Abstract
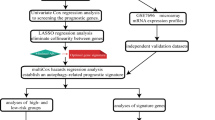
Autophagy is a highly conserved lysosomal degradation process essential in tumorigenesis. However, the involvement of autophagy-related long noncoding RNAs (lncRNAs) in low-grade glioma (LGG) remains unclear. In this study, we established an autophagy-related lncRNA prognostic signature for patients with LGG and assess its underlying functions. We used univariate Cox, least absolute shrinkage and selection operator and multivariate Cox regression models to establish an autophagy-related lncRNA prognostic signature. Kaplan–Meier survival analysis, receiver operating characteristic curve, nomogram, C-index, calibration curve and clinical decision-making curve were used to assess the predictive capability of the identified signature. A signature comprising nine autophagy-related lncRNAs (AL136964.1, ARHGEF26-AS1, PCED1B-AS1, AS104072.1, PRKCQ-AS1, LINC00957, AS125616.1, PSMB8-AS1 and AC087741.1) was identified as a prognostic model. Patients with LGG were divided into the high- and low-risk cohorts based on the median model-based risk score. The survival analysis revealed a 10-year survival rate of 9.3% (95% CI 1.91–45.3%) and 13.48% (95% CI 4.52–40.2%) in high-risk patients in the training and validation sets, respectively, and 48.4% (95% CI 24.7–95.0%) and 48.4% (95% CI 28.04–83.4%) in low-risk patients in the training and validation sets, respectively. This finding suggested a relatively low survival in high-risk patients. In addition, the lncRNA signature was independently prognostic and potentially associated with the progression of LGG. Therefore, the 9-autophagy-related-lncRNA signature may play a crucial role in the diagnosis and treatment of LGG, which may offer new avenues for tumour-targeted therapy.








Similar content being viewed by others
Data availability
All the data generated or analysed during this study have been included in this article.
References
Li Y, Deng G, Qi Y, Zhang H, Jiang H, Geng R, Ye Z, Liu B, Chen Q (2020) Downregulation of LUZP2 is correlated with poor prognosis of low-grade glioma. Biomed Res Int 2020:9716720. https://doi.org/10.1155/2020/9716720
Bao Z, Wang Y, Wang Q, Fang S, Shan X, Wang J, Jiang T (2021) Intratumor heterogeneity, microenvironment, and mechanisms of drug resistance in glioma recurrence and evolution. Front Med. https://doi.org/10.1007/s11684-020-0760-2
Bandopadhayay P, Bergthold G, London W, Goumnerova L, Morales La Madrid A, Marcus K, Guo D, Ullrich N, Robison N, Chi S, Beroukhim R, Kieran M, Manley P (2014) Long-term outcome of 4,040 children diagnosed with pediatric low-grade gliomas: an analysis of the Surveillance Epidemiology and End Results (SEER) database. Pediatr Blood Cancer 61:1173–1179. https://doi.org/10.1002/pbc.24958
Lin H, Patel S, Affleck V, Wilson I, Turnbull D, Joshi A, Maxwell R, Stoll E (2017) Fatty acid oxidation is required for the respiration and proliferation of malignant glioma cells. Neuro Oncol 19:43–54. https://doi.org/10.1093/neuonc/now128
Duffau H, Taillandier L (2015) New concepts in the management of diffuse low-grade glioma: proposal of a multistage and individualized therapeutic approach. Neuro Oncol 17:332–342. https://doi.org/10.1093/neuonc/nou153
Xue W, Zhang J, Tong H, Xie T, Chen X, Zhou B, Wu P, Zhong P, Du X, Guo Y, Yang Y, Liu H, Fang J, Wang S, Wu H, Xu K, Zhang W (2020) Effects of BMPER, CXCL10, and HOXA9 on neovascularization during early-growth stage of primary high-grade glioma and their corresponding MRI biomarkers. Front Oncol 10:711. https://doi.org/10.3389/fonc.2020.00711
Jiang L, Xiao C, Xu Q, Sun J, Chen H, Chen Y, Yin X (2017) Analysis of DTI-derived tensor metrics in differential diagnosis between low-grade and high-grade gliomas. Front Aging Neurosci 9:271. https://doi.org/10.3389/fnagi.2017.00271
Yang Y, Liu X, Cheng L, Li L, Wei Z, Wang Z, Han G, Wan X, Wang Z, Zhang J, Chen C (2020) Tumor suppressor microRNA-138 suppresses low-grade glioma development and metastasis via regulating IGF2BP2. Onco Targets Ther 13:2247–2260. https://doi.org/10.2147/ott.S232795
Zhao J, Wang L, Wei B (2020) Identification and validation of an energy metabolism-related lncRNA-mRNA signature for lower-grade glioma. Biomed Res Int 2020:3708231. https://doi.org/10.1155/2020/3708231
Louis D, Perry A, Reifenberger G, von Deimling A, Figarella-Branger D, Cavenee W, Ohgaki H, Wiestler O, Kleihues P, Ellison D (2016) The 2016 World Health Organization Classification of tumors of the central nervous system: a summary. Acta Neuropathol 131:803–820. https://doi.org/10.1007/s00401-016-1545-1
Zhang M, Wang X, Chen X, Zhang Q, Hong J (2020) Novel immune-related gene signature for risk stratification and prognosis of survival in lower-grade glioma. Front Genet 11:363. https://doi.org/10.3389/fgene.2020.00363
Fisher M, Jones D, Li Y, Guo X, Sonawane P, Waanders A, Phillips J, Weiss W, Resnick A, Gosline S, Banerjee J, Guinney J, Gnekow A, Kandels D, Foreman N, Korshunov A, Ryzhova M, Massimi L, Gururangan S, Kieran M, Wang Z, Fouladi M, Sato M, Øra I, Holm S, Markham S, Beck P, Jäger N, Wittmann A, Sommerkamp A, Sahm F, Pfister S, Gutmann D (2021) Integrated molecular and clinical analysis of low-grade gliomas in children with neurofibromatosis type 1 (NF1). Acta Neuropathol 141:605–617. https://doi.org/10.1007/s00401-021-02276-5
Tu Z, Wu L, Wang P, Hu Q, Tao C, Li K, Huang K, Zhu X (2020) N6-methylandenosine-related lncRNAs are potential biomarkers for predicting the overall survival of lower-grade glioma patients. Front Cell Dev Biol 8:642. https://doi.org/10.3389/fcell.2020.00642
Liu W, Li C, Xu W, Liu X, Tang H, Huang H (2020) Genome-wide analyses of the prognosis-related mRNA alternative splicing landscape and novel splicing factors based on large-scale low grade glioma cohort. Aging 12:13684–13700. https://doi.org/10.18632/aging.103491
Xie C, Liu X, Sham K, Lai J, Cheng C (2014) Silencing of EEF2K (eukaryotic elongation factor-2 kinase) reveals AMPK-ULK1-dependent autophagy in colon cancer cells. Autophagy 10:1495–1508. https://doi.org/10.4161/auto.29164
Jutten B, Keulers T, Peeters H, Schaaf M, Savelkouls K, Compter I, Clarijs R, Schijns O, Ackermans L, Teernstra O, Zonneveld M, Colaris R, Dubois L, Vooijs M, Bussink J, Sotelo J, Theys J, Lammering G, Rouschop K (2018) EGFRvIII expression triggers a metabolic dependency and therapeutic vulnerability sensitive to autophagy inhibition. Autophagy 14:283–295. https://doi.org/10.1080/15548627.2017.1409926
Wang H, Liu Y, Wang D, Xu Y, Dong R, Yang Y, Lv Q, Chen X, Zhang Z (2019) The upstream pathway of mTOR-mediated autophagy in liver diseases. Cells. https://doi.org/10.3390/cells8121597
Gu Y, Han J, Jiang C, Zhang Y (2020) Biomarkers, oxidative stress and autophagy in skin aging. Ageing Res Rev 59:101036. https://doi.org/10.1016/j.arr.2020.101036
Su S, Wang X, Xi X, Zhu L, Chen Q, Zhang H, Qin Y, Yang B, Che N, Cao H, Zhong W, Wang B (2021) Phellodendrine promotes autophagy by regulating the AMPK/mTOR pathway and treats ulcerative colitis. J Cell Mol Med. https://doi.org/10.1111/jcmm.16587
Zhang Y, Gao R, Zhang L, Geng Y, Chen Q, Chen X, Liu X, Mu X, Ding Y, Wang Y, He J (2021) AMPK/mTOR downregulated autophagy enhances aberrant endometrial decidualization in folate-deficient pregnant mice. J Cell Physiol. https://doi.org/10.1002/jcp.30408
Mukhopadhyay S, Panda P, Sinha N, Das D, Bhutia S (2014) Autophagy and apoptosis: where do they meet? Apoptosis 19:555–566. https://doi.org/10.1007/s10495-014-0967-2
Mariño G, Niso-Santano M, Baehrecke E, Kroemer G (2014) Self-consumption: the interplay of autophagy and apoptosis. Nat Rev Mol Cell Biol 15:81–94. https://doi.org/10.1038/nrm3735
Greabu M, Giampieri F, Imre M, Mohora M, Totan A, Pituru S, Ionescu E (2020) Autophagy, one of the main steps in periodontitis pathogenesis and evolution. Molecules. https://doi.org/10.3390/molecules25184338
Zois C, Koukourakis M (2009) Radiation-induced autophagy in normal and cancer cells: towards novel cytoprotection and radio-sensitization policies? Autophagy 5:442–450. https://doi.org/10.4161/auto.5.4.7667
Salminen A, Kaarniranta K, Kauppinen A (2013) Beclin 1 interactome controls the crosstalk between apoptosis, autophagy and inflammasome activation: impact on the aging process. Ageing Res Rev 12:520–534. https://doi.org/10.1016/j.arr.2012.11.004
Vlahakis A, Debnath J (2017) The interconnections between autophagy and integrin-mediated cell adhesion. J Mol Biol 429:515–530. https://doi.org/10.1016/j.jmb.2016.11.027
Shen H, Wang L, Xiong J, Ren C, Gao C, Ding W, Zhu D, Ma D, Wang H (2019) Long non-coding RNA CCAT1 promotes cervical cancer cell proliferation and invasion by regulating the miR-181a-5p/MMP14 axis. Cell Cycle 18:1110–1121. https://doi.org/10.1080/15384101.2019.1609829
Liu L, Zhu S, Jiang X, Ren J, Lin Y, Zhang N, Tong M, Zhang H, Zheng W, Fu H, Luo H, Lin L, Yan J, Yang T (2017) LncRNA expression in CD4+ T cells in neurosyphilis patients. Front Cell Infect Microbiol 7:461. https://doi.org/10.3389/fcimb.2017.00461
Zhao W, Lin X, Han H, Zhang H, Li X, Jiang C, Feng M (2021) Long noncoding RNA H19 contributes to the proliferation and autophagy of glioma cells through mTOR/ULK1 pathway. NeuroReport 32:352–358. https://doi.org/10.1097/wnr.0000000000001602
Lv Q, Wang L, Li D, Lin Q, Shen X, Liu H, Li M, Ji Y, Qin C, Chen S (2020) Knockdown lncRNA DLEU1 inhibits gliomas progression and promotes temozolomide chemosensitivity by regulating autophagy. Front Pharmacol 11:560543. https://doi.org/10.3389/fphar.2020.560543
Zheng J, Wang B, Zheng R, Zhang J, Huang C, Zheng R, Huang Z, Qiu W, Liu M, Yang K, Mao Z, Ji A, Yuan Y (2020) Linc-RA1 inhibits autophagy and promotes radioresistance by preventing H2Bub1/USP44 combination in glioma cells. Cell Death Dis 11:758. https://doi.org/10.1038/s41419-020-02977-x
Ma R, Zhang B, Zhang Z, Deng Q (2020) LncRNA MALAT1 knockdown inhibits cell migration and invasion by suppressing autophagy through miR-384/GOLM1 axis in glioma. Eur Rev Med Pharmacol Sci 24:2601–2615
Liu C, Fu H, Liu X, Lei Q, Zhang Y, She X, Liu Q, Liu Q, Sun Y, Li G, Wu M (2018) LINC00470 coordinates the epigenetic regulation of ELFN2 to distract GBM cell autophagy. Mol Ther 26:2267–2281. https://doi.org/10.1016/j.ymthe.2018.06.019
Zheng J, Zhou Z, Qiu Y, Wang M, Yu H, Wu Z, Wang X, Jiang X (2021) A prognostic ferroptosis-related lncRNAs signature associated with immune landscape and radiotherapy response in glioma. Front Cell Dev Biol 9:675555. https://doi.org/10.3389/fcell.2021.675555
Chen Z, Tian H, Yao D, Zhang Y, Feng Z, Yang C (2021) Identification of a ferroptosis-related signature model including mRNAs and lncRNAs for predicting prognosis and immune activity in hepatocellular carcinoma. Front Oncol 11:738477. https://doi.org/10.3389/fonc.2021.738477
Chen W, Feng Z, Huang J, Fu P, Xiong J, Cao Y, Liu Y, Tu Y, Li Z, Jie Z, Xiao T (2021) Identification of ferroptosis-related long noncoding RNA and construction of a novel prognostic signature for gastric cancer. Dis Markers 2021:7724997. https://doi.org/10.1155/2021/7724997
Charoentong P, Finotello F, Angelova M, Mayer C, Efremova M, Rieder D, Hackl H, Trajanoski Z (2017) Pan-cancer immunogenomic analyses reveal genotype-immunophenotype relationships and predictors of response to checkpoint blockade. Cell Rep 18:248–262. https://doi.org/10.1016/j.celrep.2016.12.019
Wei J, Ge X, Tang Y, Qian Y, Lu W, Jiang K, Fang Y, Hwang M, Fu D, Xiao Q, Ding K (2020) An autophagy-related long noncoding RNA signature contributes to poor prognosis in colorectal cancer. J Oncol 2020:4728947. https://doi.org/10.1155/2020/4728947
Guo J, Li Y, Duan H, Yuan L (2019) LncRNA TUBA4B functions as a competitive endogenous RNA to inhibit gastric cancer progression by elevating PTEN via sponging miR-214 and miR-216a/b. Cancer Cell Int 19:156. https://doi.org/10.1186/s12935-019-0879-x
Pellecchia S, Sepe R, Decaussin-Petrucci M, Ivan C, Shimizu M, Coppola C, Testa D, Calin G, Fusco A, Pallante P (2020) The long non-coding RNA Prader Willi/Angelman region RNA5 (PAR5) is downregulated in anaplastic thyroid carcinomas where it acts as a tumor suppressor by reducing EZH2 activity. Cancers. https://doi.org/10.3390/cancers12010235
Liang H, Su X, Wu Q, Shan H, Lv L, Yu T, Zhao X, Sun J, Yang R, Zhang L, Yan H, Zhou Y, Li X, Du Z, Shan H (2020) 2810403D21Rik/MirfLncRNA promotes ischemic myocardial injury by regulating autophagy through targeting. Autophagy 16:1077–1091. https://doi.org/10.1080/15548627.2019.1659610
Zheng J, Pang C, Du W, Wang L, Sun L, Xing Z (2020) An allele of rs619586 polymorphism in MALAT1 alters the invasiveness of meningioma via modulating the expression of collagen type V alpha (COL5A1). J Cell Mol Med 24:10223–10232. https://doi.org/10.1111/jcmm.15637
Chen Z, Wang W, Huang W, Fang K, Sun Y, Liu S, Luo X, Chen Y (2017) The lncRNA HOTAIRM1 regulates the degradation of PML-RARA oncoprotein and myeloid cell differentiation by enhancing the autophagy pathway. Cell Death Differ 24:212–224. https://doi.org/10.1038/cdd.2016.111
Gao Z, Fu P, Yu Z, Zhen F, Gu Y (2019) Comprehensive analysis of lncRNA–miRNA–mRNA network ascertains prognostic factors in patients with colon cancer. Technol Cancer Res Treat 18:1533033819853237. https://doi.org/10.1177/1533033819853237
Zhou D, Gao B, Yang Q, Kong Y, Wang W (2019) Integrative analysis of ceRNA network reveals functional lncRNAs in intrahepatic cholangiocarcinoma. BioMed Res Int 2019:2601271. https://doi.org/10.1155/2019/2601271
Qin J, Zhu T, Wu W, Chen H, He Y (2021) Long non-coding RNA PCED1B-AS1 promotes the progression of clear cell renal cell carcinoma through miR-484/ZEB1 axis. OncoTargets Ther 14:393–402. https://doi.org/10.2147/ott.S270149
Fan F, Chen K, Lu X, Li A, Liu C, Wu B (2020) Dual targeting of PD-L1 and PD-L2 by PCED1B-AS1 via sponging hsa-miR-194-5p induces immunosuppression in hepatocellular carcinoma. Hepatol Int. https://doi.org/10.1007/s12072-020-10101-6
Yao Z, Zhang Q, Guo F, Guo S, Yang B, Liu B, Li P, Li J, Guan S, Liu X (2020) Long noncoding RNA PCED1B-AS1 promotes the Warburg effect and tumorigenesis by upregulating HIF-1α in glioblastoma. Cell Transplant 29:963689720906777. https://doi.org/10.1177/0963689720906777
Yang J, Yu D, Liu X, Changyong E, Yu S (2020) LncRNA PCED1B-AS1 activates the proliferation and restricts the apoptosis of glioma through cooperating with miR-194-5p/PCED1B axis. J Cell Biochem 121:1823–1833. https://doi.org/10.1002/jcb.29417
Li M, Cui J, Niu W, Huang J, Feng T, Sun B, Yao H (2019) Long non-coding PCED1B-AS1 regulates macrophage apoptosis and autophagy by sponging miR-155 in active tuberculosis. Biochem Biophys Res Commun 509:803–809. https://doi.org/10.1016/j.bbrc.2019.01.005
Shademan M, Naseri Salanghuch A, Zare K, Zahedi M, Foroughi M, Akhavan Rezayat K, Mosannen Mozaffari H, Ghaffarzadegan K, Goshayeshi L, Dehghani H (2019) Expression profile analysis of two antisense lncRNAs to improve prognosis prediction of colorectal adenocarcinoma. Cancer Cell Int 19:278. https://doi.org/10.1186/s12935-019-1000-1
Cui G, Zhao H, Li L (2020) Long noncoding RNA PRKCQ-AS1 promotes CRC cell proliferation and migration via modulating miR-1287-5p/YBX1 axis. J Cell Biochem 121:4166–4175. https://doi.org/10.1002/jcb.29712
Li X, Sun L, Wang X, Wang N, Xu K, Jiang X, Xu S (2021) A five immune-related lncRNA signature as a prognostic target for glioblastoma. Front Mol Biosci 8:632837. https://doi.org/10.3389/fmolb.2021.632837
Zhang L, Li L, Zhang P, Cai Y, Hua D (2019) LINC00957 acted as prognostic marker was associated with fluorouracil resistance in human colorectal cancer. Front Oncol 9:776. https://doi.org/10.3389/fonc.2019.00776
Zhang H, Zhu C, He Z, Chen S, Li L, Sun C (2020) LncRNA PSMB8-AS1 contributes to pancreatic cancer progression via modulating miR-382-3p/STAT1/PD-L1 axis. J Exp Clin Cancer Res 39:179. https://doi.org/10.1186/s13046-020-01687-8
Hu T, Wang F, Han G (2020) LncRNA PSMB8-AS1 acts as ceRNA of miR-22-3p to regulate DDIT4 expression in glioblastoma. Neurosci Lett 728:134896. https://doi.org/10.1016/j.neulet.2020.134896
Shen G, Mao Y, Su Z, Du J, Yu Y, Xu F (2020) PSMB8-AS1 activated by ELK1 promotes cell proliferation in glioma via regulating miR-574-5p/RAB10. Biomed Pharmacother 122:109658. https://doi.org/10.1016/j.biopha.2019.109658
Funding
This study was not funded by anybody.
Author information
Authors and Affiliations
Contributions
AM and MT: Conceptualization, methodology, validation, investigation, supervision, software, visualization, writing—original draft, writing—reviewing and editing. YW: Validation, supervision, investigation. MA: Visualization, methodology. LJ: Data curation, software, validation. XW: Methodology, formal analysis, visualization. YM: Software, validation. YA: Data curation, visualization, validation. ZF: Investigation, validation. MK: Investigation, writing—reviewing and editing.
Corresponding author
Ethics declarations
Conflict of interest
The authors declare no conflicts of interest.
Ethical approval
Not required to this study.
Consent to participate
Not required to this study.
Consent for publication
Not required.
Additional information
Publisher's Note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary Information
Below is the link to the electronic supplementary material.
11010_2022_4368_MOESM3_ESM.tif
Supplementary Figure 3 Kaplan–Meier curves of OS differences stratified by age (A), tumour grade(B), sex(C), new tumour events(D), radiation therapy(E), tumour type(F), IDH1 R132 mutation status(G) and TP53 mutation status(H) between the high- and low-risk groups in the TCGA entire set (TIF 3914 kb)
11010_2022_4368_MOESM4_ESM.tif
Supplementary Figure 4 A. Estimation of immune and stromal cells in low-grade glioma tissues using expression data in training set. Violin plots of tumour purity for the low- and high-risk groups based on the immune score, stromal score and ESTIMATE score. B. Differences in infiltration levels of immune cell types in the high- and low-risk groups in training set. C. Comparison of immune-related functions between the high- and low-risk groups in training set. D. Comparison of the expression of immune checkpoints between the high- and low-risk groups in training set (TIF 3874 kb)
11010_2022_4368_MOESM5_ESM.tif
Supplementary Figure 5 A–D Differences in estimation of stromal and immune cells, immune cell infiltration, immune-related functions, immune checkpoints and risk scores in validation set (TIF 6324 kb)
11010_2022_4368_MOESM6_ESM.tif
Supplementary Figure 6 A–E. In the CGGA mRNA-seq-693 dataset, the overall survival probability of patients with low expression levels of ARHGEF26-AS1 and PRKCQ-AS1 was significantly lower than that of patients with high expression levels (P < 0.05). C. In the CGGA mRNA-seq-693 dataset, the overall survival probability of patients with high expression levels of PCED1B-AS1 was significantly lower than that of patients with low expression levels (P < 0.05). D. In the CGGA mRNA-seq-693 dataset, no statistical difference was observed in the overall survival probability between patients with high and low expression levels of LINC00957. E. In the GSE16011 dataset, the overall survival probability was significantly higher in patients with high PRKCQ-AS1 expression levels than in patients with low expression levels (P < 0.001) (TIF 2887 kb)
Rights and permissions
About this article
Cite this article
Maimaiti, A., Tuerhong, M., Wang, Y. et al. An innovative prognostic model based on autophagy-related long noncoding RNA signature for low-grade glioma. Mol Cell Biochem 477, 1417–1438 (2022). https://doi.org/10.1007/s11010-022-04368-6
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s11010-022-04368-6