Abstract
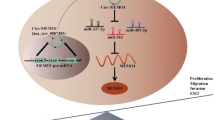
Colorectal cancer (CRC) remains a malignancy tumor with high metastasis and poor prognosis. We aimed to explore the effect of circular RNA (circRNA) hsa_circ_0006732 in the progression of CRC. Hsa_circ_0006732 expression in CRC tissues and cell lines were detected using qRT-PCR. The relationship between hsa_circ_0006732 expression and clinicopathologic characteristics of patients with CRC was analyzed. Loss-of-function assay was conducted to determine the regulatory effect of hsa_circ_0006732 on CRC cell proliferation, migration and invasion by using the CCK-8, wound-healing assay and transwell assays. Protein expression changes on epithelial mesenchymal transition (EMT)-related factors were detected by western blotting. The downstream signaling pathway was investigated by bioinformatics, dual-luciferase reporter assay. Rescue assay was further examined for prediction validation. It was found that hsa_circ_0006732 was highly expressed in CRC tissues and cell lines. Downregulation of hsa_circ_0006732 suppressed the proliferation, migration, invasion and EMT of CRC cells. Further mechanistic investigations proved that hsa_circ_0006732 functioned as a competitive endogenous RNA (ceRNA) by directly sponging of miR-127-3p, which further affected the expression of Ras-related protein Rab-3D (Rab3D). Taken together, these findings indicated that hsa_circ_0006732 might be an oncogene in CRC through the regulation of the miR-127-5p/RAB3D axis. Thus, hsa_circ_0006732 might serve as a potential therapeutic target for the treatment of CRC.





Similar content being viewed by others
Availability of data and materials
All data generated or analyzed during this study are included in this published article.
References
Mármol I, Sánchez-de-Diego C, Pradilla Dieste A, Cerrada E, Rodriguez Yoldi MJ (2017) Colorectal carcinoma: a general overview and future perspectives in colorectal cancer. Int J Mol Sci 18:197
Cronin KA, Lake AJ, Scott S, Sherman RL, Noone AM, Howlader N, Henley SJ, Anderson RN, Firth AU, Ma J (2018) Annual report to the nation on the status of cancer, part I: National cancer statistics. Cancer 124:2785–2800
Chen H, Li N, Ren J, Feng X, Lyu Z, Wei L, Li X, Guo L, Zheng Z, Zou S (2019) Participation and yield of a population-based colorectal cancer screening programme in China. Gut 68:1450–1457
Xie Y-H, Chen Y-X, Fang J-Y (2020) Comprehensive review of targeted therapy for colorectal cancer. Signal Transduct Target Ther 5:1–30
Siegel RL, Fedewa SA, Anderson WF, Miller KD, Ma J, Rosenberg PS, Jemal A (2017) Colorectal cancer incidence patterns in the United States. JNCI. https://doi.org/10.1093/jnci/djw322
Thanikachalam K, Khan G (2019) Colorectal cancer and nutrition. Nutrients 11:164. https://doi.org/10.3390/nu11010164
Vo JN, Cieslik M, Zhang Y, Shukla S, Xiao L, Zhang Y, Wu Y-M, Dhanasekaran SM, Engelke CG, Cao X (2019) The landscape of circular RNA in cancer. Cell 176(869–881):e13
Han B, Chao J, Yao H (2018) Circular RNA and its mechanisms in disease: from the bench to the clinic. Pharmacol Ther 187:31–44
Zeng X, Lin W, Guo M, Zou Q (2017) A comprehensive overview and evaluation of circular RNA detection tools. PLoS Comput Biol 13:e1005420
Hsiao K-Y, Sun HS, Tsai S-J (2017) Circular RNA–new member of noncoding RNA with novel functions. Exp Biol Med 242:1136–1141
Tian J, Xi X, Wang J, Yu J, Huang Q, Ma R, Zhang X, Li H, Wang L (2019) CircRNA hsa_circ_0004585 as a potential biomarker for colorectal cancer. Cancer Manag Res 11:5413
Yuan Y, Liu W, Zhang Y, Zhang Y, Sun S (2018) CircRNA circ_0026344 as a prognostic biomarker suppresses colorectal cancer progression via microRNA-21 and microRNA-31. Biochem Biophys Res Commun 503:870–875
Yang H, Li X, Meng Q, Sun H, Wu S, Hu W, Liu G, Li X, Yang Y, Chen R (2020) CircPTK2 (hsa_circ_0005273) as a novel therapeutic target for metastatic colorectal cancer. Mol Cancer 19:1–15
Zhou B, Zhang X, Li T, Xie R, Zhou J, Luo Y, Yang C (2020) CircZDHHC20 represses the proliferation, migration and invasion in trophoblast cells by miR-144/GRHL2 axis. Cancer Cell Int 20:1–11
Fearon ER, Vogelstein B (1990) A genetic model for colorectal tumorigenesis. Cell 61:759–767
Weinberg BA, Marshall JL, Salem ME (2017) The growing challenge of young adults with colorectal cancer. Oncology (Williston Park) 31:381–389
Rawla P, Sunkara T, Barsouk A (2019) Epidemiology of colorectal cancer: incidence, mortality, survival, and risk factors. Przeglad Gastroenterologiczny 14:89
Grasso CS, Giannakis M, Wells DK, Hamada T, Mu XJ, Quist M, Nowak JA, Nishihara R, Qian ZR, Inamura K (2018) Genetic mechanisms of immune evasion in colorectal cancer. Cancer Discov 8:730–749
Lei B, Tian Z, Fan W, Ni B (2019) Circular RNA: a novel biomarker and therapeutic target for human cancers. Int J Med Sci 16:292–301
Wang X, Ren Y, Ma S, Wang S (2020) Circular RNA 0060745, a novel circRNA, promotes colorectal cancer cell proliferation and metastasis through miR-4736 sponging. Oncol Targets Ther 13:1941
Tang J, Zhang C, Huang Y, Wang L, Xu Z, Zhang D, Zhang Y, Peng W, Feng Y, Sun Y (2021) CircRNA circ_0124554 blocked the ubiquitination of AKT promoting the skip lymph invasion on hepatic metastasis in colorectal cancer. Cell Death Dis 12:1–12
Yang J, Antin P, Berx G, Blanpain C, Brabletz T, Bronner M, Campbell K, Cano A, Casanova J, Christofori G (2020) Guidelines and definitions for research on epithelial–mesenchymal transition. Nat Rev Mol Cell Biol 21:341–352
Dongre A, Weinberg RA (2019) New insights into the mechanisms of epithelial–mesenchymal transition and implications for cancer. Nat Rev Mol Cell Biol 20:69–84
Zhang W, Zhu L, Yang G, Zhou B, Wang J, Qu X, Yan Z, Qian S, Liu R (2020) Hsa_circ_0026134 expression promoted TRIM25-and IGF2BP3-mediated hepatocellular carcinoma cell proliferation and invasion via sponging miR-127-5p. Biosci Rep. https://doi.org/10.1042/BSR20191418
Xiong G, Diao D, Lu D, Liu X, Liu Z, Mai S, Feng S, Dong X, Cai K (2020) Circular RNA circNELL2 Acts as the sponge of miR-127-5p to promote esophageal squamous cell carcinoma progression. Onco Targets Ther 13:9245
Chang S, Sun L, Feng G (2019) SP1-mediated long noncoding RNA POU3F3 accelerates the cervical cancer through miR-127-5p/FOXD1. Biomed Pharmacother 117:109133
Li L, Wan K, Xiong L, Liang S, Tou F, Guo S (2020) CircRNA hsa_circ_0087862 acts as an oncogene in non-small cell lung cancer by targeting miR-1253/RAB3D axis. Onco Targets Ther 13:2873
Jin T, Liu M, Liu Y, Li Y, Xu Z, He H, Liu J, Zhang Y, Ke Y (2020) Lcn2-derived circular RNA (hsa_circ_0088732) inhibits cell apoptosis and promotes EMT in glioma via the miR-661/RAB3D Axis. Front Oncol 10:170
Jiashi W, Chuang Q, Zhenjun Z, Guangbin W, Bin L, Ming H (2018) MicroRNA-506-3p inhibits osteosarcoma cell prolif and metastasis by suppressing RAB3D expression. Aging (Albany NY) 10:1294
Luo Y, Yu S-Y, Chen J-J, Qin J, Qiu Y-E, Zhong M, Chen M (2018) MiR-27b directly targets Rab3D to inhibit the malignant phenotype in colorectal cancer. Oncotarget 9:3830
Acknowledgements
Not applicable.
Funding
The authors declare that they have no funding support.
Author information
Authors and Affiliations
Contributions
TY, JFS conceived and designed the study. WW and DSL performed the literature search and data extraction. XXY, AJ, YDM and ZF drafted the manuscript. All authors had read and approved the final manuscript.
Corresponding author
Ethics declarations
Conflict of interest
The authors declare that they have no conflicts of interest.
Ethical approval
The experimental protocol was established, according to the ethical guidelines of the Helsinki Declaration and was approved by the Animal Ethics Committee of First Affiliated Hospital of Jinzhou Medical University.
Consent for publication
The authors agree to publication in the Journal.
Additional information
Publisher's Note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Rights and permissions
About this article
Cite this article
Yang, T., Sun, J., Wang, W. et al. Hsa_circ_0006732 regulates colorectal cancer cell proliferation, invasion and EMT by miR-127-5p/RAB3D axis. Mol Cell Biochem 477, 2751–2760 (2022). https://doi.org/10.1007/s11010-022-04458-5
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s11010-022-04458-5