Abstract
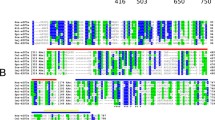
Translation initiation is the first step in three essential processes leading to protein synthesis. It is carried out by proteins called translation initiation factors and ribosomes on the mRNA. One of the critical translation initiation factors in eukaryotes is eIF4G which is a scaffold protein that helps assemble translation initiation complexes that carry out translation initiation which ultimately leads to polypeptide synthesis. Trypanosomatids are a large family of kinetoplastids, some of which are protozoan parasites that cause diseases in humans through transmission by vectors. While the protein translation mechanisms in eukaryotes and prokaryotes are well understood, the protein translation factors and mechanisms in trypanosomatids are poorly understood necessitating further studies. Unlike other eukaryotes, trypanosomatids contain five eIF4G orthologues with diversity in length and sequences. Here, I have used bioinformatics tools to look at trypanosomatid keIF4G orthologue sequences and report that there are similarities and considerable differences in their domains/motifs organization and signature amino acid sequences that are required for different functions as compared to human eIF4G. My analysis suggests that there is likely to be considerable diversity and complexity in trypanosomatid keIF4G functions as compared to other eukaryotes.








Similar content being viewed by others
Data availability
All data and material are available.
Code availability
Not applicable.
References
Ponting CP (2000) Novel eIF4G domain homologues linking mRNA translation with nonsense-mediated mRNA decay. Trends Biochem Sci 25:423–426
Gingras AC, Raught B, Sonenberg N (2001) Regulation of translation initiation by FRAP/mTOR. Genes Dev 15:807–826
Osborne MJ, Borden KL (2015) The eukaryotic translation initiation factor eIF4E in the nucleus: taking the road less traveled. Immunol Rev 263:210–223
Gruner S, Peter D, Weber R, Wohlbold L, Chung MY et al (2016) The structures of eIF4E-eIF4G complexes reveal an extended interface to regulate translation initiation. Mol Cell 64:467–479
Miras M, Truniger V, Silva C, Verdaguer N, Aranda MA et al (2017) Structure of eIF4E in complex with an eIF4G peptide supports a universal bipartite binding mode for protein translation. Plant Physiol 174:1476–1491
Stuart K, Brun R, Croft S, Fairlamb A, Gurtler RE et al (2008) Kinetoplastids: related protozoan pathogens, different diseases. J Clin Investig 118:1301–1310
Votypka J, Sukova E, Kraeva N, Ishemgulova A, Duzi I et al (2013) Diversity of trypanosomatids (Kinetoplastea: Trypanosomatidae) parasitizing fleas (Insecta: Siphonaptera) and description of a new genus Blechomonas gen. n. Protist 164:763–781
Bhattacharya A, Corbeil A, do Monte-Neto RL, Fernandez-Prada C (2020) Of drugs and trypanosomatids: new tools and knowledge to reduce bottlenecks in drug discovery. Genes (Basel) 11(7):722
Dhalia R, Reis CR, Freire ER, Rocha PO, Katz R et al (2005) Translation initiation in Leishmania major: characterisation of multiple eIF4F subunit homologues. Mol Biochem Parasitol 140:23–41
Moura DM, Reis CR, Xavier CC, da Costa Lima TD, Lima RP et al (2015) Two related trypanosomatid eIF4G homologues have functional differences compatible with distinct roles during translation initiation. RNA Biol 12:305–319
Das S (2020) Taking a re-look at cap-binding signatures of the mRNA cap-binding protein eIF4E orthologues in trypanosomatids. Mol Cell Biochem 476(2):1037–1049
Yoffe Y, Leger M, Zinoviev A, Zuberek J, Darzynkiewicz E et al (2009) Evolutionary changes in the Leishmania eIF4F complex involve variations in the eIF4E-eIF4G interactions. Nucleic Acids Res 37:3243–3253
Zinoviev A, Leger M, Wagner G, Shapira M (2011) A novel 4E-interacting protein in Leishmania is involved in stage-specific translation pathways. Nucleic Acids Res 39:8404–8415
Zinoviev A, Manor S, Shapira M (2012) Nutritional stress affects an atypical cap-binding protein in Leishmania. RNA Biol 9:1450–1460
Shrivastava R, Drory-Retwitzer M, Shapira M (2019) Nutritional stress targets LeishIF4E-3 to storage granules that contain RNA and ribosome components in Leishmania. PLoS Negl Trop Dis 13:e0007237
Freire ER, Dhalia R, Moura DM, da Costa Lima TD, Lima RP et al (2011) The four trypanosomatid eIF4E homologues fall into two separate groups, with distinct features in primary sequence and biological properties. Mol Biochem Parasitol 176:25–36
Freire ER, Vashisht AA, Malvezzi AM, Zuberek J, Langousis G et al (2014) eIF4F-like complexes formed by cap-binding homolog TbEIF4E5 with TbEIF4G1 or TbEIF4G2 are implicated in post-transcriptional regulation in Trypanosoma brucei. RNA 20:1272–1286
Freire ER, Malvezzi AM, Vashisht AA, Zuberek J, Saada EA et al (2014) Trypanosoma brucei translation initiation factor homolog EIF4E6 forms a tripartite cytosolic complex with EIF4G5 and a capping enzyme homolog. Eukaryot Cell 13:896–908
Shrivastava R, Tupperwar N, Schwartz B, Baron N, Shapira M (2021) LeishIF4E-5 is a promastigote-specific cap-binding protein in Leishmania. Int J Mol Sci 22(8):3979
Pyronnet S, Imataka H, Gingras AC, Fukunaga R, Hunter T et al (1999) Human eukaryotic translation initiation factor 4G (eIF4G) recruits mnk1 to phosphorylate eIF4E. EMBO J 18:270–279
Aravind L, Koonin EV (2000) Eukaryote-specific domains in translation initiation factors: implications for translation regulation and evolution of the translation system. Genome Res 10:1172–1184
Dobrikov M, Dobrikova E, Shveygert M, Gromeier M (2011) Phosphorylation of eukaryotic translation initiation factor 4G1 (eIF4G1) by protein kinase C{alpha} regulates eIF4G1 binding to Mnk1. Mol Cell Biol 31:2947–2959
Volpon L, Osborne MJ, Topisirovic I, Siddiqui N, Borden KL (2006) Cap-free structure of eIF4E suggests a basis for conformational regulation by its ligands. EMBO J 25:5138–5149
Peter D, Igreja C, Weber R, Wohlbold L, Weiler C et al (2015) Molecular architecture of 4E-BP translational inhibitors bound to eIF4E. Mol Cell 57:1074–1087
Marcotrigiano J, Gingras AC, Sonenberg N, Burley SK (1999) Cap-dependent translation initiation in eukaryotes is regulated by a molecular mimic of eIF4G. Mol Cell 3:707–716
Niedzwiecka A, Marcotrigiano J, Stepinski J, Jankowska-Anyszka M, Wyslouch-Cieszynska A et al (2002) Biophysical studies of eIF4E cap-binding protein: recognition of mRNA 5′ cap structure and synthetic fragments of eIF4G and 4E-BP1 proteins. J Mol Biol 319:615–635
Ptushkina M, von der Haar T, Vasilescu S, Frank R, Birkenhager R et al (1998) Cooperative modulation by eIF4G of eIF4E-binding to the mRNA 5′ cap in yeast involves a site partially shared by p20. EMBO J 17:4798–4808
Tomoo K, Matsushita Y, Fujisaki H, Abiko F, Shen X et al (2005) Structural basis for mRNA cap-binding regulation of eukaryotic initiation factor 4E by 4E-binding protein, studied by spectroscopic, X-ray crystal structural, and molecular dynamics simulation methods. Biochim Biophys Acta 1753:191–208
Siddiqui N, Tempel W, Nedyalkova L, Volpon L, Wernimont AK et al (2012) Structural insights into the allosteric effects of 4EBP1 on the eukaryotic translation initiation factor eIF4E. J Mol Biol 415:781–792
Imataka H, Sonenberg N (1997) Human eukaryotic translation initiation factor 4G (eIF4G) possesses two separate and independent binding sites for eIF4A. Mol Cell Biol 17:6940–6947
Morino S, Imataka H, Svitkin YV, Pestova TV, Sonenberg N (2000) Eukaryotic translation initiation factor 4E (eIF4E) binding site and the middle one-third of eIF4GI constitute the core domain for cap-dependent translation, and the C-terminal one-third functions as a modulatory region. Mol Cell Biol 20:468–477
da Costa Lima TD, Moura DM, Reis CR, Vasconcelos JR, Ellis L et al (2010) Functional characterization of three Leishmania poly(a) binding protein homologues with distinct binding properties to RNA and protein partners. Eukaryot Cell 9:1484–1494
Safaee N, Kozlov G, Noronha AM, Xie J, Wilds CJ et al (2012) Interdomain allostery promotes assembly of the poly(A) mRNA complex with PABP and eIF4G. Mol Cell 48:375–386
Cheng S, Liu R, Gallie DR (2013) The unique evolution of the programmed cell death 4 protein in plants. BMC Evol Biol 13:199
Dos Santos Rodrigues FH, Firczuk H, Breeze AL, Cameron AD, Walko M et al (2019) The Leishmania PABP1-eIF4E4 interface: a novel 5′–3′ interaction architecture for trans-spliced mRNAs. Nucleic Acids Res 47:1493–1504
LeFebvre AK, Korneeva NL, Trutschl M, Cvek U, Duzan RD et al (2006) Translation initiation factor eIF4G-1 binds to eIF3 through the eIF3e subunit. J Biol Chem 281:22917–22932
Jivotovskaya AV, Valasek L, Hinnebusch AG, Nielsen KH (2006) Eukaryotic translation initiation factor 3 (eIF3) and eIF2 can promote mRNA binding to 40S subunits independently of eIF4G in yeast. Mol Cell Biol 26:1355–1372
Meleppattu S, Kamus-Elimeleh D, Zinoviev A, Cohen-Mor S, Orr I et al (2015) The eIF3 complex of Leishmania-subunit composition and mode of recruitment to different cap-binding complexes. Nucleic Acids Res 43:6222–6235
Hinton TM, Coldwell MJ, Carpenter GA, Morley SJ, Pain VM (2007) Functional analysis of individual binding activities of the scaffold protein eIF4G. J Biol Chem 282:1695–1708
Villa N, Do A, Hershey JW, Fraser CS (2013) Human eukaryotic initiation factor 4G (eIF4G) protein binds to eIF3c, -d, and -e to promote mRNA recruitment to the ribosome. J Biol Chem 288:32932–32940
Kippert F, Gerloff DL (2009) Highly sensitive detection of individual HEAT and ARM repeats with HHpred and COACH. PLoS ONE 4:e7148
Schumacher MA, Bush MJ, Bibb MJ, Ramos-Leon F, Chandra G et al (2018) The crystal structure of the RsbN-sigmaBldN complex from Streptomyces venezuelae defines a new structural class of anti-sigma factor. Nucleic Acids Res 46:7467–7468
Allemand F, Mathy N, Brechemier-Baey D, Condon C (2005) The 5S rRNA maturase, ribonuclease M5, is a Toprim domain family member. Nucleic Acids Res 33:4368–4376
Funding
This work was not funded.
Author information
Authors and Affiliations
Contributions
Conceptualization, data curation, formal analysis, methodology, writing, and reviewing of this manuscript was done by SD.
Corresponding author
Ethics declarations
Conflict of interest
The author declares that there are no conflicts of interest.
Ethical approval
Not applicable.
Informed consent
Not applicable.
Consent for publication
Not applicable.
Additional information
Publisher's Note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary Information
Below is the link to the electronic supplementary material.
Rights and permissions
About this article
Cite this article
Das, S. Analysis of domain organization and functional signatures of trypanosomatid keIF4Gs. Mol Cell Biochem 477, 2415–2431 (2022). https://doi.org/10.1007/s11010-022-04464-7
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s11010-022-04464-7