) StringTie enables improved reconstruction of a transcriptome from RNA-seq reads. Nat Biotechnol 33:290–295. https://doi.org/10.1038/nbt.3122
Love MI, Huber W, Anders S (2014) Moderated estimation of fold change and dispersion for RNA-seq data with DESeq2. Genome Biol 15:550. https://doi.org/10.1186/s13059-014-0550-8
Ginestet C (2011) ggplot2: elegant graphics for data analysis. J R Stat Soc a Stat 174:245–245. https://doi.org/10.1111/j.1467-985X.2010.00676_9.x
Yu G, Wang LG, Han Y, He QY (2012) clusterProfiler: an R package for comparing biological themes among gene clusters. OMICS 16:284–287. https://doi.org/10.1089/omi.2011.0118
Ashburner M, Ball CA, Blake JA, Botstein D, Butler H, Cherry JM, Davis AP, Dolinski K, Dwight SS, Eppig JT, Harris MA, Hill DP, Issel-Tarver L, Kasarskis A, Lewis S, Matese JC, Richardson JE, Ringwald M, Rubin GM, Sherlock G (2000) Gene ontology: tool for the unification of biology. The Gene Ontology Consortium. Nat Genet 25:25–29. https://doi.org/10.1038/75556
Kanehisa M, Goto S (2000) KEGG: kyoto encyclopedia of genes and genomes. Nucleic Acids Res 28:27–30. https://doi.org/10.1093/nar/28.1.27
Szklarczyk D, Gable AL, Lyon D, Junge A, Wyder S, Huerta-Cepas J, Simonovic M, Doncheva NT, Morris JH, Bork P, Jensen LJ, Mering CV (2019) STRING v11: protein-protein association networks with increased coverage, supporting functional discovery in genome-wide experimental datasets. Nucleic Acids Res 47:D607–D613. https://doi.org/10.1093/nar/gky1131
Kohl M, Wiese S, Warscheid B (2011) Cytoscape: software for visualization and analysis of biological networks. Methods Mol Biol 696:291–303. https://doi.org/10.1007/978-1-60761-987-1_18
Li C, Tang Z, Zhang W, Ye Z, Liu F (2021) GEPIA2021: integrating multiple deconvolution-based analysis into GEPIA. Nucleic Acids Res 49:W242–W246. https://doi.org/10.1093/nar/gkab418
Ponten F, Jirstrom K, Uhlen M (2008) The human protein atlas—a tool for pathology. J Pathol 216:387–393. https://doi.org/10.1002/path.2440
Li T, Fu J, Zeng Z, Cohen D, Li J, Chen Q, Li B, Liu XS (2020) TIMER2.0 for analysis of tumor-infiltrating immune cells. Nucleic Acids Res 48:W509–W514. https://doi.org/10.1093/nar/gkaa407
Suh JH, Won KY, Kim GY, Bae GE, Lim SJ, Sung JY, Park YK, Kim YW, Lee J (2015) Expression of tumoral FOXP3 in gastric adenocarcinoma is associated with favorable clinicopathological variables and related with Hippo pathway. Int J Clin Exp Pathol 8:14608–14618
Vidal AF, Cruz AM, Magalhaes L, Pereira AL, Anaissi AK, Alves NC, Albuquerque PJ, Burbano RM, Demachki S, Ribeiro-dos-Santos A (2016) hsa-miR-29c and hsa-miR-135b differential expression as potential biomarker of gastric carcinogenesis. World J Gastroenterol 22:2060–2070. https://doi.org/10.3748/wjg.v22.i6.2060
Xu XW, Yang XM, Zhao WJ, Zhou L, Li DC, Zheng YH (2018) DNM1L, a key prognostic predictor for gastric adenocarcinoma, is involved in cell proliferation, invasion, and apoptosis. Oncol Lett 16:3635–3641. https://doi.org/10.3892/ol.2018.9138
Yan SM, He F, Luo RZ, Wu HN, Huang MY, Huang CY, Li Y, Zhou ZW (2016) Decreased expression of BRCA1-associated protein 1 predicts unfavorable survival in gastric adenocarcinoma. Tumor Biol 37:6125–6133. https://doi.org/10.1007/s13277-015-3983-0
Sun QL, Lin P, Zhang JY, Li XY, Yang LG, Huang JG, Zhou ZJ, Liu P, Liu NQ (2015) Expression of fibroblast growth factor 10 is correlated with poor prognosis in gastric adenocarcinoma. Tohoku J Exp Med 236:311–318. https://doi.org/10.1620/tjem.236.311
Takahashi K, Kaira K, Shimizu A, Sato T, Takahashi N, Ogawa H, Yoshinari D, Yokobori T, Asao T, Takeyoshi I, Oyama T (2016) Clinical significance of beta2-adrenergic receptor expression in patients with surgically resected gastric adenocarcinoma. Tumour Biol 37:13885–13892. https://doi.org/10.1007/s13277-016-5139-2
Borg D, Hedner C, Nodin B, Larsson A, Johnsson A, Eberhard J, Jirstrom K (2016) Expression of podocalyxin-like protein is an independent prognostic biomarker in resected esophageal and gastric adenocarcinoma. BMC Clin Pathol 16:13. https://doi.org/10.1186/s12907-016-0034-8
Wu Q, Li X, Yang H, Lu C, You J, Zhang Z (2014) Extracellular matrix protein 1 is correlated to carcinogenesis and lymphatic metastasis of human gastric cancer. World J Surg Oncol 12:132. https://doi.org/10.1186/1477-7819-12-132
Smolka AJ, Schubert ML (2017) Helicobacter pylori-induced changes in gastric acid secretion and upper gastrointestinal disease. Curr Top Microbiol Immunol 400:227–252. https://doi.org/10.1007/978-3-319-50520-6_10
Li Y, Sun R, Zhao X, Sun B (2021) RUNX2 promotes malignant progression in gastric cancer by regulating COL1A1. Cancer Biomark 31:227–238. https://doi.org/10.3233/CBM-200472
Wang Q, Yu J (2018) MiR-129-5p suppresses gastric cancer cell invasion and proliferation by inhibiting COL1A1. Biochem Cell Biol 96:19–25. https://doi.org/10.1139/bcb-2016-0254
Tan Y, Chen Q, Xing Y, Zhang C, Pan S, An W, Xu H (2021) High expression of COL5A2, a member of COL5 family, indicates the poor survival and facilitates cell migration in gastric cancer. Biosci Rep. https://doi.org/10.1042/BSR20204293
Yang JD, Ma L, Zhu Z (2019) SERPINE1 as a cancer-promoting gene in gastric adenocarcinoma: facilitates tumour cell proliferation, migration, and invasion by regulating EMT. J Chemother 31:408–418. https://doi.org/10.1080/1120009X.2019.1687996
Cheng Y, Sun H, Wu L, Wu F, Tang W, Wang X, Lv C (2020) VUp-regulation of VCAN promotes the proliferation, invasion and migration and serves as a biomarker in gastric cancer. Onco Targets Ther 13:8665–8675. https://doi.org/10.2147/OTT.S262613
Klatt AR, Nitsche DP, Kobbe B, Morgelin M, Paulsson M, Wagener R (2000) Molecular structure and tissue distribution of matrilin-3, a filament-forming extracellular matrix protein expressed during skeletal development. J Biol Chem 275:3999–4006. https://doi.org/10.1074/jbc.275.6.3999
Chapman KL, Mortier GR, Chapman K, Loughlin J, Grant ME, Briggs MD (2001) Mutations in the region encoding the von Willebrand factor A domain of matrilin-3 are associated with multiple epiphyseal dysplasia. Nat Genet 28:393–396. https://doi.org/10.1038/ng573
Song H, Liu L, Song Z, Ren Y, Li C, Huo J (2018) P4HA3 is epigenetically activated by slug in gastric cancer and its deregulation is associated with enhanced metastasis and poor survival. Technol Cancer Res Treat 17:1533033818796485. https://doi.org/10.1177/1533033818796485
Wang T, Wang YX, Dong YQ, Yu YL, Ma K (2020) Prolyl 4-hydroxylase subunit alpha 3 presents a cancer promotive function in head and neck squamous cell carcinoma via regulating epithelial-mesenchymal transition. Arch Oral Biol 113:104711. https://doi.org/10.1016/j.archoralbio.2020.104711
Nakasuka F, Tabata S, Sakamoto T, Hirayama A, Ebi H, Yamada T, Umetsu K, Ohishi M, Ueno A, Goto H, Sugimoto M, Nishioka Y, Yamada Y, Tomita M, Sasaki AT, Yano S, Soga T (2021) TGF-beta-dependent reprogramming of amino acid metabolism induces epithelial-mesenchymal transition in non-small cell lung cancers. Commun Biol 4:782. https://doi.org/10.1038/s42003-021-02323-7
Funding
This work was supported by the Natural Nature Science Foundation of China (81700709), Research Foundation of Jilin Provincial Science & Technology Development (20180520105JH), Systems Biology Research on Genome and Transcriptome of Stem Cells (2017030) of Jilin Province Sunbird Regenerative Medical Engineering Co., Ltd.
Author information
Authors and Affiliations
Contributions
Concept and design: ML, LS, CY, YH. Bioinformatics analysis: ML, MB. Biological assays: ML, MB, YW, SY, LZ. Drafting, revision and final approval of the manuscript: ML, MB, YW, SY, LZ, LS, CY, YH.
Corresponding author
Ethics declarations
Conflict of interest
The authors declare that there is no conflict of interest to indicate.
Additional information
Publisher's Note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary Information
Below is the link to the electronic supplementary material.
Rights and permissions
Springer Nature or its licensor (e.g. a society or other partner) holds exclusive rights to this article under a publishing agreement with the author(s) or other rightsholder(s); author self-archiving of the accepted manuscript version of this article is solely governed by the terms of such publishing agreement and applicable law.
About this article
Cite this article
Li, M., Bai, M., Wu, Y. et al. Transcriptome sequencing identifies prognostic genes involved in gastric adenocarcinoma. Mol Cell Biochem 478, 2891–2906 (2023). https://doi.org/10.1007/s11010-023-04705-3
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s11010-023-04705-3